Beyond Cre-Lox: How Next-Gen PCE Systems Are Revolutionizing Chromosome Engineering
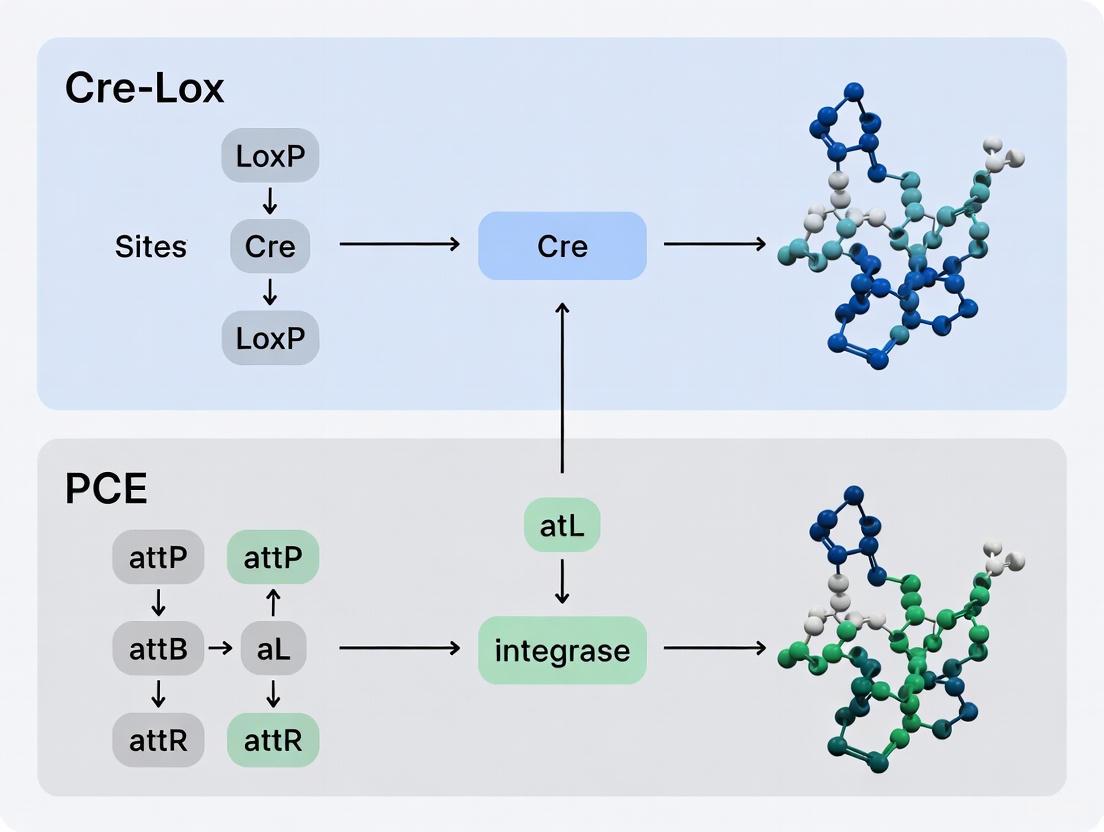
This article provides a comprehensive analysis for researchers and drug development professionals on the pivotal shift from the foundational Cre-Lox system to the novel Programmable Chromosome Engineering (PCE) platforms.
Beyond Cre-Lox: How Next-Gen PCE Systems Are Revolutionizing Chromosome Engineering
Abstract
This article provides a comprehensive analysis for researchers and drug development professionals on the pivotal shift from the foundational Cre-Lox system to the novel Programmable Chromosome Engineering (PCE) platforms. We explore the fundamental principles, historical limitations of traditional recombinase technologies, and the groundbreaking methodologies underpinning new systems like PCE and RePCE. The content details their applications in achieving megabase-scale edits, directly compares their efficiency and precision against established tools, and discusses the immediate implications for therapeutic development, crop engineering, and synthetic biology, offering a forward-looking perspective on the future of large-scale genome manipulation.
The Foundations of Large-Scale DNA Editing: From Cre-Lox to a New Era
The Cre-Lox system, derived from bacteriophage P1, has served as a cornerstone of genetic manipulation for decades, enabling researchers to achieve precise spatial and temporal control over gene expression in model organisms [1] [2]. This technology functions through the Cre recombinase enzyme, which recognizes and catalyzes recombination between specific 34-base pair DNA sequences known as loxP sites [2]. The orientation of these loxP sites dictates the genetic outcome: sites in the same direction cause deletion of the intervening DNA, while sites in opposite orientations lead to inversion [1]. The system's enduring utility is evidenced by its extensive application across thousands of publications, facilitating the generation of conditional knockout mice, lineage tracing, and complex genetic models that have profoundly advanced biomedical research [1] [3].
Despite its proven versatility, the classic Cre-Lox system faces significant challenges, including recombination reversibility, efficiency limitations with large DNA fragments, and residual "scar" sequences left after editing [4] [5]. These limitations have prompted intensive efforts to redesign and optimize the system. Recent breakthroughs have led to the development of advanced technologies like Programmable Chromosome Engineering (PCE) systems, which integrate engineered Cre variants with novel editing strategies to overcome these historical constraints [4] [5] [6]. This guide objectively compares the performance of the foundational Cre-Lox tool with its modern successors, providing experimental data and methodologies to inform researchers' experimental design.
Performance Comparison: Cre-Lox vs. Programmable Chromosome Engineering (PCE) Systems
The following tables summarize key performance characteristics based on recent, systematic studies and the latest technological developments.
Table 1: Efficiency and Precision Comparison
| Performance Parameter | Classic Cre-Lox System | New PCE Systems (PCE/RePCE) |
|---|---|---|
| Optimal Recombination Distance | 1-4 kb (wildtype loxP); 1-3 kb (mutant loxP) [1] [3] | Kilobase to megabase scales [4] [6] |
| Maximum Deletion Demonstrated | Not systematically quantified for large scales | 4 Mb in human cells [4] [5] |
| Maximum Insertion Demonstrated | Limited by efficiency decline | 18.8 kb targeted integration [4] [5] |
| Maximum Inversion Demonstrated | Not systematically quantified for large scales | 12 Mb in human cells; 315 kb in rice [4] [5] [6] |
| Recombination Reversibility | High, due to symmetric Lox sites [4] [5] | >10-fold reduction with novel asymmetric Lox variants [4] [5] |
| Editing Scar | Residual LoxP sites remain [4] [5] | Scarless editing via Re-pegRNA strategy [4] [5] |
Table 2: Experimental and Practical Factors
| Factor | Classic Cre-Lox System | New PCE Systems |
|---|---|---|
| Cre Recombinase Efficiency | Wildtype baseline [4] | AiCErec-engineered variant with 3.5x higher efficiency [4] [7] [5] |
| Key Influencing Factors | Cre-driver strain, inter-loxP distance, zygosity, breeder age (8-20 weeks optimal) [1] [3] | Programmable insertion position and orientation of Lox sites [4] |
| Critical Failure Distance | ≥15 kb (wildtype loxP); ≥7 kb (mutant lox71/66) [1] [3] | Not reported for upper limits |
| Typical Editing Workflow | Multi-step breeding or complex plasmid design [8] | Direct, programmable editing in plant and animal cells [4] [6] |
| Reported Mosaicism | Significant, especially with increasing loxP distance [1] [3] | Presumed higher efficiency reduces mosaicism; requires further validation |
Optimized Experimental Protocols
Protocol: Systematic Analysis of Cre-Mediated RecombinationIn Vivo
This protocol is adapted from a comprehensive 2025 study optimizing Cre-Lox in mice [1] [3].
- Objective: To quantitatively assess the efficiency of Cre-Lox recombination and the degree of mosaicism influenced by inter-loxP distance and Cre-driver strain.
- Key Reagents:
- Mouse Strains: 11 novel floxed strains generated at the Rosa26 locus on a C57BL/6J background, with inter-loxP distances of 0.8, 4, 8, 10, 12, and 15 kb.
- Cre-Drivers: Three robust Cre-driver strains (Ella-cre, CMV-cre, Sox2-cre).
- Methodology:
- Breeding Scheme: Cross female Cre-driver mice with male R26-floxed mice. This leverages Cre activity in the female germline to ensure all F1 progeny are exposed to Cre recombinase.
- Genotyping & Analysis: Genotype 8-55 F1 offspring from 1-8 litters. Categorize outcomes as:
- Complete Recombination: Successful deletion in all cells.
- Mosaicism: Mixed population of recombined and non-recombined cells.
- No Recombination: Floxed allele remains intact.
- Data Quantification: Calculate the percentage of offspring in each category for every Cre-driver and inter-loxP distance combination.
- Key Control: Include a positive control strain (e.g., Ella-cre) known to mediate widespread, though often mosaic, recombination.
Protocol: Programmable Chromosome Engineering for Large-Scale Mutagenesis
This protocol is based on the landmark 2025 study describing PCE systems [4] [5].
- Objective: To achieve scarless, precise manipulation of large DNA segments (insertions, deletions, inversions) in plant and animal cells.
- Key Reagents:
- Engineered Enzymes: AiCErec-optimized Cre variant (3.5x wildtype efficiency).
- Novel Recombination Sites: Asymmetric Lox variants with minimized reversibility.
- Editing System: Prime editors and specifically designed Re-pegRNAs for scar removal.
- Methodology:
- Target Site Preparation: Use prime editing to insert programmed Lox sites with specific orientations into the genome at desired locations.
- Recombination: Express the high-efficiency AiCErec-Cre variant in the cells to catalyze recombination between the inserted Lox sites, executing the desired large-scale edit (e.g., deletion, inversion).
- Scar Removal (RePCE): Employ Re-pegRNAs to direct prime editors to precisely reverse the residual Lox sites back to the original genomic sequence, resulting in a seamless edit.
- Validation: Use long-read sequencing and PCR to confirm the precise structural variation and the absence of unwanted sequence alterations.
- Proof-of-Concept Application: Creation of herbicide-resistant rice by programming a precise 315-kb inversion at the endogenous locus [4] [6].
Workflow and System Diagrams
Cre-Lox Recombination Workflow and Outcomes
Cre-Lox Mediated Gene Deletion: This diagram illustrates the fundamental process of Cre-Lox mediated gene knockout. A "floxed" allele, containing a target gene flanked by loxP sites in the same orientation, is exposed to Cre recombinase. The enzyme catalyzes a recombination event between the two loxP sites, resulting in the excision and deletion of the floxed gene, leaving behind a single residual loxP site in the genome [1] [2] [9].
PCE System Workflow for Scarless Editing
Programmable Chromosome Engineering (PCE/RePCE) Workflow: This diagram outlines the multi-step process for precise, large-scale genome engineering using PCE systems. The process begins with the programmable insertion of Lox sites. The AI-optimized Cre recombinase is then induced to perform the large DNA manipulation. For applications requiring ultimate precision, the RePCE system adds a final step where residual Lox "scars" are seamlessly removed, restoring the original sequence [4] [5].
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagents for Cre-Lox and Advanced Recombinase Research
| Reagent / Tool | Function & Application | Examples / Key Types |
|---|---|---|
| Cre Recombinase | Executes recombination at loxP sites. | Wildtype Cre, iCre (improved mammalian expression), CreERT2 (tamoxifen-inducible) [2] [9]. |
| Engineered Cre Variants | Enhanced efficiency and specificity for advanced applications. | AiCErec-Cre (3.5x efficiency) [4] [7]. |
| loxP Sites | 34 bp target sequences for Cre recombinase. | Wildtype loxP, lox66/lox71 (asymmetric, for irreversible recombination) [4] [2]. |
| Novel Lox Variants | Reduce reaction reversibility in next-gen systems. | Asymmetric Lox sites from PCE systems [4] [5]. |
| Floxed Alleles | Target gene or sequence flanked by loxP sites. | Conditional knockout mice (e.g., at Rosa26 locus), reporter strains [1] [3] [9]. |
| Cre-Driver Lines | Genetically defined models that express Cre in specific cell types/times. | Ella-cre, CMV-cre, Sox2-cre (ubiquitous); tissue-specific promoters (e.g., Ctsk-Cre) [1] [3] [9]. |
| TAx9 Sequence | Synthetic DNA element that prevents spurious Cre recombination in plasmid propagation in E. coli. | Used in single-plasmid Cre-loxP system construction [8]. |
| Prime Editors & Re-pegRNAs | Enable precise insertion of lox sites and subsequent scar removal in PCE systems. | Components of the PCE and RePCE toolkits [4] [5]. |
The Cre-Lox system remains an indispensable tool in the genetic engineer's arsenal, with its performance parameters and optimization strategies now more clearly defined than ever [1] [3]. However, the emergence of Programmable Chromosome Engineering (PCE) systems marks a significant evolutionary leap, addressing long-standing limitations of efficiency, scalability, and precision [4] [6]. While the classic Cre-Lox system continues to be the pragmatic choice for many conditional genetic manipulations within its optimal size range, PCE technologies unlock new possibilities for synthetic biology, crop improvement, and modeling complex genetic disorders. For the research and drug development community, this comparison underscores a transitional moment: the legacy of Cre-Lox provides a robust foundation, but its engineered successors are poised to redefine the frontiers of precise genome manipulation.
Recombinase systems are powerful tools in genetic engineering that enable precise manipulation of DNA structure and function through site-specific recombination. These systems facilitate fundamental genomic operations—excision, inversion, and translocation—by leveraging specialized enzymes that recognize and catalyze recombination at specific DNA target sequences. The Cre-lox system, derived from bacteriophage P1, represents one of the most versatile and widely adopted recombinase platforms, revolutionizing our ability to study gene function in complex organisms [10]. Since its development in the 1980s, Cre-lox technology has been effectively used to modify genomes in plants, insects, fish, and mammals including mice [10].
The core mechanism involves a single enzyme, Cre recombinase, that recombines a pair of short target sequences called loxP sites [11]. Each loxP site consists of 34 base pairs, featuring two 13 bp inverted repeats that serve as Cre binding regions and an asymmetric 8 bp core sequence that confers directionality [12]. The strategic placement and orientation of these loxP sites relative to each other determines the specific genetic outcome—whether a DNA segment will be excised, inverted, or translocated [10]. This elegant simplicity, requiring no additional co-factors for efficient recombination, has made the Cre-lox system an indispensable tool for mammalian gene editing [12].
The expanding genetic toolbox now includes several alternative recombinase systems, each operating on a similar conceptual framework but with distinct recognition sites and functional characteristics. Among these are the Flp-frt system from S. cerevisiae, the Dre-rox system from the D6 bacteriophage, and various inducible versions such as CreERT that respond to exogenous stimuli like tamoxifen [10]. These systems can be broadly categorized into constitutive systems, where recombination occurs continuously, and inducible systems, where temporal control enables researchers to trigger genetic changes at specific developmental timepoints [10]. The advent of intersectional genetics, which strategically leverages multiple recombinase systems simultaneously, has further enhanced our capability to manipulate genomes with precision unattainable by any individual system alone [10].
Core Mechanisms of DNA Rearrangement
Molecular Principles of Site-Specific Recombination
The fundamental mechanism underlying recombinase systems involves a carefully orchestrated process of DNA cleavage and rejoining at specific target sequences. Cre recombinase, a 38-kDa tyrosine recombinase, functions as the molecular scissors that execute these precise genetic operations [12]. The process initiates when Cre proteins bind to the 13 bp palindromic sequences within two separate loxP sites, forming a protein-DNA complex that brings the target sites into proximity [11]. Each Cre recombinase subunit then catalyzes a strand cleavage and exchange reaction through a Holliday junction intermediate, eventually resulting in complete recombination of the DNA between the recognition sites [11].
The specific genomic alteration that occurs depends entirely on the relative position and orientation of the loxP sites, which dictate the structural outcome of the recombination event [10]. This elegant system enables researchers to program different genetic rearrangements by strategically placing loxP sequences in predetermined configurations within the genome. The recombination process is remarkably efficient and does not require additional cellular factors or energy cofactors like ATP, making it highly suitable for genetic engineering applications across diverse biological systems [12].
Table 1: Molecular Components of Major Recombinase Systems
| Recombinase System | Origin | Recognition Site | Site Length | Key Features |
|---|---|---|---|---|
| Cre-loxP | Bacteriophage P1 | loxP | 34 bp | Most widely used; high efficiency in mammalian cells |
| Flp-frt | S. cerevisiae | frt | 48 bp | Temperature-sensitive; lower efficiency than Cre |
| Dre-rox | D6 bacteriophage | rox | 32 bp | Significant homology with Cre; minimal cross-reactivity |
Excision, Inversion, and Translocation Mechanisms
The strategic placement of loxP sites enables three primary types of DNA rearrangements, each with distinct experimental applications:
DNA Excision: When two loxP sites flank a DNA segment in the same orientation on the same chromosome, Cre-mediated recombination results in the excision of the intervening sequence as a circular molecule, while the original DNA molecule is left with a single loxP site [10]. This excision mechanism forms the basis for conditional gene knockout strategies, where essential genes can be deleted in specific tissues or at specific developmental stages without causing embryonic lethality [11]. The efficiency of excision is highly dependent on the distance between loxP sites, with optimal recombination occurring when sites are spaced less than 4 kb apart [1].
DNA Inversion: When loxP sites are oriented in opposite directions on the same chromosome arm, Cre recombinase mediates the inversion of the intervening DNA segment [10]. This inversion mechanism allows researchers to switch the orientation of genomic elements, enabling studies of promoter directionality, chromatin organization effects on gene expression, and the functional consequences of reversing specific DNA regions. Unlike excision events which are permanent, inversion events are theoretically reversible, though in practice the equilibrium often favors one orientation.
DNA Translocation: When loxP sites are located on different DNA molecules (such as separate chromosomes) and oriented in the same direction, Cre recombinase facilitates translocation events through reciprocal exchange between chromosomes [10]. This enables modeling of chromosomal rearrangements relevant to human diseases, including balanced translocations found in various cancers and genetic disorders. Translocation events can generate novel fusion genes or disrupt endogenous genes at breakpoints, replicating the genetic alterations observed in pathological conditions.
Diagram 1: Core recombination mechanisms showing how loxP site orientation and location determine genetic outcomes.
Comparative Analysis of Recombinase Systems
Performance Metrics and Efficiency Data
Recent systematic studies have quantitatively evaluated the performance parameters of Cre-lox and alternative recombinase systems, providing researchers with essential data for experimental design. A comprehensive 2025 analysis of Cre-mediated recombination in mice created 11 novel strains with conditional alleles at the Rosa26 locus to determine factors influencing recombination efficiency [1]. The findings demonstrated that the choice of Cre-driver strain plays a pivotal role in recombination efficiency, irrespective of the inter-loxP distance [1]. Furthermore, recombination was most successful when loxP sites were separated by distances ranging from 1 to 4 kb for wildtype loxP sites, or 1 to 3 kb when working with mutant loxP variants [1].
The efficiency of recombination is also influenced by the zygosity of the floxed allele, with heterozygous floxed alleles showing more efficient recombination than homozygous alleles when crossed with Cre-driver mice [1]. Additionally, the age of the Cre-driver mouse at breeding significantly impacts outcomes, with optimal recombination observed in breeders aged 8-20 weeks [1]. These findings provide a crucial framework for optimizing Cre-mediated recombination, enhancing experimental precision, improving reproducibility, and ultimately reducing the time and cost of genome modification.
Table 2: Performance Comparison of Recombinase Systems in Mammalian Cells
| System | Recombination Efficiency | Temperature Sensitivity | Inducible Versions | Cross-Reactivity |
|---|---|---|---|---|
| Cre-loxP | High efficiency; optimal with 1-4 kb distance | Stable at 37°C | CreERT, CreERT2 (tamoxifen) | Minimal with other systems |
| Flp-frt | Lower efficiency than Cre | Sensitive (>30°C); variants (FLPe, FLPo) improved | Limited availability | Minimal with other systems |
| Dre-rox | Efficient in mice | Stable at 37°C | Limited availability | Generally absent; some reports at high expression |
Advanced and Next-Generation Recombinase Systems
The expanding repertoire of recombinase tools now includes sophisticated inducible systems that provide precise temporal control over recombination events. The CreERT system utilizes a Cre-estrogen receptor (ERT) fusion protein that remains sequestered in the cytoplasm until the administration of tamoxifen induces nuclear translocation and recombination activity [10]. An improved version, CreERT2, demonstrates approximately ten-fold greater sensitivity to the synthetic ligand 4-hydroxytamoxifen (4-OHT) in vivo compared to the original CreERT [12]. This enhanced sensitivity allows for more precise temporal control with lower inducer concentrations, reducing potential side effects.
Recent innovations have pushed the boundaries of precision through optogenetic recombinase systems. The newly developed REDMAPCre represents a cutting-edge red-light-controlled split-Cre system that enables rapid activation (1-second illumination) and achieves an 85-fold increase in reporter expression over background levels [13]. This system addresses limitations of previous photoactivatable Cre tools, including low recombination efficiency, background activation, slow activation kinetics, and poor tissue penetration [13]. REDMAPCre utilizes 660 nm red light, which offers improved tissue penetration compared to blue/violet light-activated systems, enabling non-invasive DNA recombination in deep tissues without requiring implantable optical fibers [13].
The Tet-on and Tet-off systems provide complementary methods for controlling recombinase expression through antibiotic regulators. In the Tet-off system, a tetracycline-controlled transactivator protein (tTA) activates transcription of Cre in the absence of doxycycline, while the Tet-on system utilizes a reverse transactivator (rtTA) that requires doxycycline to activate Cre expression [10]. These systems enable reversible and quantitative control of recombination events, particularly useful for studying gene function during extended processes like development or disease progression.
Experimental Protocols and Methodologies
Standard Protocol for Cre-lox Recombination in Mice
The implementation of Cre-lox technology for conditional gene manipulation in mice follows a well-established breeding scheme with specific methodological considerations:
Strain Generation: Two separate mouse lines must be generated or obtained—a Cre-driver strain in which Cre recombinase is expressed under the control of a tissue-specific promoter, and a "floxed" strain in which the target gene is flanked by loxP sites [12].
Breeding Scheme: Female Cre driver mice are typically bred with male floxed mice to produce F1 offspring. This breeding scheme is preferred based on evidence that the Cre transgene shows higher activity in the female germline [1].
Genotype Validation: Between 1 to 8 litters, consisting of 8 to 55 total offspring, should be genotyped to assess the percentage of three possible outcomes: complete recombination, mosaicism, or no recombination [1]. PCR analysis with primers flanking the loxP sites typically confirms successful recombination.
Efficiency Optimization: To maximize recombination efficiency, breeders should be maintained at 8-20 weeks of age, and heterozygous floxed alleles should be used when possible [1]. The inter-loxP distance should be maintained at ≤4 kb for wildtype loxP sites or ≤3 kb for mutant loxP sites.
Diagram 2: Standard workflow for generating conditional knockout mice using Cre-lox technology.
Protocol for Inducible Cre-lox Recombination
For temporal control of genetic recombination, the CreERT system requires specific methodological adaptations:
Tamoxifen Preparation: Tamoxifen is typically dissolved in corn oil or sunflower oil at concentrations of 10-20 mg/mL. For the more sensitive CreERT2 variant, lower concentrations (1-5 mg/mL) may be sufficient [12].
Administration Route: Tamoxifen can be administered via intraperitoneal injection (most common), oral gavage, or in some cases through diet. Injection regimens vary from a single dose to multiple injections over consecutive days, depending on the experimental requirements.
Dosage Optimization: For CreERT2, effective doses typically range from 0.1-1 mg per 10g body weight for intraperitoneal injection [12]. Dosage should be optimized for each specific Cre line and target tissue.
Timing Considerations: The timing of tamoxifen administration depends on the biological question. For developmental studies, pregnant females may be injected at specific gestation timepoints; for adult studies, recombination is typically analyzed 2-14 days after the final tamoxifen injection.
Control Experiments: Critical controls include vehicle-only treated animals (oil without tamoxifen) and animals lacking the Cre transgene treated with tamoxifen, to control for potential non-specific effects of the compound.
The Scientist's Toolkit: Essential Research Reagents
Successful implementation of recombinase technologies requires access to specific biological reagents and methodological resources. The table below outlines essential components for designing and executing recombinase-based experiments.
Table 3: Essential Research Reagents for Recombinase-Based Studies
| Reagent Category | Specific Examples | Function/Application | Key Considerations |
|---|---|---|---|
| Cre-Driver Lines | Ella-cre, CMV-cre, Sox2-cre (constitutive); Various tissue-specific promoters (Table 1) | Provide spatial control of recombination | Choice significantly impacts efficiency; test multiple lines |
| Floxed Alleles | Rosa26 locus-targeted; Gene-specific conditional alleles | DNA target for recombination | Optimal recombination with 1-4 kb inter-loxP distance |
| Inducible Systems | CreERT, CreERT2 (tamoxifen); Tet-on/Tet-off (doxycycline) | Temporal control of recombination | CreERT2 offers 10x higher sensitivity than CreERT |
| Reporters | lacZ, Fluorescent proteins (GFP, tdTomato) | Visualization of recombination events | Critical for quantifying efficiency and patterns |
| Inducing Agents | Tamoxifen (4-OHT), Doxycycline | Activation of inducible systems | Dose optimization required for each system |
| Validation Tools | PCR primers flanking loxP sites; Antibodies for protein detection | Confirmation of successful recombination | Essential for quantifying mosaicism |
Recombinase systems have fundamentally transformed genetic engineering by providing precise control over DNA excision, inversion, and translocation events. The core mechanics of these systems—leveraging site-specific recombination between strategically positioned recognition sequences—enable researchers to address biological questions with unprecedented precision. While the Cre-lox system remains the gold standard in the field, emerging technologies like REDMAPCre offer exciting new capabilities for spatiotemporal control with minimal background activity [13].
The future of recombinase technologies lies in enhancing precision and reducing limitations. Current challenges include mosaicism, unpredictable recombination efficiency, and off-target effects [1]. Next-generation systems are addressing these limitations through improved engineering, such as the development of high-fidelity Cre variants with reduced cryptic site recognition and optimized inducible systems with faster kinetics and reduced background. The integration of recombinase systems with other genome editing technologies, particularly CRISPR-based approaches, promises to further expand their utility for complex genetic manipulations.
As these tools continue to evolve, they will undoubtedly unlock new possibilities for modeling human diseases, deciphering gene function, and developing novel therapeutic strategies. The systematic optimization of parameters influencing recombination efficiency—including inter-loxP distance, Cre-driver selection, and breeding conditions—will enhance reproducibility and reliability across research applications [1]. By providing researchers with an expanding toolkit for precise genetic manipulation, recombinase systems will continue to drive fundamental discoveries in biology and medicine.
For decades, the Cre-Lox system has served as a cornerstone of genetic engineering, enabling researchers to manipulate genes with spatial and temporal control. This technology, derived from the P1 bacteriophage, utilizes Cre recombinase to catalyze DNA recombination at specific loxP sites, facilitating conditional gene knockouts, insertions, and other complex genomic rearrangements. Despite its widespread adoption and transformative impact, the Cre-Lox system has faced significant challenges related to efficiency and precision that have limited its application in advanced chromosome engineering research. A comprehensive understanding of these historical limitations is crucial for contextualizing the development of next-generation genome editing tools, including newly described PCE (Precise Chromosome Engineering) systems. This guide objectively compares the traditional Cre-Lox system's performance against emerging alternatives, providing researchers with experimental data and methodological insights to inform their experimental designs.
Historical Limitations of Cre-Lox: A Systematic Analysis
Efficiency Constraints in Cre-Mediated Recombination
The efficiency of Cre-Lox recombination has been systematically evaluated across multiple studies, revealing critical limitations that impact experimental outcomes.
Table 1: Factors Affecting Cre-Lox Recombination Efficiency
| Factor | Impact on Efficiency | Experimental Findings |
|---|---|---|
| Inter-loxP Distance | Critical determinant | Recombination most successful with loxP sites spaced <4 kb; complete failure with wildtype loxP sites ≥15 kb or mutant lox71/66 sites ≥7 kb [1]. |
| Cre-Driver Strain | Pivotal role | Choice of strain significantly influences recombination efficiency regardless of inter-loxP distance [1]. |
| Zygosity of Floxed Allele | Moderate effect | Heterozygous floxed alleles yield more efficient recombination than homozygous alleles [1]. |
| Breeder Age | Contributing factor | Optimal recombination observed in breeders aged 8-20 weeks [1]. |
| Type of loxP Sites | Variable impact | Wildtype loxP sites generally more efficient than mutant variants [1]. |
Efficiency limitations extend beyond these parameters. The system demonstrates variable and often incomplete recombination with inter-loxP distances in the kilobase range, leading to mosaic animals where recombination occurs only in a subset of target cells. This mosaicism presents particular challenges for generating uniform animal models and can lead to unexpected phenotypic outcomes [1].
Precision Limitations and Off-Target Effects
The precision of the traditional Cre-Lox system is compromised by several biological and technical factors that affect its specificity and reliability.
Promoter Fidelity Issues: The specificity of Cre expression heavily depends on the cell-type fidelity of promoters driving its expression. Many tissue-specific promoters utilized to drive Cre expression are also active in unexpected cell types or at different developmental stages. For instance, pancreatic endocrine and ductal cell-specific promoters have been documented to express in brain neurons, liver, stomach, and intestines, resulting in lack of specificity of genetic recombination [14].
Ectopic Recombination: The problem of off-target recombination is not limited to pancreatic systems. A Lys2 promoter, widely used to delete genes in the myeloid lineage, is also active in type 2 pneumocytes in the lungs, potentially confounding respiratory studies [14].
Intercellular Transfer: Microvesicles-mediated transfer of Cre mRNA between neighboring cells can lead to unfaithful cellular labeling, causing false interpretation of lineage tracing and cell-specific knockout studies [14].
Background Activity in Plasmid Propagation: A fundamental technical limitation involves the difficulty of creating single plasmids containing both Cre recombinase and floxed sites, as Cre-mediated recombination spontaneously occurs when the plasmid is amplified in Escherichia coli bacterial cells. This has historically impeded the development of all-in-one genetic tools [8].
Experimental Methodologies for Assessing Limitations
Understanding the experimental approaches used to quantify these limitations provides valuable context for evaluating comparative studies.
Systematic Efficiency Analysis
A comprehensive multi-locus analysis of Cre-mediated recombination examined 12 different loci across different chromosomes. Researchers created 11 novel floxed strains with different inter-loxP site lengths integrated into the R26 locus using a high-efficiency Bxb1 recombinase system. The workflow involved breeding female Cre driver mice (Ella-cre, CMV-cre, and Sox2-cre) with male R26-floxed mice to produce F1 offspring. Between 1-8 litters (total 8-55 offspring) were genotyped to assess the percentage of three outcomes: complete recombination, mosaicism, or no recombination [1].
Table 2: Experimental Recombination Efficiency by Distance
| Inter-loxP Distance | Cre Driver Strain | Complete Recombination | Mosaicism | No Recombination |
|---|---|---|---|---|
| 0.8 kb | Ella-cre | 54% | 33% | 13% [1] |
| 4 kb | Ella-cre | 48% | 35% | 17% [1] |
| 8 kb | Ella-cre | 22% | 41% | 37% [1] |
| 15 kb | Ella-cre | 0% | 28% | 72% [1] |
Precision Assessment Methods
Studies evaluating precision limitations typically employ lineage tracing and reporter systems. One approach involves crossing Cre-driver lines with fluorescent reporter strains (e.g., Rosa26-lacZ or Rosa26-EYFP) and examining reporter expression across multiple tissues through immunohistochemistry and fluorescence imaging. Unanticipated reporter expression in non-target tissues indicates promoter infidelity or ectopic recombination [14].
Emerging Solutions and Comparative Systems
Recent technological innovations have sought to address these historical limitations through novel approaches to genome engineering.
TAx9 Technology for Improved Plasmid Construction
The TAx9 system represents a significant advancement for creating all-in-one Cre-LoxP plasmids. This artificial nucleic acid sequence (TATATATATATATATATA) effectively blocks spontaneous Cre-mediated recombination in E. coli cells when positioned upstream of tissue-specific Cre-driver sites. In practical applications, TAx9 technology enabled the creation of tamoxifen-inducible retinal pigment epithelium cell-labeled newts and skeletal muscle fiber cell-labeled mice in the F0 generation, dramatically reducing the time and cost for establishing genetically modified organisms [8].
Optogenetic Control with REDMAPCre
The REDMAPCre system addresses precision limitations through red-light-controlled split-Cre recombinase based on the ΔPhyA/FHY1 interaction. This technology enables rapid activation (1-second illumination) and achieves an 85-fold increase in reporter expression over background levels. Key advantages include:
- Spatiotemporal Precision: Enables remote control of recombination with high temporal resolution [13]
- Deep-Tissue Penetration: Red light (660 nm) offers improved tissue penetration compared to blue-light systems [13]
- Minimal Background Activity: Uses phycocyanobilin as a covalently bound chromophore, ensuring negligible background activity without illumination [13]
Multifunctional Reporter Systems
Advanced reporter systems have been developed that enable simultaneous evaluation of fluorescence, bioluminescence, and cell-killing assays within a single gene-modified line. These systems, successfully implemented in rat models using CRISPR/Cas12a, provide more comprehensive functional readouts while minimizing animal use [15].
Essential Research Reagent Solutions
Table 3: Key Research Reagents for Cre-Lox and Advanced Systems
| Reagent | Function | Applications |
|---|---|---|
| Cre Driver Strains | Provide Cre recombinase expression under specific promoters | Tissue-specific gene knockout; fate mapping [1] |
| Floxed Allele Strains | Contain loxP-flanked target sequences | Conditional gene deletion; lineage tracing [1] |
| TAx9 Element | Blocks spontaneous Cre recombination in E. coli | All-in-one plasmid construction; rapid F0 generation model creation [8] |
| REDMAPCre System | Red-light-controlled split-Cre recombinase | Spatiotemporal control of recombination; deep-tissue activation [13] |
| Cre-Dependent Reporters | Express markers upon Cre-mediated recombination | Visualization of recombination events; lineage tracing [15] |
| Tamoxifen | Induces nuclear translocation of CreERT2 | Temporal control of recombination; inducible gene knockout [16] |
Visualizing Experimental Workflows
The historical limitations of the Cre-Lox system in efficiency and precision have represented a significant "ceiling" for chromosome engineering research. Constraints in recombination efficiency related to inter-loxP distance, Cre-driver strain selection, and zygosity, coupled with precision issues including mosaicism, promoter infidelity, and ectopic recombination, have necessitated the development of advanced engineering solutions. Emerging technologies such as TAx9 for stable all-in-one plasmid construction and REDMAPCre for optogenetic control address these limitations while maintaining the fundamental utility of recombinase-mediated genome engineering. Understanding these historical constraints provides valuable context for evaluating next-generation PCE systems and their potential to overcome the limitations that have challenged researchers for decades.
For decades, life sciences research has been able to manipulate single genes with increasing precision. However, a significant frontier remained largely unconquered: the precise engineering of chromosomal segments at the megabase scale. Many genetic diseases, complex traits, and agronomically valuable characteristics arise from large-scale structural variations—deletions, duplications, inversions, and translocations spanning thousands to millions of base pairs. The lack of tools to manipulate these large DNA segments efficiently and precisely created a critical bottleneck in functional genomics, synthetic biology, and therapeutic development.
This article compares the established Cre-Lox system with a new generation of Programmable Chromosome Engineering (PCE) systems, examining how they address this fundamental unmet need and the quantitative performance gains they offer to researchers and drug development professionals.
The Bottleneck: Limitations of Traditional and Cre-Lox Systems
Traditional methods for large DNA modification, such as those relying on double-strand breaks (DSBs) and non-homologous end joining (NHEJ), are prone to introducing undesired insertions, deletions, and cellular damage, making them unreliable for precise, large-scale edits [4]. While the Cre-Lox system offered a pathway for large-segment manipulation, its adoption for megabase-scale engineering was hampered by several biological and technical constraints:
- Reversible Recombination: The inherent symmetry of standard Lox sites leads to forward and reverse reactions, making edits unstable and often reversing them to the original state [4] [17] [18].
- Limited Recombinase Efficiency: The activity of the wild-type Cre enzyme was often insufficient for reliably manipulating very large DNA segments [4] [7].
- Residual "Scar" Sequences: After recombination, leftover Lox sites in the genome could compromise gene function and lead to unintended secondary recombination events [4] [18].
- Inefficiency with Large Distances: Systematic studies in mice showed that recombination efficiency decreases dramatically as the distance between Lox sites increases. Complete recombination consistently failed with inter-LoxP distances of 15 kb or more [1].
The Paradigm Shift: Programmable Chromosome Engineering (PCE) Systems
To overcome these historical limitations, researchers developed PCE systems through three key innovations [4] [17] [18]:
- Engineered Recombination Sites: A high-throughput platform created novel, asymmetric Lox variants that reduce reversible recombination by over 10-fold, effectively locking edits in place [4] [17].
- AI-Optimized Recombinase (AiCErec): An AI-assisted protein engineering approach was used to optimize the Cre recombinase's multimerization interface, creating a variant with 3.5 times the recombination efficiency of the wild-type enzyme [4] [18].
- Scar-Free Editing Strategy (Re-pegRNA): This method leverages prime editors to precisely replace residual Lox sites with the original genomic sequence post-recombination, enabling seamless "scarless" edits [4] [17].
These innovations culminated in two integrated platforms, PCE and RePCE, designed for efficient, precise, and scarless manipulation of DNA from kilobase to megabase scales [4].
Direct Comparison: Cre-Lox vs. PCE Systems
The table below summarizes the performance and operational characteristics of traditional Cre-Lox systems versus the new PCE systems, based on published experimental data.
Table 1: Performance Comparison of Cre-Lox and PCE Systems
| Feature | Traditional Cre-Lox System | New PCE/RePCE Systems |
|---|---|---|
| Editing Precision | Leaves residual Lox "scars" [4] | Scarless editing via Re-pegRNA [4] [18] |
| Reaction Reversibility | High (reversible reactions) [4] | >10-fold reduction in reversibility [17] [18] |
| Recombinase Efficiency | Wild-type Cre efficiency [4] | 3.5x higher efficiency with AiCErec variant [4] [7] |
| Maximum Efficient Deletion | Fails at ≥ 15 kb in mice [1] | Up to 4 Mb demonstrated in plants [4] [17] |
| Maximum Efficient Inversion | Limited to smaller scales | Up to 12 Mb demonstrated [17] [18] |
| Targeted Insertion Capacity | Limited | Up to 18.8 kb [4] [17] |
| Key Limitation Overcome | Reversibility, scars, low efficiency | Stability, precision, and scale of edits |
Experimental Evidence: Protocols and Workflows
Key Workflow for PCE-Mediated Megabase-Scale Editing
The following diagram illustrates the core experimental workflow for achieving precise, large-scale edits using the PCE system.
Detailed Experimental Protocols:
Step 1: Insertion of Asymmetric Lox Variants
- Methodology: Researchers use prime editing with specifically designed pegRNAs (prime-editing guide RNAs) to install the novel, asymmetric Lox sites at precise genomic locations. This step is DSB-free, avoiding the cellular damage associated with traditional CRISPR-Cas9 cutting [4]. The Lox variants used are designed for high-efficiency forward recombination and minimal reverse reaction.
Step 2: Expression of the AiCErec Cre Variant
- Methodology: The engineered Cre recombinase, optimized via the AiCErec platform, is expressed in the cell. The AiCErec method integrates general inverse folding models with structural and evolutionary constraints to optimize the recombinase's multimerization interface, resulting in a variant with significantly enhanced activity [4] [18].
Step 3: Catalyzing Defined Recombination
Step 4: Scarless Editing via Re-pegRNA (RePCE)
- Methodology: In the RePCE system, a final step uses Re-pegRNAs to perform "re-prime editing" on the residual Lox sites left after recombination. This precisely reverts the Lox sequence back to the original genomic sequence, leaving no exogenous DNA behind and ensuring a seamless, scarless edit [4] [17].
The Scientist's Toolkit: Essential Research Reagents
The following table details key reagents and tools that are fundamental to implementing PCE systems, based on the materials cited in the foundational research.
Table 2: Key Research Reagent Solutions for PCE Systems
| Reagent/Tool | Function in the Experiment |
|---|---|
| Asymmetric Lox Variants | Engineered recombination sites that minimize reverse recombination, ensuring stable edits [4] [17]. |
| AiCErec-Optimized Cre Recombinase | High-efficiency enzyme variant that catalyzes recombination of large DNA segments with 3.5x greater efficiency than wild-type Cre [4] [18]. |
| Prime Editor (PE) System | A "search-and-replace" editing system used to install Lox sites without creating double-strand breaks [4]. |
| Re-pegRNA | A specially designed pegRNA that directs the prime editor to remove residual Lox sites after recombination, enabling scarless editing [4] [18]. |
| Bxb1 Recombinase System | An alternative, high-efficiency recombinase system used in some protocols for efficiently integrating large constructs flanked by Lox sites into specific genomic loci (e.g., Rosa26) [1]. |
The development of PCE systems represents a transformative advance in genome engineering. By directly addressing the historic limitations of the Cre-Lox system—instability, inefficiency, and lack of precision—these tools have unlocked the ability to manipulate chromosomes at a scale that truly matches the scope of many biological questions and challenges.
For basic researchers, this enables the functional study of large structural variants and the dissection of complex genetic networks. For applied fields, it dramatically accelerates the development of improved crop varieties [17] [7] and brings future therapeutic interventions for chromosomal disorders closer to reality. As these tools are adopted and refined, they are poised to become the new standard for precision chromosome engineering, pushing the boundaries of what is possible in synthetic biology and medicine.
Inside the PCE Breakthrough: Mechanisms and Real-World Applications
For decades, the Cre-Lox system has been a cornerstone of genetic engineering, enabling researchers to manipulate genes with spatial and temporal control in model organisms. This technology, derived from the P1 bacteriophage, utilizes Cre recombinase to catalyze recombination at specific loxP sites, allowing for gene deletions, inversions, and translocations [10]. Its application has been revolutionary, facilitating the generation of conditional knockout mice and advancing our understanding of gene function in fields from neuroscience to skeletal biology [19] [9]. However, this powerful tool has been hampered by persistent limitations: reversible recombination reactions that can undo edits, low efficiency especially with large DNA fragments, and the residual "scar" sequences (loxP sites) left behind after editing, which can compromise genomic integrity and precision [5] [20].
The recent development of the Programmable Chromosome Engineering (PCE) system represents a quantum leap forward. A team of researchers led by Prof. Gao Caixia has engineered a triad of innovations that systematically address the historical shortcomings of Cre-Lox technology [5] [21]. This guide deconstructs the PCE system, providing an objective, data-driven comparison with traditional Cre-Lox systems for researchers and drug development professionals evaluating the next generation of chromosome engineering tools.
Core Innovation 1: Asymmetric Lox Sites to Curb Reversible Recombination
A fundamental flaw of the traditional Cre-Lox system is the symmetry of standard loxP sites. This symmetry means the recombination reaction is readily reversible, often leading to an equilibrium where a significant proportion of cells fail to retain the desired genetic modification, thus reducing overall editing efficiency [5].
- PCE Innovation: The PCE system introduces novel, asymmetric lox site variants. This critical redesign disrupts the symmetry of the reaction, heavily favoring the forward (editing) direction.
- Performance Data: This engineering triumph results in a reduction of reversible recombination activity by over 10-fold, bringing it to levels nearly indistinguishable from negative controls [5].
Core Innovation 2: AiCErec - An AI-Engineered High-Efficiency Recombinase
Even with optimal lox sites, the efficiency of the wild-type Cre recombinase enzyme itself is a limiting factor, particularly for large-scale chromosomal manipulations. The tetrameric nature of Cre has made it difficult to optimize through traditional protein engineering approaches [5].
- PCE Innovation: The researchers developed AiCErec, an AI-informed protein engineering method. This platform integrates inverse folding models with structural and evolutionary constraints to intelligently guide the evolution of the Cre recombinase.
- Performance Data: The AiCErec platform yielded a superior Cre variant with a recombination efficiency 3.5 times that of the wild-type Cre enzyme [5] [20]. This dramatic enhancement is crucial for enabling robust editing across kilobase to megabase scales.
Core Innovation 3: Re-pegRNA for Scarless Editing Precision
A persistent issue in genome editing is the retention of foreign sequences, such as the 34 bp loxP sites, after recombination. These "scars" can potentially interfere with gene function and preclude truly seamless edits [5].
- PCE Innovation: The PCE system incorporates a Re-pegRNA-mediated scarless editing strategy. This method leverages the high precision of prime editors. After the initial recombination is complete, specifically designed pegRNAs (prime editing guide RNAs) direct the editing machinery to the residual lox sites, precisely replacing them with the original genomic sequence.
- Performance Data: This process ensures seamless genome modifications, leaving no trace of the editing machinery behind and fully restoring genomic integrity at the edited site [5].
Performance Comparison: PCE vs. Cre-Lox Systems
The following tables summarize key quantitative differences and editing capabilities between traditional Cre-Lox and the new PCE systems, based on published data.
Table 1: Direct Performance Metric Comparison
| Performance Metric | Traditional Cre-Lox System | New PCE System | Reference |
|---|---|---|---|
| Recombination Efficiency | Baseline (Wild-type Cre) | 3.5x improvement | [5] [20] |
| Reaction Reversibility | High (reversible equilibrium) | >10-fold reduction | [5] |
| Editing Scalability | Declines over a few kilobases [1] | Kilobase to Megabase scale | [5] [21] |
| Editing Precision | Leaves residual lox "scars" | Scarless edits | [5] |
Table 2: Demonstrated Editing Capabilities of PCE Systems
| Editing Type | Scale Demonstrated | Experimental Context |
|---|---|---|
| Targeted Insertion | Up to 18.8 kb | Plant and human cells [5] |
| Sequence Replacement | 5 kb | Plant and human cells [5] |
| Chromosomal Inversion | 315 kb (rice); 12 Mb (human) | Herbicide-resistant rice; human disease-related sites [5] [21] |
| Chromosomal Deletion | 4 Mb | Plant and human cells [5] |
| Chromosomal Translocation | Whole chromosomes | Plant and human cells [5] |
Experimental Workflow: From Design to Scarless Edit
The following diagram illustrates the streamlined experimental workflow for achieving a scarless edit using the RePCE system, integrating its three core innovations.
The Scientist's Toolkit: Essential Reagents for PCE
Table 3: Key Research Reagent Solutions for PCE System Implementation
| Reagent / Tool | Function in the PCE System |
|---|---|
| Asymmetric Lox Site Plasmids | Engineered DNA sequences that serve as the target for recombination, designed to minimize reaction reversibility. |
| AiCErec-Optimized Cre Recombinase | The high-efficiency, AI-engineered recombinase enzyme that catalyzes the large-scale DNA manipulation. |
| Re-pegRNA Constructs | Specialized guide RNAs that direct the prime editing machinery to remove residual lox sites after recombination, enabling scarless edits. |
| Prime Editor (PE) System | The core editing machinery (e.g., fusion of reverse transcriptase and Cas9 nickase) used in conjunction with Re-pegRNA to achieve the final, scarless sequence replacement. |
| Delivery Vector (e.g., AAV, Lentivirus) | A vehicle for efficiently introducing the PCE system components into the target cells (plant or human). |
The PCE system is not merely an incremental improvement but a fundamental redesign of recombinase-based editing. By tackling the three historic Achilles' heels of the Cre-Lox system—reversibility, inefficiency, and scarring—this triad of innovations (asymmetric lox sites, AiCErec, and Re-pegRNA) unlocks a new regime of precision and scale in chromosome engineering. For researchers in molecular breeding, therapeutic development, and synthetic biology, the PCE platform offers a transformative toolset for ambitious projects, from creating complex disease models with large structural variations to engineering crops with multi-gene trait stacks, all with unprecedented efficiency and fidelity.
For decades, the Cre-Lox system has been a cornerstone of genetic engineering, enabling conditional gene expression and basic genome modifications across model organisms. However, its translational potential has been hampered by fundamental limitations: reversible recombination reactions, suboptimal recombination efficiency, and residual "scar" sequences left in the genome after editing. Recent breakthroughs from Gao Caixia's lab have culminated in the development of Programmable Chromosome Engineering (PCE) systems—a suite of technologies that overcome these historical constraints through artificial intelligence-assisted recombinase engineering (AiCErec). This guide provides a comprehensive comparison between traditional Cre-Lox systems and these new PCE technologies, offering experimental data and methodologies for researchers navigating the evolving landscape of precision genome editing.
Technical Breakdown: Systematic Optimization of Recombinase Systems
Core Limitations of Traditional Cre-Lox Systems
The conventional Cre-Lox system, while versatile, presents significant challenges for precise chromosomal engineering. The inherent symmetry of LoxP sites creates bidirectional recombination activity, causing undesirable reversals of genetic modifications. Furthermore, wild-type Cre recombinase exhibits limited recombination efficiency, while the tetrameric nature of the enzyme complicates protein engineering efforts. After recombination, residual Lox sites remain in the genome, potentially disrupting gene function and regulatory elements [4] [17].
Systematic studies in mouse models have quantified additional constraints, revealing that recombination efficiency is highly dependent on inter-loxP distance. Effective recombination typically requires distances under 4 kb for standard loxP sites and under 3 kb for mutant lox variants. Complete recombination failure occurs with wildtype loxP sites spaced ≥15 kb apart or mutant lox71/66 sites spaced ≥7 kb apart [1]. Efficiency is further influenced by Cre-driver strain selection, zygosity of floxed alleles, and breeder age [1].
The AiCErec Engineering Framework
The AiCErec platform represents a paradigm shift in recombinase optimization, integrating three key technological innovations:
Asymmetric Lox Variants: Using high-throughput recombination site modification, researchers developed novel Lox variants with >10-fold reduction in reversible recombination while maintaining high forward recombination efficiency [4] [21] [17].
AI-Optimized Cre Recombinase: The AiCErec method employs a computational approach combining general inverse folding models with structural and evolutionary constraints to optimize Cre's multimerization interface, generating variants with 3.5-fold higher recombination efficiency than wild-type Cre [4] [6] [22].
Scarless Editing Strategy: Through Re-pegRNA-mediated re-prime editing, residual Lox sites are precisely replaced with the original genomic sequence, enabling truly seamless genome modifications [4] [17].
Table 1: Performance Comparison Between Wild-Type Cre and AiCErec-Optimized Systems
| Feature | Wild-Type Cre-Lox | AiCErec-Optimized System |
|---|---|---|
| Recombination Efficiency | Baseline | 3.5-fold increase [4] [21] |
| Reversibility | High (inherently symmetric) | >10-fold reduction [4] [17] |
| Editing Precision | Leaves residual Lox "scars" | Scarless editing via Re-pegRNA [4] [22] |
| Optimal Inter-site Distance | <4 kb (standard loxP) [1] | Kilobase to megabase scales [4] [21] |
| Primary Applications | Small-scale edits, conditional knockouts | Large-scale chromosomal manipulations [4] [6] |
Experimental Data and Workflow Analysis
Chromosomal-Scale Editing Capabilities
The PCE and RePCE systems enable unprecedented manipulation of chromosomal architecture, achieving editing efficiencies of up to 26.2% for precise, large-scale modifications [4]. The technology has been successfully demonstrated across both plant and human cell systems with the following documented achievements:
Table 2: Documented Chromosomal Engineering Feats Using PCE Systems
| Edit Type | Scale Demonstrated | Biological System | Application Outcome |
|---|---|---|---|
| Targeted Insertion | Up to 18.8 kb | Plants & human cells [4] [17] | Precise integration of large DNA cargo |
| DNA Replacement | Up to 5 kb | Plants & human cells [4] [17] | Complete gene replacement |
| Chromosomal Inversion | Up to 12 Mb | Plants & human cells [4] [21] | Creation of herbicide-resistant rice [6] [22] |
| Chromosomal Deletion | Up to 4 Mb | Plants & human cells [4] [17] | Large-scale genomic rearrangements |
| Chromosome Translocation | Whole chromosomes | Plants & human cells [4] [17] | Synthetic chromosome biology |
Experimental Workflow for AiCErec Implementation
The following diagram illustrates the integrated workflow for implementing AiCErec-enhanced chromosome engineering:
AiCErec-Enhanced Chromosome Engineering Workflow
Detailed Methodologies for Key Experimental Procedures
High-Throughput Recombination Site Engineering
The development of asymmetric Lox variants employed a rapid site-directed mutagenesis approach using specifically designed primers to generate diverse Lox site libraries. Two plasmid libraries (P1 and P2), each containing a single Lox site, were constructed and recombined in vitro. The resulting recombined plasmids were transformed into E. coli, and the Lox sites were sequenced via high-throughput methods to identify variants with reduced reversibility [4]. This systematic approach identified novel Lox variants with significantly improved performance characteristics compared to conventional symmetric sites.
AI-Assisted Recombinase Optimization Protocol
The AiCErec methodology adapts the previously developed AiCE platform, which integrates general inverse folding models with structural and evolutionary constraints specifically for recombinase engineering [4] [17]. The experimental protocol involves:
- Computational Analysis: Using structural modeling to identify key residues at the multimerization interface of Cre recombinase that influence catalytic efficiency without compromising structural integrity.
- Variant Generation: Creating Cre mutants with optimized interfacial interactions through site-directed mutagenesis.
- Efficiency Validation: Testing candidate variants in both plant (rice) and mammalian cell systems, measuring recombination efficiency via sequencing-based assays and functional outcomes. This process yielded the high-efficiency Cre variant with 3.5-times wild-type activity [4].
Scarless Editing Implementation
The Re-pegRNA technology integrates prime editing with recombinase systems to eliminate residual recombination sites. The protocol involves:
- Designing Re-pegRNAs: These specifically target residual Lox sites post-recombination for precise reversion to the original genomic sequence.
- Prime Editor Delivery: Co-delivering prime editors with Re-pegRNAs into cells following initial recombinase-mediated editing.
- Validation: Using sequencing to confirm precise replacement of Lox sites with native sequences, ensuring truly scarless edits [4] [17]. This approach expands targetable sites and provides a seamless platform for chromosome engineering.
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Research Reagents for Chromosome Engineering Studies
| Reagent / Tool | Function / Application | Example Use Case |
|---|---|---|
| AiCErec-Optimized Cre Variants | High-efficiency DNA recombination | 3.5x more efficient large fragment integration [4] [21] |
| Asymmetric Lox Variants | Directional recombination with minimal reversibility | Enabling stable megabase-scale chromosomal edits [4] [17] |
| Re-pegRNAs | Guide RNAs for scarless removal of residual sites | Precise restoration of original sequence post-editing [4] [22] |
| Prime Editing Systems | Precise small-scale edits to facilitate larger rearrangements | Installing Lox sites for subsequent recombinase steps [4] |
| Bxb1 Recombinase System | High-efficiency integration of large constructs | Rapid generation of floxed strains for recombination studies [1] |
| REDMAPCre System | Optogenetic control of recombination with red light | Spatiotemporal precision in recombination activation [13] |
The development of AiCErec and PCE systems represents a transformative advancement in genome engineering capabilities. While traditional Cre-Lox systems remain valuable for straightforward conditional knockout studies and smaller-scale modifications, the new PCE technologies offer unprecedented precision and scale for chromosomal engineering. The quantitative improvements—3.5-fold enhancement in recombination efficiency, >10-fold reduction in reversibility, and the capacity for megabase-scale manipulations—position these technologies as essential tools for future research in functional genomics, agricultural biotechnology, and therapeutic development.
For research teams considering implementation, the choice between systems depends on experimental objectives: traditional Cre-Lox suffices for standard conditional mutagenesis, while PCE systems enable complex chromosomal rearrangements, synthetic biology applications, and precise gene stacking for crop improvement. The integration of AI-guided protein engineering with advanced DNA editing technologies marks a new era in precision genetic manipulation, offering researchers previously unimaginable control over chromosomal architecture and function.
The Cre-loxP system, derived from bacteriophage P1, has been a cornerstone tool in genetic engineering for decades. This site-specific recombination system enables precise genomic modifications—including excision, inversion, integration, and translocation of DNA sequences—across diverse organisms, from bacteria to mammals [23] [24]. The system's core components consist of the Cre recombinase enzyme and its 34-base pair recognition sequence, loxP [25]. The loxP site is structured with two 13-bp inverted repeats flanking an asymmetric 8-bp spacer region that confers directionality to recombination events [23]. Cre functions as a tetramer, binding to two loxP sites to facilitate recombination within the spacer region through a Holliday junction intermediate [23].
Despite its widespread adoption, the traditional Cre-lox system has faced a fundamental limitation: the inherent symmetry of standard loxP sites leads to reversible recombination reactions [6] [5]. This reversibility means that after the desired genetic modification occurs, Cre can catalyze the reverse reaction, potentially reverting the edited genome to its original state and compromising the stability of genetic edits [26]. This challenge, combined with difficulties in engineering the tetrameric Cre complex and the problem of residual loxP "scars" remaining in the genome after editing, has limited the system's broader application [6] [5]. This article examines how novel asymmetric lox site designs overcome these limitations and compares their performance with traditional symmetric systems within the emerging context of Programmable Chromosome Engineering (PCE).
Breaking Symmetry: The Molecular Design of Asymmetric lox Sites
Fundamental Limitations of Symmetric lox Sites
Traditional loxP sites are palindromic in their inverted repeat elements, creating a symmetrical structure that allows Cre recombinase to bind and catalyze recombination with equal probability in both forward and reverse directions [23]. This molecular symmetry is the root cause of the reversibility problem that has plagued conventional Cre-lox applications. The 8-bp asymmetric core spacer, while providing directionality, does not prevent the reverse reaction once the initial recombination is complete [23]. In practical terms, this means that excision events can be followed by re-integration, and inversion events can spontaneously revert to the original orientation, creating genetic instability that is particularly problematic for long-term studies and therapeutic applications.
Previous efforts to address this limitation focused on developing mutant lox sites (lox66 and lox71) with alterations in their inverted repeats [25]. When these asymmetric mutants recombine, they form a double-mutant lox72 site with significantly reduced affinity for Cre recombinase, thereby minimizing reverse reactions [25]. While this approach demonstrated the potential of engineered asymmetry, it provided only a partial solution and lacked the comprehensive design principles needed for broad application across different genetic engineering contexts.
The Asymmetric lox Site Design Revolution
Recent breakthroughs have fundamentally reimagined lox site architecture through purposefully engineered asymmetric designs. Researchers led by Professor Gao Caixia at the Chinese Academy of Sciences have developed novel lox variants by systematically addressing the symmetry problem at the molecular level [6] [5]. Their approach involved creating a high-throughput platform for rapid recombination site modification and implementing a strategic asymmetric lox site design that disrupts the palindromic symmetry critical for bidirectional recombination [5] [26].
The key innovation lies in engineering lox variants where the structural symmetry is broken at the nucleotide level, creating molecular architectures that favor forward recombination while imposing steric or energetic barriers to the reverse reaction [6]. This design principle represents a significant departure from previous approaches that primarily focused on modifying the spacer region while maintaining symmetrical inverted repeats. The resulting asymmetric lox variants demonstrate a reduction in reversible recombination activity by over 10-fold compared to traditional loxP sites, approaching the background level of negative controls while maintaining high efficiency for forward recombination [6] [5]. This breakthrough effectively decouples the forward and reverse reactions that were intrinsically linked in the native system.
Comparative Performance: Quantitative Analysis of Editing Systems
Table 1: Performance Comparison of Traditional Cre-lox versus Advanced PCE Systems
| Editing Parameter | Traditional Cre-lox | PCE Systems (with Asymmetric lox sites) |
|---|---|---|
| Reversibility Rate | High (inherent to system) | >10-fold reduction (near background levels) [6] [5] |
| Recombination Efficiency | Baseline (wild-type Cre) | 3.5× improvement with AiCErec-engineered Cre [6] [5] |
| Editing Scale Range | Kilobase to megabase [24] | Kilobase to megabase scale [6] [21] |
| Maximum Integration Size | Not specified in results | 18.8 kb [5] [26] |
| Sequence Replacement Capacity | Not specified in results | 5 kb complete replacement [5] [26] |
| Maximum Deletion | Not specified in results | 4 Mb chromosomal deletions [5] [26] |
| Maximum Inversion | Not specified in results | 12 Mb chromosomal inversions [6] [5] |
| Editing Precision | Leaves residual loxP scars | Scarless editing via Re-pegRNA strategy [6] [5] |
Table 2: Key Experimental Achievements Enabled by Asymmetric lox Technology
| Organism/System | Achievement | Scale | Functional Outcome |
|---|---|---|---|
| Rice (Oryza sativa) | Precise chromosomal inversion | 315 kb | Herbicide-resistant germplasm [6] [5] |
| Human Cells | Chromosomal inversion | 12 Mb | Targeted manipulation of disease-related loci [21] |
| Human Cells | Chromosomal deletion | 4 Mb | Large-scale genomic removal [5] [26] |
| Human Cells | Whole-chromosome translocation | Entire chromosomes | Chromosome-level engineering [5] [26] |
| Plant and Animal Cells | Targeted DNA integration | 18.8 kb | Large fragment insertion [5] [26] |
Experimental Paradigms: Methodologies for Evaluating Asymmetric lox Systems
In Vitro Recombination Assay for Asymmetric lox Variants
The foundational methodology for evaluating novel asymmetric lox sites employs systematic in vitro recombination assays that directly quantify recombination efficiency and directionality [23]. The experimental workflow begins with the construction of plasmid substrates containing pairs of asymmetric lox sites flanking a reporter cassette, typically featuring a ∼1-kb DNA spacer fragment between strategically positioned restriction sites (e.g., BamHI-EcoRI) [23]. Researchers then incubate these substrate constructs with heterotetrameric Cre recombinase complexes—mixtures of wild-type and mutant Cre proteins—that enable recombination between asymmetric sites [23].
Following the recombination reaction, the DNA is extracted and subjected to restriction enzyme digestion and gel electrophoresis analysis. Successful recombination is quantified by examining the characteristic banding patterns that indicate specific recombination products. This approach enables direct comparison of recombination efficiency between symmetric and asymmetric lox configurations. The heterotetrameric Cre complex is essential for this process, as it facilitates functional cooperation between different Cre variants that collectively recognize the asymmetric site architectures [23]. This methodology provided the first experimental evidence that asymmetric lox sites could recombine efficiently when paired with appropriate Cre complexes, establishing the foundational principle for subsequent developments.
High-Throughput Screening Platform for lox Variant Characterization
To systematically characterize numerous asymmetric lox designs, researchers developed a high-throughput screening platform for rapid evaluation of recombination site modifications [6] [5]. This innovative approach involves synthesizing libraries of lox variants with systematic nucleotide substitutions, particularly focusing on the spacer region where asymmetry is engineered. Each variant is cloned into reporter constructs that generate measurable signals—typically fluorescent or selectable markers—upon successful recombination.
The core of this methodology involves transient transfection of lox variant constructs into model cell lines followed by Cre expression and subsequent quantification of recombination outcomes using flow cytometry or antibiotic selection. This enables parallel assessment of hundreds of lox designs for their recombination efficiency, orthogonality, and reversibility profiles. The platform incorporates negative selection counterscreens to identify variants with reduced reverse recombination activity, precisely quantifying the extent to which each asymmetric design minimizes undesirable backward reactions [5]. This high-throughput methodology was instrumental in identifying the novel asymmetric lox variants that demonstrate a greater than 10-fold reduction in reversible recombination compared to traditional loxP sites [6] [5].
AiCErec: AI-Driven Recombinase Engineering
A pivotal innovation complementing asymmetric lox development is AiCErec (AI-informed Constraints for protein Engineering for recombinases), an advanced protein engineering framework that integrates general inverse folding models with structural and evolutionary constraints [6] [5] [26]. This methodology begins with comprehensive structural analysis of the Cre tetrameric complex, identifying key interfacial residues that govern multimerization and DNA recognition. Researchers then employ machine learning algorithms to predict amino acid substitutions that optimize cooperative binding to asymmetric lox sites while maintaining structural stability.
The experimental validation phase involves directed evolution of Cre variants using the AiCErec model to guide library design, followed by iterative screening for enhanced recombination activity on asymmetric lox substrates [5]. The most promising candidates undergo thorough characterization of their recombination efficiency, specificity, and oligomerization properties. This approach yielded engineered Cre variants with 3.5 times the recombination efficiency of wild-type Cre [6] [5], providing the enhanced enzymatic activity necessary to maximize the performance of asymmetric lox systems.
Figure 1: Integrated Development Workflow for Advanced lox Systems. This diagram illustrates the systematic approach combining asymmetric lox design, AI-driven protein engineering, high-throughput screening, scarless editing strategies, and biological validation to overcome traditional Cre-lox limitations.
The Research Toolkit: Essential Reagents for Chromosome Engineering
Table 3: Essential Research Reagents for Advanced Chromosome Engineering
| Reagent/Solution | Function in Experimental Workflow | Key Features/Benefits |
|---|---|---|
| Asymmetric lox Variants | Engineered recognition sites for Cre recombinase | >10-fold reduction in reverse recombination; maintained forward recombination efficiency [6] [5] |
| AiCErec-Engineered Cre Variants | Enhanced recombinase enzymes | 3.5× higher recombination efficiency than wild-type Cre; optimized for asymmetric sites [6] [5] |
| Re-pegRNA System | Prime editing components for scarless removal of residual lox sites | Precise replacement of residual sites with original genomic sequence; enables seamless edits [6] [5] |
| Heterotetrameric Cre Complexes | Mixed recombinase complexes for asymmetric site recognition | Enable recombination between asymmetric lox sites; functional cooperation between Cre variants [23] |
| Programmable Chromosome Engineering (PCE) Platform | Integrated system for large-scale DNA manipulations | Enables kilobase to megabase scale edits; supports insertions, deletions, inversions, translocations [6] [21] |
| RePCE Platform | Enhanced version with scarless editing capability | Incorporates Re-pegRNA for precise removal of residual sequences; improved editing precision [5] [26] |
| High-Throughput lox Screening Platform | System for rapid evaluation of lox variants | Enables parallel assessment of recombination efficiency and reversibility for multiple designs [6] [5] |
The PCE Revolution: Programmable Chromosome Engineering with Asymmetric lox Sites
The integration of asymmetric lox sites with other technological advances has enabled the development of Programmable Chromosome Engineering (PCE) systems, representing a paradigm shift in large-scale genome manipulation [6] [21] [5]. PCE combines three key innovations: (1) asymmetric lox sites that minimize reversibility, (2) AiCErec-engineered Cre variants with enhanced efficiency, and (3) Re-pegRNA technology for scarless editing [5] [26]. This integrated approach enables unprecedented control over chromosomal modifications across scales ranging from kilobases to megabases.
The PCE platform demonstrates remarkable versatility in supporting diverse editing operations. It facilitates targeted integration of large DNA fragments up to 18.8 kb, complete replacement of 5-kb sequences, chromosomal inversions spanning 12 Mb, chromosomal deletions of 4 Mb, and even whole-chromosome translocations [5] [26]. The RePCE variant further enhances editing precision by employing specifically designed pegRNAs to perform re-prime editing on residual lox sites, precisely replacing them with the original genomic sequence to ensure truly seamless genome modifications [6] [5]. This capability addresses the long-standing challenge of unwanted "scar" sequences remaining in the genome after editing, which could potentially interfere with gene function or regulation.
Figure 2: Addressing Traditional Cre-lox Limitations with PCE Innovations. This diagram maps the specific solutions provided by Programmable Chromosome Engineering systems against the fundamental limitations of traditional Cre-lox technology.
The development of asymmetric lox sites represents a transformative advancement in genome engineering technology, effectively addressing the long-standing challenge of reversible recombination that has limited the traditional Cre-lox system. When integrated with AI-enhanced Cre variants and scarless editing technologies in PCE platforms, these asymmetric systems enable unprecedented precision and scale in chromosomal manipulations. The quantitative evidence demonstrates clear superiority over conventional approaches, with order-of-magnitude improvements in reversibility reduction and significant enhancements in recombination efficiency.
For researchers pursuing complex genetic engineering projects—particularly those requiring large-scale chromosomal rearrangements, multiple sequential edits, or long-term genetic stability—asymmetric lox-based systems now offer a compelling alternative to both traditional Cre-lox and CRISPR-based approaches. The successful application of this technology in creating herbicide-resistant rice through a precise 315-kb inversion [6] [5] exemplifies its transformative potential for crop improvement, while achievements in human cells, including 12-Mb inversions at disease-related loci [21], highlight its therapeutic relevance. As these technologies continue to evolve, asymmetric lox systems are poised to become indispensable tools in the advanced genome engineering toolkit, enabling new possibilities in basic research, therapeutic development, and agricultural biotechnology.
The ability to perform precise, scarless genome editing represents a paramount goal in genetic engineering, with profound implications for therapeutic development and functional genomics. For decades, the Cre-Lox system has served as a cornerstone technology for conditional gene manipulation, enabling spatial and temporal control of gene expression through recombination-mediated DNA rearrangement [1]. While powerful, this system faces inherent limitations including reversible recombination, unpredictable efficiency, and the persistent incorporation of exogenous "landing pad" sequences that create permanent scars in the genome [4] [5]. The recent convergence of prime editing technology with engineered recombinase systems has catalyzed a transformative shift toward truly scarless chromosomal engineering. This comparison guide examines the emerging paradigm of Programmable Chromosome Engineering (PCE) systems utilizing Re-pegRNA technology against traditional Cre-Lox approaches, providing researchers with experimental data and methodological frameworks to inform their genome editing strategies.
Technological Foundations: From Cre-Lox to Programmable Systems
The Cre-Lox System: Established Workhorse with Limitations
The Cre-Lox system, derived from bacteriophage P1, utilizes the Cre recombinase enzyme to catalyze recombination between specific 34-base pair LoxP sites [1]. The system's versatility enables targeted DNA excision, inversion, or integration depending on the orientation and placement of Lox sites. However, its fundamental architecture introduces several constraints that impact experimental outcomes and therapeutic applications.
The system exhibits distance-dependent efficiency degradation, with optimal recombination occurring when LoxP sites are spaced within 1-4 kilobases (kb) for wildtype sites and 1-3 kb for mutant variants [1]. Critical limitations include reversible recombination reactions due to symmetrical Lox sites, low recombination efficiency in complex genomic contexts, and residual Lox sequences that remain after editing, potentially disrupting native gene regulation [4] [5]. These "scars" can alter gene expression patterns and confound phenotypic interpretations, particularly in sensitive applications like disease modeling and gene therapy.
Prime Editing and Re-pegRNA: A Programmable Platform for Scarless Editing
Prime editing represents a "search-and-replace" genome editing technology that directly writes new genetic information into a target DNA site without requiring double-strand breaks (DSBs) [27]. The system employs a prime editing guide RNA (pegRNA) that both specifies the target site and encodes the desired edit, along with a fusion protein consisting of a Cas9 nickase and reverse transcriptase [28].
The recent development of Re-pegRNA (Re-prime editing guide RNA) technology addresses the critical challenge of residual recombination sites. This innovation enables the precise replacement of leftover Lox sites with the original genomic sequence, achieving truly scarless genome modifications [4] [5]. When integrated with engineered recombinase systems, Re-pegRNA facilitates the Programmable Chromosome Engineering (PCE) platform, which enables flexible programming of insertion positions and orientations for different Lox sites, allowing precise manipulation of DNA fragments ranging from kilobase to megabase scale [5].
Table 1: Core Component Comparison Between Editing Systems
| Component | Cre-Lox System | PCE/RePCE System |
|---|---|---|
| Core Enzyme | Cre recombinase | Engineered Cre variant + Prime editor |
| Targeting Molecule | None (pre-defined LoxP sites) | pegRNA/Re-pegRNA |
| Recognition Site | 34 bp LoxP sequence | pegRNA-defined genomic locus + Lox sites |
| Template Source | None (excision/inversion) or Donor vector (integration) | pegRNA-encoded template + Prime editor |
| Outcome | Site-specific recombination | Programmable, scarless edits |
Performance Comparison: Quantitative Analysis of Editing Outcomes
Editing Efficiency and Precision
Recent studies directly comparing optimized Cre-Lox systems with PCE platforms reveal significant differences in editing efficiency and precision across various genomic contexts. The conventional Cre-Lox system demonstrates highly variable recombination efficiency (0-54%) in mouse models, influenced by factors including inter-LoxP distance, genomic location, and Cre-driver strain [1]. This variability presents substantial challenges for experimental reproducibility and therapeutic applications.
In contrast, PCE systems incorporating AI-engineered Cre variants (AiCErec) demonstrate 3.5-fold higher recombination efficiency compared to wildtype Cre, while novel asymmetric Lox variants reduce reversible recombination by 10-fold [4] [21]. The integrated RePCE platform achieves remarkable precision across diverse editing modalities with minimal off-target effects, enabling scarless insertions up to 18.8 kb with efficiencies reaching 26.2% in plant and human cell applications [4].
Table 2: Quantitative Performance Metrics Across Editing Platforms
| Editing Metric | Traditional Cre-Lox | Optimized Cre-Lox | PCE/RePCE System |
|---|---|---|---|
| Max Insertion Size | ~4 kb (efficient) | ~7 kb (mutant Lox) | 18.8 kb |
| Deletion Capacity | ~15 kb (limited efficiency) | Not reported | 4 Mb |
| Inversion Capacity | Not systematically quantified | Not reported | 12 Mb |
| Editing Efficiency | 0-54% (distance-dependent) | 3.5× wildtype Cre | Up to 26.2% |
| Scar Formation | Persistent Lox sites | Persistent Lox sites | Scarless via Re-pegRNA |
| Reversibility | High (symmetrical Lox) | 10-fold reduction | Programmatically controlled |
Versatility in Editing Types and Scalability
The PCE platform demonstrates unprecedented versatility across multiple dimensions of genome engineering. While traditional Cre-Lox primarily facilitates excision and inversion events, PCE systems enable a comprehensive suite of precise chromosomal modifications including targeted integrations, complete sequence replacements, megabase-scale inversions, and interchromosomal translocations [4] [5]. This expanded capability directly addresses a critical bottleneck in synthetic biology and therapeutic development—the need for flexible, large-scale DNA manipulation without genotoxic double-strand breaks.
The scalability of PCE systems represents another significant advancement, supporting manipulations ranging from single-base substitutions to chromosomal megabase rearrangements within a unified experimental framework [21]. This continuum of precision across scales enables researchers to engineer everything as subtle as disease-associated single nucleotide polymorphisms to as dramatic as pathogenic chromosomal translocations, all while maintaining scarless outcomes through the Re-pegRNA mechanism.
Experimental Protocols: Methodological Framework for Scarless Editing
PCE Workflow Implementation
The implementation of Programmable Chromosome Engineering follows a systematic workflow that integrates prime editing, recombinase-mediated recombination, and scar removal:
Target Site Selection and Lox Site Engineering: Identify target genomic loci and design asymmetric Lox sites (e.g., Lox66/Lox71, LoxKR3/Lox71) to minimize reverse recombination [4]. These engineered variants demonstrate approximately 10-fold reduction in reversibility while maintaining high forward recombination efficiency.
Prime Editing-Mediated Lox Integration: Utilize prime editors to install Lox sites at target genomic positions. The optimized protocol employs:
- Editor Architecture: PEmax or PE4max systems for enhanced expression and nuclear localization
- Delivery Method: Lentiviral or plasmid-based delivery in human cells; Agrobacterium-mediated transformation in plants
- Culture Conditions: For mammalian cells, maintain at 37°C with 5% CO₂; include selection markers (e.g., puromycin) for 3-5 days post-transfection [4]
Recombinase-Mediated Chromosome Engineering: Introduce the engineered Cre variant (AiCErec-optimized) to catalyze recombination between installed Lox sites:
- Expression System: Use a strong constitutive promoter (e.g., CAG for mammalian cells, Ubiquitin for plants)
- Dosage Optimization: Titrate Cre expression to balance efficiency against potential cytotoxicity
- Temporal Control: Consider inducible systems (e.g., estrogen receptor fusion) for precise timing of recombination [4] [5]
Scar Removal via Re-pegRNA: Design and deliver Re-pegRNAs to precisely replace residual Lox sites with native genomic sequence:
- Re-pegRNA Design: The scaffold includes a primer binding site (PBS) of 10-15 nucleotides and an RT template encoding the original genomic sequence without Lox elements
- Editing Validation: Implement PCR screening with flanking primers and Sanger sequencing to confirm scarless outcomes [4]
Validation and Quality Control
Rigorous validation of scarless editing outcomes requires a multi-layered analytical approach:
Genomic DNA Analysis: Isolate genomic DNA from target tissues (not just tail biopsies) and perform PCR with primers flanking the edited region to distinguish recombined from unrecombined alleles [29].
Sequencing Confirmation: Implement Sanger sequencing of PCR amplicons to verify precise sequence restoration at former Lox sites. For complex edits, utilize long-read sequencing (Oxford Nanopore, PacBio) to capture full context.
Functional Assessment: Evaluate transcriptional and translational outcomes through RT-qPCR (assay regions upstream, within, and downstream of edited exons) and Western blot analysis where appropriate [29].
Cellular Phenotyping: For therapeutic applications, assess viability, proliferation, and disease-relevant functional endpoints to confirm intended phenotypic consequences without off-target effects.
Research Reagent Solutions: Essential Tools for Implementation
Successful implementation of scarless chromosome engineering requires carefully selected molecular tools and reagents. The following table catalogues essential research solutions for both traditional and advanced editing approaches:
Table 3: Essential Research Reagents for Chromosome Engineering
| Reagent Category | Specific Examples | Function & Application |
|---|---|---|
| Recombinase Enzymes | Wildtype Cre, AiCErec-engineered Cre variants | Catalyzes site-specific recombination between Lox sites |
| Editing Plasmids | PEmax, PE4max, pegRNA expression vectors | Prime editor component delivery and pegRNA expression |
| Lox Site Variants | LoxP, Lox66/Lox71, LoxKR3/Lox71, asymmetric Lox | Recombination site recognition with reduced reversibility |
| Delivery Systems | Lentiviral vectors, AAV, Agrobacterium (plants) | Efficient editor delivery to target cells |
| Selection Markers | Puromycin, Hygromycin, Fluorescent reporters | Enrichment of successfully edited cells |
| Validation Tools | Lox-specific primers, flanking PCR assays, Sanger sequencing | Confirmation of editing efficiency and precision |
| Cell Lines/Models | HEK293T, K562, mouse embryonic stem cells | Optimized model systems for editing validation |
The evolution from traditional Cre-Lox systems to programmable PCE platforms represents a paradigm shift in chromosome engineering capabilities. While established Cre-Lox methodologies remain valuable for straightforward excision and inversion experiments in permissive genomic contexts, PCE and RePCE systems offer superior efficiency, scalability, and precision for complex editing applications requiring scarless outcomes. The integration of Re-pegRNA technology directly addresses the longstanding challenge of residual recombination sites, enabling truly seamless genome modifications across kilobase to megabase scales.
Researchers should consider strategic implementation of these technologies based on specific experimental needs. For foundational genetic studies requiring tissue-specific or inducible modifications, traditional Cre-Lox systems provide proven, well-characterized platforms. However, for therapeutic development, synthetic biology applications, and precision breeding initiatives where scarless outcomes are imperative, PCE systems offer transformative potential. As these technologies continue to mature, their convergence with other emerging modalities like CRISPR-associated transposase systems and base editing will further expand the frontiers of programmable genome engineering, enabling unprecedented precision in rewriting the code of life.
Chromosome engineering, the ability to programmatically rearrange genetic material, represents a frontier in genetic research with profound implications for modeling disease and developing novel therapies. For decades, the Cre-Lox recombination system has been the cornerstone of precise genetic manipulation, enabling conditional gene knockout, inversion, and excision in a wide range of organisms [3] [10]. However, this established technology faces inherent limitations in programmability and scale. Recently, a new class of programmable chromosome engineering (PCE) systems, notably bridge recombinases, has emerged with demonstrated capability for megabase-scale rearrangements in human cells [30]. This comparison guide provides an objective analysis of both systems' performance, supported by experimental data, to inform researchers selecting tools for chromosome engineering applications.
Technology Comparison: Mechanisms and Capabilities
Cre-LoxP Recombination System
The Cre-Lox system, derived from bacteriophage P1, utilizes the Cre recombinase enzyme to catalyze recombination between specific 34-base pair sequences known as loxP sites [10] [31]. The orientation and placement of these loxP sites determine the recombination outcome: excision when sites face the same direction, inversion when oriented oppositely, and translocation when located on different DNA strands [10]. This system has been extensively optimized over decades, with studies establishing that recombination efficiency is highly dependent on inter-loxP distance, with 1-4 kb being optimal for complete recombination using wildtype loxP sites [3]. The system's versatility is enhanced through engineered variants like Cre-ERT, which permits temporal control through tamoxifen induction [10].
Bridge Recombinase Systems
Bridge recombinases represent a breakthrough in programmable genome architecture. These systems utilize a bridge RNA (bRNA) that directly mediates recombination between a donor DNA sequence and a target genomic locus without creating double-strand breaks [30]. The technology was recently engineered for human cell application through systematic optimization of the bacterial ISCro4 recombinase, including bRNA structural stabilization and protein engineering via deep mutational scanning [30]. This engineering cascade produced a system capable of facilitating megabase-scale rearrangements with efficiencies independent of DNA segment size, overcoming a critical limitation of previous technologies [30].
Comparative Performance Analysis
Table 1: Direct Comparison of Key Performance Metrics
| Performance Parameter | Cre-Lox System | Bridge Recombinase System |
|---|---|---|
| Programmability | Requires pre-inserted loxP sites | Fully programmable via bRNA design |
| Maximum Demonstrated Rearrangement Size | ~15 kb (with wildtype loxP) [3] | 0.93 Mb (inversion), 134 kb (excision) [30] |
| Recombination Efficiency | Highly dependent on inter-loxP distance (optimal 1-4 kb) [3] | Up to 20.2% insertion efficiency; size-independent [30] |
| Temporal Control | Available via inducible systems (e.g., Cre-ERT) [10] | Not yet demonstrated |
| Tissue Specificity | Available via tissue-specific promoters [31] | Not yet demonstrated |
| Multiplexing Capacity | Limited without multiple recombinase systems | Theoretically high via bRNA design |
| Clinical Translation | Extensive preclinical use | Early proof-of-concept (e.g., BCL11A enhancer excision) [30] |
Table 2: Experimental Optimization Parameters for Cre-Lox
| Optimization Factor | Optimal Condition | Impact on Efficiency |
|---|---|---|
| Inter-loxP Distance | 1-4 kb (wildtype loxP) [3] | No recombination at ≥15 kb [3] |
| loxP Variant | Mutant lox71/66 for 1-3 kb distances [3] | Expanded range with mutant sites |
| Cre-Driver Strain | Strain-dependent [3] | Profound impact on recombination efficiency |
| Animal Age | 8-20 weeks [3] | Critical factor for breeding efficiency |
| Zygosity | Heterozygous floxed alleles [3] | More efficient than homozygous |
| Genomic Context | Maintain ≤4 kb inter-loxP distance [3] | Ensures efficiency across chromosomal locations |
Experimental Applications and Protocols
Cre-Lox System: Herbicide-Resistant Rice Development
The Cre-Lox system has been successfully applied in plant biotechnology for generating marker-free transgenic crops. A specific protocol for developing herbicide-resistant rice illustrates this application:
Vector Construction: A double fluorescent protein-expressing Ac/Ds transposon vector was constructed containing the gene of interest (GOI) for herbicide resistance, flanked by loxP sites, along with GFP and mCherry markers and Cre recombinase under a constitutive promoter [32].
Transformation and Selection: Rice calli were transformed via Agrobacterium-mediated transformation, with initial selection using hygromycin. T-DNAs integrated into the plant genome, and Cre-mediated recombination excised the marker genes flanked by loxP sites [32].
Marker Excision: Successful recombination events were identified by screening for loss of fluorescent markers while retaining herbicide resistance. PCR confirmation verified complete excision of selection markers, producing marker-free transgenic plants [32].
Efficiency Assessment: The double fluorescent system provided reliable counterselection, with herbicide resistance confirmed through application assays and molecular analysis showing decreased expression of target genes in resistant lines [32].
Bridge Recombinase: Megabase-Scale Rearrangements
The protocol for implementing bridge recombinase systems demonstrates their unique capabilities:
System Engineering: The native bacterial bridge recombinase system was optimized through (1) ortholog screening identifying ISCro4 as the most active in human cells, (2) bRNA engineering by splitting into two molecules and structural stabilization, and (3) protein evolution via deep mutational scanning to identify enhancing mutations [30].
Targeting Strategy: A "reverse targeting" approach was implemented where the bRNA was programmed to use a sequence unique to the donor plasmid to target the genome, and a sequence from the genome to target the donor. This strategy reduced off-target integrations by over 90% [30].
Large-Scale Rearrangement: For megabase inversions, bRNAs were designed to target boundaries of the target region, with successful demonstration of 0.93 Mb inversion in human cells. Efficiency was quantified via PCR and sequencing across junction sites [30].
Therapeutic Validation: As proof-of-concept, the system was deployed to excise the BCL11A enhancer (for sickle cell disease therapy) and pathogenic trinucleotide repeat expansions, with efficiency quantified by functional rescue assays [30].
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Research Reagents for Chromosome Engineering
| Reagent/Solution | Function | Example Applications |
|---|---|---|
| Cre Driver Strains | Provide tissue-specific or inducible Cre expression | Spatial-temporal gene knockout studies [3] |
| Floxed Allele Models | Contain loxP-flanked target sequences | Conditional gene deletion; excision studies [31] |
| Bridge RNA (bRNA) | Programs target specificity for bridge recombinases | Megabase-scale inversions, excisions [30] |
| Optimized ISCro4 | Engineered bridge recombinase with enhanced activity | Large payload integration in human cells [30] |
| Reporter Models (e.g., GFP/mCherry) | Visualize recombination events | Efficiency quantification; lineage tracing [32] |
| Mutant loxP Variants | Enhance recombination efficiency for specific distances | Applications requiring 1-3 kb rearrangements [3] |
| Inducible Systems (Cre-ERT) | Enable temporal control of recombination | Developmental studies; avoiding embryonic lethality [10] |
Visualizing Mechanisms and Workflows
Diagram 1: Fundamental Mechanisms of Action (Width: 760px)
Diagram 2: Experimental Selection Workflow (Width: 760px)
The comparative analysis presented here reveals complementary strengths of Cre-Lox and bridge recombinase systems. The Cre-Lox system remains the preferred choice for applications requiring precise, conditional gene manipulation in specific tissues or developmental stages, particularly for rearrangements under 15 kb. Its extensive characterization, availability of diverse reporter strains, and inducible controls make it ideal for physiological studies in model organisms.
Conversely, bridge recombinase systems represent a transformative technology for chromosome engineering at unprecedented scales, enabling programmable megabase rearrangements that were previously infeasible. Their programmability via bRNA design eliminates the need for laborious precursor strain generation. While still in early stages of biological characterization, bridge recombinases offer particular promise for modeling chromosomal rearrangement disorders and therapeutic genome architecture interventions.
Strategic selection between these systems should be guided by experimental goals: Cre-Lox for precision physiology in established model systems, and bridge recombinases for large-scale genomic engineering and applications requiring multi-locus targeting without pre-installed recognition sites. As both technologies continue to evolve, their convergence may ultimately provide the comprehensive toolkit needed for complete mastery of genome architecture.
Addressing Technical Hurdles: A Comparative Look at Optimization Strategies
The Cre-Lox system, derived from bacteriophage P1, has been an indispensable tool in genetic engineering for decades, enabling precise spatial and temporal control over gene expression in a wide range of organisms [1]. At its core, the system consists of two components: the Cre recombinase enzyme and its 34-base pair loxP DNA target sites [33]. The utility of this system extends far beyond simple gene knockout studies, supporting complex genetic manipulations including conditional mutagenesis, chromosomal rearrangements, and lineage tracing in models ranging from plants to mammals [34] [1]. However, despite its widespread adoption and versatility, the Cre-Lox system has been plagued by a fundamental biochemical limitation: the reversibility of its recombination reaction [4].
This reversibility problem stems from the inherent symmetry of wild-type loxP sites. Each loxP site consists of two 13-bp inverted repeats flanking an 8-bp asymmetric spacer region that confers directionality [33]. When Cre recombinase acts on two identical loxP sites, it catalyzes a conservative recombination reaction that is fully reversible [34]. In practical terms, this means that the system can oscillate between pre- and post-recombination states, leading to unstable genetic modifications and heterogeneous experimental outcomes. This equilibrium poses particular challenges for applications requiring permanent genetic changes, such as the creation of stable transgenic lines or therapeutic gene editing. The scientific community has long recognized this limitation, prompting nearly three decades of research to engineer solutions that bias the system toward irreversible outcomes.
Traditional loxP and the Roots of Reversibility
The Biochemical Basis of Reversibility
The molecular mechanism of Cre-Lox recombination follows a well-characterized pathway that explains its inherent reversibility. Cre recombinase functions as a tetrameric complex, with each monomer binding to one of the 13-bp inverted repeats of two loxP sites [33]. The reaction proceeds through a Holliday junction intermediate where strand cleavage and exchange occur within the 8-bp spacer region [33]. Because wild-type loxP sites are perfectly symmetrical, the reaction is energetically favorable in both directions, resulting in a thermodynamic equilibrium between the pre- and post-recombination states [4] [34]. This fundamental property means that Cre-mediated excision, inversion, or integration events can—and do—reverse themselves in the presence of continued Cre expression.
The consequences of this reversibility are particularly evident in applications involving gene inversion. Unlike excision events—where one loxP site is removed from the genome, making the reaction irreversible—inversion events leave two loxP sites in the genome that can undergo repeated recombination cycles for as long as Cre is present [34]. This continuous cycling between states creates cellular mosaicism, where individual cells within a population exhibit different genetic states, complicating phenotypic analysis and data interpretation [1]. As noted in one study investigating Cre-mediated genetic inversion in mice, "inversion, on the other hand, yields two loxP sites that are indistinguishable from the original loxP pair due to the conservative nature of the recombination event, and recombination continues as long as Cre is present" [34].
Early Attempts to Overcome Reversibility
The first significant effort to address the reversibility problem came in 1995 with the development of the first-generation mutant lox sites, lox66 and lox71 [33]. These variants featured asymmetric mutations in the distal end of either the left (lox66) or right (lox71) inverted repeats (Table 1). The underlying strategy was elegant in its conception: when lox66 and lox71 recombine, they produce one wild-type loxP site and one double-mutant lox72 site with mutations in both inverted repeats [33] [34].
Table 1: Traditional lox Site Sequences
| Site Name | Sequence (5' to 3' leading strand) | Characteristics |
|---|---|---|
| loxP | ATAACTTCGTATA GCATACAT TATACGAAGTTAT | Wild-type site; fully reversible recombination |
| lox66 | ATAACTTCGTATA GCATACAT TATACGAAcggta | Mutation in last 5 bp of right inverted repeat |
| lox71 | taccgTTCGTATA GCATACAT TATACGAAGTTAT | Mutation in first 5 bp of left inverted repeat |
| lox72 | taccgTTCGTATA GCATACAT TATACGAAcggta | Double mutant with reduced Cre affinity |
The lox72 site was found to have markedly reduced affinity for Cre recombinase—up to 8-fold lower according to some studies [34]. This reduction in affinity was intended to create a system where the forward reaction (lox66 × lox71) would be strongly favored over the reverse reaction (loxP × lox72). Early applications in mouse embryonic stem cells appeared promising, with one study reporting that "inversion appeared irreversible in ES cell colonies" [34].
However, these first-generation solutions suffered from significant limitations. The reduction in reverse reaction rate was often insufficient for applications requiring absolute stability, particularly in systems with high or prolonged Cre expression. Additionally, the creation of a wild-type loxP site as a recombination product meant that the potential for unwanted recombination events persisted. Perhaps most problematic was the nomenclature confusion that emerged in the literature, with many studies incorrectly annotating these mutant sites, leading to difficulties in protocol replication and vector design [33]. Despite these limitations, lox66/lox71 represented an important proof-of-concept that asymmetric lox sites could partially mitigate the reversibility problem.
The New Generation: Advanced Low-Reversion Variants
Systematic Identification of Novel Recombination Sites
The latest breakthrough in addressing Cre-Lox reversibility comes from a comprehensive research effort published in 2025, which established a high-throughput platform for rapid recombination site (RS) modification [4]. This systematic approach represented a significant departure from earlier piecemeal efforts, enabling the screening and characterization of numerous novel RS variants. The researchers developed a robust strategy to rapidly engineer RSs that minimize reversibility while preserving high recombination efficiency [4].
Through this methodical analysis, the team identified novel Lox variants that demonstrate a 10-fold reduction in reversible recombination activity compared to traditional systems, with reversibility rates approaching the background level of negative controls [4] [6]. This dramatic improvement in forward reaction bias represents the most significant advancement in Cre-Lox stability since the initial development of lox66/lox71. Unlike earlier approaches that focused on random design and testing, this new methodology established systematic principles for RS optimization, addressing a fundamental limitation noted in the literature that "current Lox site optimization relies on random design and testing, often yielding suboptimal results due to the absence of systematic principles" [4].
AI-Assisted Recombinase Engineering (AiCErec)
Complementing the improved recombination sites, the researchers employed artificial intelligence to optimize the Cre recombinase enzyme itself [4] [6]. Their AI-assisted recombinase engineering (AiCErec) platform integrated general inverse folding models with structural and evolutionary constraints to precisely optimize Cre's multimerization interface [4]. This approach addressed a critical limitation of traditional Cre engineering efforts, which had primarily focused on altering target specificity rather than enhancing enzymatic activity [4].
The AiCErec platform yielded a engineered Cre variant with 3.5 times the recombination efficiency of the wild-type enzyme [4] [6] [7]. This enhancement is particularly significant because the tetrameric nature of Cre recombinase had previously complicated efforts to improve its activity through conventional protein engineering approaches [6]. The combination of high-efficiency Cre variants with low-reversion RSs creates a synergistic system that simultaneously addresses both the kinetic and thermodynamic limitations of traditional Cre-Lox systems.
Scarless Editing Strategies
A persistent limitation of traditional Cre-Lox systems has been the retention of "scars"—residual lox sites that remain in the genome after recombination. These leftover sequences can potentially interfere with gene expression or serve as substrates for unwanted recombination events [4] [6]. The new programmable chromosome engineering (PCE) systems address this problem through a scar-free editing strategy that leverages the high editing efficiency of prime editors [4].
This approach uses specifically designed Re-pegRNAs to perform re-prime editing on residual Lox sites, precisely replacing them with the original genomic sequence and ensuring truly seamless genome modifications [4] [6]. By eliminating both the reversibility problem and the scar sequence issue, this technology represents a comprehensive solution to the major limitations that have hampered traditional Cre-Lox applications for decades.
Direct Comparison: Performance Metrics and Experimental Data
Quantitative Assessment of Recombination Efficiency and Stability
Table 2: Performance Comparison of Traditional vs. New lox Systems
| Parameter | Traditional loxP | First-Generation lox66/71 | New Low-Reversion Variants |
|---|---|---|---|
| Reversibility Rate | High (fully reversible) | Moderate (up to 8-fold reduction) | 10-fold reduction vs. traditional |
| Recombination Efficiency | Baseline | Comparable to wild-type | 3.5× improvement with AiCErec Cre |
| Editing Precision | Leaves loxP scars | Leaves lox72 and loxP scars | Scarless via Re-pegRNA system |
| Editing Scale | Kilobase scale with decreasing efficiency | Limited by reversibility | Kilobase to megabase scale (up to 18.8 kb insertions, 4 Mb deletions) |
| Stability of Modifications | Low (reversible while Cre present) | Moderate (leakage possible) | High (approaching background levels) |
The performance advantages of the new low-reversion systems are demonstrated across multiple experimental applications. In one notable proof-of-concept, researchers used the technology to create herbicide-resistant rice through a precise 315-kb chromosomal inversion—a feat that would have been exceptionally challenging with traditional systems due to both efficiency and stability limitations [4] [6] [7]. The system has also demonstrated robust capability for targeted integration of large DNA fragments up to 18.8 kb, complete replacement of 5-kb DNA sequences, chromosomal inversions spanning 12 Mb, and chromosomal deletions of 4 Mb [4].
Experimental Workflow for Assessing Recombination Efficiency
The following diagram illustrates the key experimental workflow for evaluating lox variant performance:
Experimental Workflow for Evaluating lox Variant Performance
This high-throughput assessment method, adapted from published studies [4] [35], enables systematic quantification of recombination efficiency and reversibility across hundreds of lox variants simultaneously. The process involves constructing comprehensive lox variant libraries, cloning them into reporter plasmids, and transforming them into a host system such as yeast [35]. Following Cre induction—for example, using a β-estradiol-inducible system—the plasmid DNA is extracted and subjected to deep sequencing to precisely quantify recombination rates [35]. This method allows researchers to calculate both forward and reverse recombination rates, providing a comprehensive picture of reaction bias and efficiency.
Table 3: Essential Research Reagents for Advanced Cre-Lox Studies
| Reagent/Resource | Function/Application | Key Characteristics |
|---|---|---|
| AiCErec-engineered Cre variants | High-efficiency recombination | 3.5× recombination efficiency vs. wild-type Cre [4] |
| Novel low-reversion Lox variants | Stable genome editing | 10-fold reduction in reversible recombination [4] |
| Prime editing systems | Scarless editing via Re-pegRNA | Precise replacement of residual Lox sites [4] |
| Bxb1 recombinase system | High-efficiency transgene integration | Enables precise insertion of lox-flanked constructs [1] |
| β-estradiol inducible Cre (CreEBD) | Controlled recombinase expression | Enables temporal control of recombination [35] |
| Reporter plasmid libraries | Evaluation of lox variant efficiency | Enables high-throughput screening [35] |
The research toolkit for advanced chromosome engineering has expanded significantly with the development of these new technologies. The core components include the engineered Cre variants for enhanced efficiency, novel low-reversion Lox sites for stability, and complementary editing systems that enable scarless modifications [4]. The Bxb1 recombinase system has proven particularly valuable for efficiently integrating large lox-flanked constructs into specific genomic loci, enabling systematic studies of how inter-lox distance affects recombination efficiency [1]. Inducible Cre systems remain essential for controlling the timing of recombination events, while comprehensive reporter libraries facilitate the high-throughput characterization of new lox variants [35].
When applying these tools, researchers should consider several technical factors that influence experimental outcomes. Studies have demonstrated that inter-lox distance significantly impacts recombination efficiency, with optimal results achieved when lox sites are spaced 1-4 kb apart for wild-type sites or 1-3 kb for mutant sites [1] [3]. Additionally, the choice of Cre-driver strain, zygosity of floxed alleles, and age of Cre-driver animals all contribute to variability in recombination efficiency and should be carefully controlled in experimental design [1].
The development of low-reversion lox variants represents a transformative advancement in genome engineering technology, effectively addressing the fundamental reversibility problem that has limited traditional Cre-Lox systems for decades. Through systematic recombination site engineering, AI-assisted protein design, and innovative scarless editing strategies, researchers have created a new generation of tools that enable efficient, stable, and precise chromosomal modifications across kilobase to megabase scales [4] [6].
These technological breakthroughs have profound implications for diverse research applications. In basic research, they enable more predictable and reproducible genetic manipulations, reducing experimental variability and enhancing data reliability. For agricultural biotechnology, the ability to create stable, large-scale chromosomal modifications opens new possibilities for crop improvement, as demonstrated by the herbicide-resistant rice model [4] [7]. In therapeutic development, the improved precision and stability of these systems may eventually support novel gene therapy approaches targeting structural variations implicated in genetic disorders.
While the PCE and RePCE systems currently represent the state-of-the-art in chromosome engineering, further refinements are undoubtedly forthcoming. The integration of these technologies with other emerging modalities—including advanced delivery systems and computational prediction models—will continue to expand the boundaries of achievable genetic modifications. As these tools become more widely adopted and validated across different biological systems, they are poised to replace traditional Cre-Lox systems as the preferred platform for complex genome engineering applications, ultimately accelerating progress across basic research, agricultural science, and therapeutic development [7].
For decades, the Cre-Lox system has been a cornerstone of genetic engineering, enabling precise chromosomal manipulations through its site-specific recombination mechanism. However, researchers working with this system have consistently faced a critical bottleneck: the limited recombination efficiency of the wild-type Cre recombinase, which becomes particularly problematic when manipulating large DNA segments. This efficiency constraint has hindered applications ranging from complex genetic modeling to therapeutic development. A breakthrough has now been achieved with the development of Programmable Chromosome Engineering (PCE) systems, which directly address this limitation through AI-driven protein engineering. The centerpiece of this advancement is an engineered recombinase that demonstrates 3.5 times the recombination efficiency of wild-type Cre, a quantitative leap that fundamentally transforms what's possible in chromosome engineering [6] [36] [5].
Historical Limitations of the Cre-Lox System
The traditional Cre-Lox system's inefficiency stems from several fundamental characteristics of its molecular components. The tetrameric nature of the Cre recombinase enzyme creates inherent complexity for engineering efforts, as modifications to one subunit can disrupt the delicate quaternary structure essential for function [6] [36]. Additionally, the symmetrical design of conventional Lox sites leads to reversible recombination reactions, where a significant portion of enzymatic activity is wasted on undoing and redoing edits rather than driving them to completion [4] [5]. This reversibility problem means that even when Cre functions optimally, a substantial portion of its catalytic activity fails to produce stable genetic modifications. Beyond these core issues, the system historically left behind residual Lox sites after recombination, creating permanent "scars" in the genome that could potentially interfere with normal gene function and cellular processes [36] [26].
The PCE Breakthrough: A Multi-Faceted Engineering Approach
The development of PCE systems addressed these limitations through three interconnected innovations, with the enhanced recombinase standing as the centerpiece achievement.
Core Innovation 1: AiCErec - AI-Driven Recombinase Engineering
The most significant advancement in PCE technology is the AiCErec (AI-informed Constraints for protein Engineering for recombinases) platform, which systematically optimized Cre's structure-function relationship [6] [36] [26].
Experimental Methodology: The researchers employed a protein-directed evolution system that integrated general inverse folding models with structural and evolutionary constraints [36] [5]. This approach enabled precise optimization of Cre's multimerization interface - the critical region where individual Cre subunits assemble into the functional tetrameric complex [6] [4]. Rather than random mutagenesis, the AI platform strategically identified mutations that would stabilize subunit interactions while enhancing DNA-binding affinity and catalytic activity. The engineered variants were then screened using high-throughput assays to quantify recombination efficiency against standard genetic targets [4].
Quantitative Outcome: This targeted engineering produced a Cre variant with recombination efficiency 3.5 times greater than wild-type Cre [6] [36] [26]. This enhancement directly translates to higher success rates for all types of chromosomal manipulations, from small insertions to megabase-scale rearrangements.
Supporting Innovation 2: Asymmetric Lox Site Design
Complementing the recombinase engineering, the team developed novel asymmetric Lox variants that fundamentally changed the reaction dynamics [6] [4].
Experimental Methodology: Using a high-throughput platform for rapid recombination site modification, researchers systematically designed and tested Lox sequences with asymmetric architectures that favored forward recombination [36] [5]. This platform enabled rapid screening of countless Lox variants, identifying those with optimal binding characteristics for the engineered recombinase while minimizing backward reactions.
Quantitative Outcome: The optimized Lox variants demonstrated a reduction in reversible recombination activity by over 10-fold, bringing it nearly to background levels observed in negative controls while maintaining high-efficiency forward recombination [6] [36] [26].
Supporting Innovation 3: Scarless Editing Strategy
To address the problem of residual recombination sites, the team developed Re-pegRNA, a method that leverages prime editing technology [6] [36].
Experimental Methodology: This approach uses specifically designed prime-editing guide RNAs (Re-pegRNAs) to perform "re-prime editing" on residual Lox sites after the primary recombination event [36] [5]. The system precisely replaces the leftover Lox sequences with the original genomic sequence, effectively erasing all traces of the engineering process.
Quantitative Outcome: The result is truly scarless genome editing, with the edited genomic sequence indistinguishable from native sequence architecture while achieving editing efficiencies up to 26.2% for complex manipulations [4] [21].
Comparative Performance Analysis
The quantitative advantages of the PCE system become evident when comparing its performance metrics against traditional Cre-Lox systems across various manipulation types.
Table 1: Comparison of Editing Capabilities Between Traditional Cre-Lox and PCE Systems
| Editing Type | Traditional Cre-Lox Limitations | PCE System Performance | Efficiency Gain |
|---|---|---|---|
| DNA Insertion | Limited efficiency for large fragments | Targeted integration up to 18.8 kb | ~3.5× with novel recombinase [36] [5] |
| DNA Replacement | Challenging beyond small segments | Complete replacement of 5-kb sequences | Enhanced by asymmetric Lox sites [36] [5] |
| Chromosomal Inversion | Limited scale and efficiency | Inversions spanning 12 Mb in human cells, 315 kb in rice | Enabled by reduced reversibility [36] [21] |
| Chromosomal Deletion | Size-constrained | Deletions of 4 Mb achieved | Scarless editing prevents residual sequences [36] [5] |
| Translocation | Technically challenging | Whole-chromosome translocations demonstrated | High-efficiency recombination enables complex rearrangements [6] [36] |
Table 2: Direct Comparison of Cre vs. Engineered PCE Recombinase Properties
| Property | Wild-Type Cre Recombinase | PCE Engineered Recombinase |
|---|---|---|
| Recombination Efficiency | Baseline (1×) | 3.5× higher than wild-type [6] [36] [26] |
| Reversibility | High (symmetrical Lox sites) | >10× reduction with asymmetric Lox sites [6] [36] [26] |
| Editing Precision | Leaves residual Lox "scars" | Scarless via Re-pegRNA system [6] [36] [5] |
| Target Programmability | Limited | Flexible programming of insertion positions/orientations [6] [36] |
Visualization of the PCE Engineering Strategy
The following diagram illustrates the multi-faceted engineering approach that enabled the breakthrough in recombinase activity:
Molecular Mechanism of Enhanced Recombination
The structural basis for the 3.5-fold efficiency improvement lies in specific modifications to the recombinase's functional architecture:
Essential Research Reagents for PCE Implementation
Table 3: Key Research Reagents for PCE System Implementation
| Reagent/Cell Line | Function in PCE Workflow | Experimental Application |
|---|---|---|
| Engineered Cre Variant | Catalyzes high-efficiency recombination | Core component for all DNA manipulations; 3.5× more efficient than wild-type [6] [36] [26] |
| Asymmetric Lox Variants | Target sites for recombination | Enable irreversible editing; reduce reversibility >10× [6] [36] [26] |
| Re-pegRNA Constructs | Removes residual Lox sites | Creates scarless edits; used in final editing phase [6] [36] [5] |
| Prime Editor Components | Facilitates precise sequence replacement | Required for Re-pegRNA scarless editing step [36] [5] |
| Plant/Animal Cell Lines | Host organisms for editing validation | Rice for plant models; human cell lines for therapeutic research [6] [36] [21] |
The development of PCE systems with their engineered recombinase represents a paradigm shift in large-scale genome manipulation. By achieving a 3.5-fold enhancement in recombination efficiency through AI-driven protein engineering, while simultaneously addressing the historical limitations of reversibility and scarring, this technology has effectively overcome the critical bottlenecks that constrained traditional Cre-Lox systems. The quantitative advances in editing scale - from kilobase to megabase manipulations - open new frontiers in functional genomics, agricultural biotechnology, and therapeutic development. As these tools become widely adopted, researchers can now approach chromosomal-scale engineering with unprecedented precision and efficiency, accelerating the pace of discovery across the life sciences.
The ability to precisely manipulate large segments of DNA represents a frontier in genetic engineering, with profound implications for therapeutic development, agricultural improvement, and basic biological research. For decades, the Cre-Lox system has served as a cornerstone technology for these endeavors, enabling researchers to execute deletions, inversions, and integrations of genetic material. However, a persistent limitation has hampered its broader application: the "scar issue." Following recombination, all conventional Cre-Lox reactions leave behind residual loxP sites—non-native 34-base pair sequences that remain embedded in the genome at the editing site [4] [10]. These scars are not merely aesthetic imperfections; they can potentially disrupt native gene function, alter regulatory elements, and serve as unintended substrates for subsequent recombination events, leading to genomic instability and unpredictable phenotypic outcomes [4].
The quest for scarless editing has driven the field toward innovation. This guide provides an objective comparison between the traditional Cre-Lox system and a new generation of Programmable Chromosome Engineering (PCE) systems, focusing on their performance regarding editing precision, efficiency, and the critical issue of residual sequences. We frame this comparison within a broader thesis that while Cre-Lox technology laid the essential groundwork for targeted genomic rearrangements, newly developed PCE systems represent a transformative advance by achieving seamless, scar-free modifications across kilobase to megabase scales [4] [21] [6].
The Established Standard: Cre-Lox Recombinase System
The Cre-Lox system, derived from bacteriophage P1, consists of Cre recombinase and its specific 34 bp DNA target sites, loxP [10]. The technology's versatility is demonstrated in the table below, which summarizes key recombination outcomes based on the orientation and location of loxP sites.
Table 1: Outcomes of Cre-Lox Recombination Based on loxP Site Configuration
| loxP Site Arrangement | Orientation | Primary Recombination Outcome |
|---|---|---|
| cis (on same DNA strand) | Same direction | Excision/Deletion of flanked sequence |
| cis (on same DNA strand) | Opposite directions | Inversion of flanked sequence |
| trans (on different DNA strands) | Same direction | Translocation of DNA segments |
A fundamental and inherent limitation of the wild-type system is its reversibility. Because the loxP sites are symmetrical and the reaction is bidirectional, the excised DNA fragment can readily re-integrate, preventing stable, permanent edits [4]. Furthermore, after an excision event, a single residual loxP "scar" remains in the genome. While this site is only 34 bp, its presence in a coding or regulatory region can have unintended consequences, and it presents a substrate for potentially deleterious future recombination events in the presence of Cre recombinase [4] [9].
The New Paradigm: Programmable Chromosome Engineering (PCE) Systems
To address the core limitations of Cre-Lox, researchers have developed a suite of technologies termed Programmable Chromosome Engineering (PCE). These systems are not merely incremental improvements but a fundamental re-engineering of the editing machinery [4] [6]. Two primary systems, PCE and RePCE, have been developed, integrating three key innovations:
- Engineered Recombination Sites: Through high-throughput screening, novel Lox variants were identified that exhibit 10-fold lower reversibility while maintaining high forward recombination efficiency, ensuring edits are stable and permanent [4] [21].
- AI-Optimized Recombinase (AiCErec): An AI-assisted protein engineering approach was used to optimize the Cre multimerization interface, resulting in Cre variants with 3.5 times the recombination efficiency of the wild-type enzyme [4] [6].
- Scar-Free Editing Strategy (RePCE): This strategy harnesses prime editing. Specifically designed Re-pegRNAs are used to perform "re-prime editing" on residual Lox sites after the primary recombination event, precisely replacing them with the original genomic sequence. This leaves no foreign DNA behind, achieving true scarlessness [4].
The following workflow diagram illustrates how these components integrate to overcome the traditional limitations of the Cre-Lox system.
Performance Comparison: Quantitative Data Analysis
The theoretical advantages of PCE systems are substantiated by robust experimental data. The table below provides a direct, objective comparison of key performance metrics between the traditional Cre-Lox system and the new PCE systems, based on results reported in plant and human cells [4] [21].
Table 2: Direct Performance Comparison: Traditional Cre-Lox vs. PCE Systems
| Editing Parameter | Traditional Cre-Lox | New PCE/RePCE Systems |
|---|---|---|
| Recombination Efficiency | Baseline (Wild-type) | 3.5x increase (via AiCErec-engineered variant) |
| Reaction Reversibility | High (inherently symmetrical) | 10-fold reduction (via novel Lox variants) |
| Editing Precision | Leaves residual loxP "scars" | Scarless (via Re-pegRNA strategy) |
| Maximum Insertion Size | Limited, inefficient for large segments | Up to 18.8 kb |
| Maximum Deletion Size | Limited, highly mosaic over large distances | Up to 4 Mb |
| Maximum Inversion Size | Inefficient over long ranges | Up to 12 Mb (in human cells) |
| Editing Efficiency | Variable, often low for large edits | Up to 26.2% (for various modification types) |
The capability of PCE systems to execute massive, precise edits is a game-changer. For context, a systematic analysis of the traditional Cre-Lox system in mice found that optimal recombination requires loxP sites to be spaced less than 4 kb apart for wild-type sites and less than 3 kb for mutant lox71/66 sites. Complete recombination consistently failed with wild-type loxP sites spaced ≥ 15 kb apart [1]. In stark contrast, the PCE system has demonstrated the ability to create a precise 315-kb inversion in rice to confer herbicide resistance—a feat that was previously prohibitively difficult [4] [6]. This directly translates to practical benefits in breeding and research, reducing the number of seeds that need to be genetically edited to find one with a desired trait from roughly 1,000 to 100, saving significant time and resources [7].
Experimental Protocols for Key Applications
Protocol: Scarless Large DNA Insertion Using RePCE
The following detailed methodology outlines the workflow for achieving scarless insertion of large DNA fragments, a core application of the RePCE system [4].
- Targeting Vector Design and Construction: Clone the large DNA fragment (up to 18.8 kb) to be inserted into a donor plasmid. This fragment must be flanked by the novel, engineered Lox sites that exhibit low reversibility.
- Re-pegRNA Design and Synthesis: Design a prime editing guide RNA (Re-pegRNA) that specifically targets the genomic region where the residual Lox site will remain post-recombination. The Re-pegRNA's template must encode the original, native genomic sequence to facilitate perfect restoration.
- Co-delivery into Cells: Co-transfect the target cells (plant or human) with:
- The donor plasmid containing the floxed insert.
- A plasmid expressing the AiCErec-optimized Cre recombinase variant.
- The CRISPR-based prime editing system (PE protein and the synthesized Re-pegRNA).
- Screening and Validation: Isolate and expand transfected cells. Screen for successful integration using PCR across the 5' and 3' junctions. Confirm the absence of residual Lox sites via Sanger sequencing of the edited junctions, verifying seamless and scarless integration.
Protocol: Programmable Chromosome Inversion
This protocol describes the steps for creating large-scale chromosomal inversions, useful for modeling structural variants and studying gene regulation [4].
- Dual Lox Site Insertion: Using prime editors, precisely integrate two engineered Lox sites in an inverted orientation at the desired boundaries of the genomic segment to be inverted. The distance between these sites can range from kilobases to megabases.
- Induction of Recombinase Expression: Introduce the expression construct for the high-efficiency AiCErec Cre variant into the cells. This can be done via transient transfection or through a stable, inducible expression system.
- Recombination and Inversion: The expressed Cre variant will recognize the paired Lox sites and catalyze the inversion of the intervening DNA segment. The use of low-reversibility Lox sites ensures the inversion is stable and not readily reverted.
- Karyotype and Molecular Analysis: Validate the inversion using a combination of long-range PCR, Southern blotting, and for very large inversions (e.g., the 12 Mb inversion in human cells), karyotyping or optical mapping.
The Scientist's Toolkit: Essential Research Reagent Solutions
The implementation of these advanced chromosome engineering technologies requires a specific set of molecular tools and reagents. The table below details key components essential for researchers looking to adopt PCE systems.
Table 3: Key Research Reagents for Programmable Chromosome Engineering
| Reagent / Tool | Type/Description | Primary Function in PCE/RePCE |
|---|---|---|
| AiCErec-engineered Cre | Protein / DNA plasmid | High-efficiency catalyzes recombination; 3.5x more efficient than wild-type [4] [21]. |
| Engineered Lox Sites | DNA sequence (e.g., novel Lox variants) | Target for Cre recombinase; provides 10-fold lower reversibility for stable edits [4]. |
| Re-pegRNA | Synthetic guide RNA | Directs prime editor to remove residual Lox sites, enabling scarless genome modifications [4] [6]. |
| Prime Editor (PE) | Protein / RNA complex (PE protein + pegRNA) | Executes the "search-and-replace" editing to seamlessly restore original sequence post-recombination [4]. |
| Bxb1 Recombinase System | Integrase / attB & attP sites | Facilitates high-efficiency, site-specific integration of large transgene constructs for building complex alleles, useful for preparing floxed target sites [1]. |
The objective comparison presented in this guide clearly demonstrates a paradigm shift in large-scale genome engineering. The traditional Cre-Lox system, while foundational, is intrinsically limited by inefficiency, reversibility, and the unavoidable problem of genomic scars. The emergence of Programmable Chromosome Engineering (PCE and RePCE) systems marks a significant leap forward. By integrating AI-assisted protein engineering, high-throughput site optimization, and scar-free editing strategies, these technologies overcome historical bottlenecks [4] [6].
The quantitative data speaks unequivocally: PCE systems offer dramatically enhanced efficiency, the ability to manipulate DNA across megabase scales, and, most critically, the capacity for seamless, scarless modifications. This precision is paramount for applications in therapeutic development, where any residual foreign DNA could pose safety risks, and in advanced crop breeding, where the goal is to create clean, market-acceptable products with precise trait improvements [4] [7]. As these tools are adopted and refined, they are poised to accelerate progress in synthetic biology, disease modeling, and the development of next-generation cell and gene therapies, effectively rewriting the code of what is possible in genetic manipulation.
The ability to manipulate DNA at scale represents a critical frontier in genetic research, with profound implications for therapeutic development, agricultural biotechnology, and fundamental biological discovery. For decades, the Cre-Lox system has served as a cornerstone technology for conditional genetics and targeted genomic modifications. However, its utility has been constrained by significant scale limitations, particularly when manipulating DNA sequences beyond a few kilobases. Recent breakthroughs have yielded a new class of technologies—Programmable Chromosome Engineering (PCE) systems—that dramatically expand the scope of possible edits from kilobases to megabases. This comparison guide examines the technical capabilities, experimental performance, and practical applications of both established Cre-Lox systems and emerging PCE platforms, providing researchers with a comprehensive framework for selecting appropriate tools for large-scale genetic engineering projects.
The fundamental challenge in large-scale genome editing lies in achieving precise, efficient manipulations without introducing unintended genomic damage or leaving behind residual "scar" sequences that can compromise function or regulation. While CRISPR-Cas systems have revolutionized point mutations and small insertions/deletions, they face limitations in efficiency and precision when dealing with larger modifications. Site-specific recombinase systems like Cre-Lox offer an alternative pathway but have historically been hampered by reversibility reactions, efficiency limitations, and residual recombination sites that remain in the edited genome. The new PCE systems address these limitations through integrated engineering approaches that combine computational protein design, high-throughput screening, and precision editing technologies.
Fundamental Mechanisms and Applications
The Cre-Lox system, derived from bacteriophage P1, utilizes the Cre recombinase enzyme to catalyze recombination between specific 34-base pair DNA sequences known as Lox sites. The orientation and relative position of these Lox sites determine the outcome of recombination: excision when sites are directly repeated, inversion when sites are inverted relative to each other, and translocation when sites are on different DNA molecules [10]. This system has become indispensable in mouse genetics, enabling spatiotemporal control of gene expression through conditional knockout models where critical gene segments are flanked by Lox sites ("floxed") and removed upon Cre-mediated recombination [1].
The Cre-Lox system functions without generating DNA double-strand breaks, instead relying on a Holliday junction intermediate during strand exchange [37]. This mechanism avoids the toxicity associated with double-strand break repair and makes the technology particularly valuable in systems with inefficient native repair pathways. Beyond basic gene deletion, advanced applications include the Brainbow system for multicolor neuronal labeling, inducible systems using Cre-ERT fusions activated by tamoxifen, and intersectional genetics combining multiple recombinase systems for enhanced precision [1] [10].
Documented Scale Limitations
Despite its versatility, the Cre-Lox system faces well-documented constraints when targeting large genomic regions. Systematic analyses in mouse models reveal that recombination efficiency declines significantly as the distance between Lox sites increases. Optimal recombination occurs with inter-Lox distances of less than 4 kb for standard LoxP sites and 3 kb for mutant Lox variants, with complete failure of recombination observed at distances ≥15 kb for wildtype LoxP sites and ≥7 kb for mutant Lox71/66 sites [1].
Additional factors compounding these scale limitations include:
- Reversible recombination: The inherent symmetry of LoxP sites creates thermodynamic equilibrium where forward and reverse reactions compete, limiting the completeness of desired edits [6].
- Mosaicism: In vivo applications frequently produce mosaic animals with incomplete recombination, even when inter-Lox distances fall within the "efficient" range [1].
- Genomic context dependence: Recombination efficiency varies significantly across different chromosomal locations, with some loci proving particularly recalcitrant to editing [1].
- Residual "scar" sequences: Conventional Cre-Lox editing leaves behind Lox sites after recombination, potentially interfering with normal gene function and regulation [6].
Table 1: Factors Influencing Cre-Lox Recombination Efficiency
| Factor | Impact on Efficiency | Optimal Range |
|---|---|---|
| Inter-Lox Distance | Efficiency declines with increasing distance | <4 kb (standard LoxP), <3 kb (mutant Lox) |
| Cre Driver Strain | Significant variability between strains | Ella-cre shows broad efficiency |
| Genomic Context | Locus-dependent effects | Consistent at fixed loci (e.g., Rosa26) |
| Zygosity | Heterozygous floxed alleles recombine more efficiently | Heterozygous preferred over homozygous |
| Animal Age | Age-dependent efficiency changes | 8-20 weeks optimal for breeders |
The PCE Breakthrough: Engineering at Scale
A research team led by Prof. GAO Caixia at the Chinese Academy of Sciences has recently developed Programmable Chromosome Engineering (PCE and RePCE) systems that overcome the fundamental limitations of traditional Cre-Lox technology [6] [21] [22]. This integrated platform combines three key innovations to enable precise, scarless manipulations at unprecedented scales:
Asymmetric Lox variants: Through high-throughput screening, the team developed novel Lox site variants that reduce reversible recombination by over 10-fold while maintaining high forward recombination efficiency [6] [22].
AI-informed protein engineering (AiCErec): Leveraging a protein-directed evolution system integrating general inverse folding models with structural constraints, the researchers created Cre variants with 3.5 times the recombination efficiency of wild-type Cre [6] [21].
Scarless editing strategy (Re-pegRNA): By combining prime editing with specifically designed pegRNAs, the system precisely replaces residual Lox sites with the original genomic sequence, ensuring truly seamless modifications [6] [21].
These innovations collectively enable flexible programming of insertion positions and orientations for different Lox sites, allowing precise manipulation of DNA fragments ranging from kilobase to megabase scales in both plant and animal cells [21].
Demonstrated Editing Capabilities
The PCE systems have demonstrated unprecedented editing capabilities across multiple systems:
Table 2: Documented Editing Capabilities of PCE Systems
| Edit Type | Scale Demonstrated | Experimental System |
|---|---|---|
| Targeted Insertion | Up to 18.8 kb | Plant and animal cells |
| Sequence Replacement | Complete 5-kb replacement | Plant and animal cells |
| Chromosomal Inversion | 315 kb to 12 Mb | Rice (315 kb), human cells (12 Mb) |
| Chromosomal Deletion | Up to 4 Mb | Plant and animal cells |
| Whole-Chromosome Translocation | Full chromosome scale | Plant and animal cells |
As a proof of concept, the researchers created herbicide-resistant rice by introducing a precise 315-kb inversion in the genome, demonstrating the technology's potential for crop improvement [6] [22]. In human cells, the system achieved a 12-Mb inversion at disease-relevant loci, highlighting its therapeutic potential [21].
Direct Performance Comparison
Quantitative Metrics and Experimental Validation
Direct comparison of key performance metrics reveals the dramatic advances represented by PCE systems:
Table 3: Performance Comparison: Cre-Lox vs. PCE Systems
| Parameter | Traditional Cre-Lox | PCE Systems |
|---|---|---|
| Maximum Efficient Deletion | 4 kb | 4 Mb |
| Maximum Efficient Insertion | 1-3 kb (via HDR) | 18.8 kb |
| Recombination Efficiency | Distance-dependent decline | Maintained across scales |
| Editing Precision | Leaves residual Lox sites | Scarless (no residual sequences) |
| Reversibility | Significant reversible reactions | >10-fold reduction |
| Chromosomal Rearrangements | Limited to small inversions | Up to megabase inversions/translocations |
The PCE systems achieve these advances through their engineered components. The AiCErec-evolved Cre recombinase demonstrates 3.5-fold higher activity than wild-type Cre, directly addressing the efficiency limitations that constrained traditional applications [6]. The asymmetric Lox variants fundamentally alter the reaction equilibrium, favoring forward reactions and thereby ensuring more complete editing [6] [21].
Experimental Workflows and Methodologies
The experimental workflow for PCE systems builds upon but significantly enhances traditional recombinase approaches:
PCE System Experimental Workflow
The key methodological differentiators in PCE workflows include:
Asymmetric Lox site deployment: Novel Lox variants with reduced symmetry are integrated to minimize reversible reactions.
High-efficiency Cre delivery: The AiCErec-engineered Cre variants are delivered using optimized expression systems.
Dual-stage editing: Primary recombination is followed by Re-pegRNA-mediated scar removal to eliminate residual Lox sites.
Multiplexed validation: Edited lines undergo comprehensive genotyping and phenotyping to confirm intended modifications.
This workflow contrasts with traditional Cre-Lox approaches, which typically involve symmetric Lox sites, wild-type Cre, and no scar-removal step, resulting in the documented limitations.
Research Reagent Solutions
Successful implementation of these technologies requires specific reagent systems and tools:
Table 4: Essential Research Reagents for Chromosome Engineering
| Reagent Category | Specific Examples | Function & Application |
|---|---|---|
| Recombinase Enzymes | Wild-type Cre, AiCErec-engineered Cre variants, FLPo, Dre | Catalyze site-specific recombination with varying efficiencies and specificities |
| Recognition Sites | LoxP, asymmetric Lox variants, LoxPsym, frt, rox | Serve as recombination targets with different orthogonality and efficiency profiles |
| Delivery Vectors | AAV, lentiviral, plasmid-based expression systems | Enable efficient delivery of editing components to target cells |
| Selection Systems | Antibiotic resistance, fluorescent reporters, metabolic markers | Facilitate identification and isolation of successfully edited cells |
| Validation Tools | PCR screening primers, Southern blot protocols, sequencing assays | Confirm intended edits and assess off-target effects |
For orthogonal applications requiring multiple independent recombination events, researchers can leverage recently developed LoxPsym variants. One study established 16 orthogonal LoxPsym sites that enable multiplexed genome engineering without cross-reactivity in prokaryotic and eukaryotic hosts [37]. These symmetrical Lox variants expand the Cre-Lox toolbox for complex applications including metabolic engineering and genetic circuit construction.
For spatially and temporally controlled activation, optogenetic Cre systems such as REDMAPCre provide enhanced precision. This recently developed red-light-controlled split-Cre system enables rapid activation (1-second illumination) with an 85-fold increase in reporter expression over background, facilitating precise genomic manipulation in deep tissues without invasive fiber implantation [13].
Applications and Implications for Research
Therapeutic Development
The ability to perform megabase-scale chromosomal engineering opens new avenues for modeling human diseases caused by large structural variations, including genomic disorders such as DiGeorge syndrome (3 Mb deletion) and Smith-Magenis syndrome (4 Mb deletion). Where previous technologies could only approximate these conditions through partial gene knockouts, PCE systems enable precise recapitulation of the actual genetic alterations, creating more clinically relevant models for therapeutic screening and validation.
The documented 12-Mb inversion in human cells at disease-relevant loci demonstrates the potential for creating accurate models of chromosomal rearrangement disorders [21]. Furthermore, the scarless nature of RePCE editing ensures that therapeutic constructs can be integrated without residual exogenous sequences that might trigger immune responses or regulatory complications in clinical applications.
Agricultural Biotechnology
The development of herbicide-resistant rice through a precise 315-kb chromosomal inversion showcases the transformative potential of PCE systems for crop improvement [6] [22]. Traditional transgenic approaches often face regulatory hurdles and public skepticism due to the incorporation of foreign DNA sequences. PCE technology enables the creation of improved crop varieties through precise chromosomal rearrangements without introducing exogenous DNA, potentially streamlining regulatory approval and public acceptance.
The scalability of PCE systems makes them particularly valuable for manipulating complex traits controlled by large genomic regions, such as yield components, stress resistance, and nutritional quality. The ability to perform megabase-scale manipulations in plants positions this technology as a powerful tool for addressing food security challenges in the face of climate change.
Synthetic Biology
The expanded scale of precise DNA manipulation enables more ambitious synthetic biology projects, including whole-pathway integration, chromosome synthesis, and genome-scale reorganization. The orthogonal LoxPsym system [37] complements PCE technologies by enabling multiple independent recombination events in a single organism, essential for complex genome engineering projects.
These capabilities support the construction of increasingly sophisticated genetic circuits and the development of microbial chassis strains optimized for industrial production of biofuels, pharmaceuticals, and specialty chemicals. The avoidance of double-strand breaks during editing makes these technologies particularly valuable for applications requiring high viability and genome stability.
The emergence of PCE systems represents a paradigm shift in large-scale genome engineering, effectively addressing the fundamental limitations that have constrained traditional Cre-Lox technology. The quantitative data demonstrate unambiguous advantages across all measured parameters—efficiency, scale, precision, and versatility.
For research applications requiring manipulations below 4 kb, particularly in established model systems with well-characterized Cre drivers, traditional Cre-Lox approaches remain sufficient and may offer practical advantages in terms of accessibility and established protocols. However, for projects involving larger genomic regions, requiring complete removal of exogenous sequences, or aiming for chromosomal-scale rearrangements, PCE systems provide unequivocal technical superiority.
The integration of AI-informed protein engineering, asymmetric recombination sites, and scarless editing strategies in PCE platforms establishes a new standard for genome engineering capabilities. As these technologies become more widely adopted and optimized, they are poised to accelerate progress across biomedical research, therapeutic development, and agricultural biotechnology by enabling precise manipulation of the genome at its natural structural scales.
PCE vs. Cre-Lox: A Data-Driven Comparison for Modern Research
For researchers in chromosome engineering, selecting the right gene-editing tool hinges on precise performance data. This guide provides a quantitative, evidence-based comparison between the established Cre-Lox system and the novel Programmable Chromosome Engineering (PCE) systems, focusing on editing efficiency and scale.
Quantitative Performance Comparison: Cre-Lox vs. PCE Systems
The table below summarizes key performance metrics from recent studies, offering a direct comparison of capabilities.
| Performance Metric | Traditional Cre-Lox System | Novel PCE Systems |
|---|---|---|
| Optimal Deletion Efficiency | Near-complete recombination with inter-loxP distance < 4 kb [1] | Not explicitly quantified for small scales, but enables large-scale edits previously impossible [26] [17]. |
| Maximum Demonstrated Deletion | ~4 Mb in mouse ES cells (deletion efficiency decreases with distance) [24] | 4 Mb chromosomal deletion [26] [17] [6] |
| Maximum Demonstrated Integration | Typically limited to a few kb for efficient integration | Targeted integration of DNA fragments up to 18.8 kb [26] [17] |
| Maximum Demonstrated Inversion | Not specified for large scales in provided results | Chromosomal inversion spanning 12 Mb [26] [17] [6] |
| Key Limitation | Reversible reaction; residual loxP "scars"; efficiency drops over distance [26] [1] [17] | Engineered to be scarless and have reduced reverse reaction [26] [17] [6] |
| Protein Engineering | Wild-type Cre recombinase | AI-engineered Cre variant with 3.5x higher recombination efficiency than wild-type [26] [17] [6] |
Experimental Protocols and Methodologies
Understanding the experimental designs that yielded these metrics is crucial for evaluating the data.
Systematic Optimization of Cre-Lox in Mice
A 2025 study systematically analyzed factors affecting Cre-Lox recombination efficiency to establish a robust optimization framework [1].
- Strain Generation: Researchers generated 11 novel conditional allele strains on a C57BL/6J background, targeting the Rosa26 locus. A high-efficiency Bxb1 recombinase system was used to integrate constructs with varying inter-loxP distances (from 0.8 kb to 15 kb), including both wild-type and mutant loxP sites [1].
- Efficiency Testing: Female mice from three common Cre-driver strains (Ella-cre, CMV-cre, Sox2-cre) were bred with male mice carrying the "floxed" alleles. The resulting F1 offspring were genotyped to determine the percentage showing complete recombination, mosaicism, or no recombination. This design allowed for the direct comparison of efficiency across different drivers and genetic distances [1].
- Key Parameters Measured: The study quantified the impact of inter-loxP distance, Cre-driver strain, zygosity of the floxed allele, age of the breeder, and the type of loxP site (wild-type vs. mutant) on recombination efficiency and mosaicism [1].
Development and Validation of PCE Systems
Professor GAO Caixia's team addressed the fundamental limitations of Cre-Lox through three key innovations, validated in plant and animal cells [26] [17] [6].
- Asymmetric lox Site Design: The team developed a high-throughput platform to modify recombination sites. They engineered novel, asymmetric lox variants that reduced the undesirable reverse recombination reaction by over 10-fold while maintaining high forward recombination efficiency [26] [17].
- AI-Driven Protein Engineering: The researchers employed an AI-informed protein engineering (AiCE) model to create AiCErec, a method for optimizing recombinases. This led to an engineered Cre variant with a recombination efficiency 3.5 times greater than the wild-type enzyme [26] [17] [6].
- Scarless Editing Strategy (Re-pegRNA): To remove residual lox sites after editing, the team harnessed prime editing technology. They designed specialized pegRNAs (Re-pegRNA) to perform "re-prime editing," precisely replacing the lox site with the original genomic sequence, ensuring seamless modifications [26] [17].
The integration of these methods created the two core platforms, PCE and RePCE, which were then tested for large-scale edits. The workflow for creating herbicide-resistant rice involved introducing a precise 315-kb inversion in the rice genome using these technologies [17] [6].
Logical Workflow of PCE Technology Development
The following diagram illustrates the logical progression of innovations that led to the advanced capabilities of PCE systems.
The Scientist's Toolkit: Key Research Reagent Solutions
This table details essential reagents and their functions for working with Cre-Lox and PCE technologies, as featured in the cited research.
| Research Reagent / Tool | Function in Experimentation |
|---|---|
| Cre-driver Strains (e.g., Ella-cre, CMV-cre) [1] | Genetically engineered mouse lines that express Cre recombinase in specific tissues or cell types, enabling spatial control of gene editing. |
| Floxed Allele Strains [16] [1] | Mouse strains where a target DNA sequence is flanked by two loxP sites. Crossing them with Cre-driver strains triggers recombination at the target locus. |
| Asymmetric lox Variants [26] [17] | Engineered recombination sites that minimize reverse reaction rates, stabilizing edits and increasing forward recombination efficiency for advanced systems like PCE. |
| Engineered Cre Recombinase (AiCErec) [26] [17] [6] | An AI-optimized variant of the Cre enzyme with significantly higher (3.5x) recombination efficiency than the wild-type protein. |
| Re-pegRNA [26] [17] | A specially designed prime editing guide RNA used to remove residual recombination sites after editing, enabling seamless "scarless" genome modifications. |
| Bxb1 Recombinase System [1] | An alternative recombinase system used for high-efficiency integration of large constructs (e.g., floxed reporters) into specific genomic loci, facilitating strain generation. |
| Orthogonal LoxPsym Sites [37] | A set of 16 non-cross-reacting, symmetrical lox variants that enable multiple, independent recombination events within a single organism without interference. |
For decades, the Cre-Lox system has served as a cornerstone tool for precise genome engineering in both plant and mammalian cells. This technology, derived from bacteriophage P1, enables researchers to perform targeted DNA manipulations including deletions, insertions, inversions, and translocations. The system operates through the Cre recombinase enzyme which recognizes and catalyzes recombination between specific 34-base pair DNA sequences known as Lox sites. The orientation and position of these Lox sites determine the specific genetic outcome—inversion when sites are in opposite orientations, deletion when in the same orientation, and translocation when on different DNA molecules. Despite its widespread adoption and considerable utility across molecular biology, synthetic biology, and therapeutic development, the Cre-Lox system has been hampered by three persistent limitations: reversible recombination reactions stemming from symmetrical Lox sites that can undo desired edits, the tetrameric nature of Cre recombinase that complicates protein engineering efforts, and residual Lox sites remaining after recombination that compromise editing precision by leaving behind unwanted "scar" sequences.
The recent development of Programmable Chromosome Engineering (PCE) systems by Gao Caixia's team at the Chinese Academy of Sciences represents a paradigm shift in large-scale DNA manipulation. These technologies specifically address the historical limitations of Cre-Lox while dramatically expanding the scale and precision of possible genetic edits. This analysis comprehensively compares the editing fidelity, efficiency, and scalability of traditional Cre-Lox systems against emerging PCE technologies, providing researchers with experimental data and methodological insights to inform their genome engineering strategies.
Technological Foundations: From Cre-Lox to Programmable Chromosome Engineering
The Cre-Lox System: Established Workhorse with Defined Limitations
The Cre-Lox system functions through a well-characterized mechanism wherein Cre recombinase recognizes LoxP sites, each consisting of two 13 bp inverted repeats flanking an 8 bp directional spacer region. Two Cre enzymes bind as dimers to the inverted repeats of a LoxP site, interacting with a similar dimer bound to another LoxP site to form a tetrameric complex. This synaptic complex facilitates DNA cleavage and strand exchange through a Holliday junction intermediate, resulting in recombination between the target sites. The system's key advantage lies in its ability to mediate these rearrangements without generating toxic DNA double-strand breaks, unlike CRISPR-Cas systems that induce breaks and rely on endogenous repair mechanisms [38].
However, systematic studies reveal significant constraints in Cre-Lox efficiency. Research examining Cre-mediated recombination in mice demonstrates that efficiency critically depends on inter-LoxP distance, with optimal recombination occurring when sites are spaced less than 4 kb apart for wildtype LoxP and 3 kb for mutant LoxP variants. Complete recombination failure was observed with wildtype LoxP sites spaced ≥15 kb apart or mutant Lox71/66 sites spaced ≥7 kb apart [1]. Additional factors influencing efficiency include the specific Cre-driver strain, zygosity of the floxed allele, and the genomic context of the target locus, all contributing to unpredictable and often mosaic recombination outcomes that complicate experimental design and interpretation [1].
PCE Systems: Engineering Solutions to Historical Limitations
The Programmable Chromosome Engineering (PCE and RePCE) platforms integrate three groundbreaking innovations that collectively address the fundamental constraints of traditional Cre-Lox technology:
Asymmetric Lox Variants: Through high-throughput recombination site modification, researchers developed novel Lox variants featuring asymmetric designs that reduce reversible recombination activity by over 10-fold while retaining high-efficiency forward recombination. This innovation ensures editing permanence by preventing the undo of genetic modifications [5] [17].
AI-Optimized Recombinase (AiCErec): Leveraging an AI-informed protein engineering system termed AiCE (AI-informed Constraints for protein Engineering), the team optimized Cre's multimerization interface, generating engineered variants with 3.5 times the recombination efficiency of wild-type Cre recombinase [21] [22].
Scarless Editing Strategy (Re-pegRNA): By combining prime editing technology with recombinase function, the researchers developed Re-pegRNA molecules that precisely replace residual Lox sites with the original genomic sequence after recombination, ensuring truly scarless genome modifications without unwanted leftover sequences [17] [6].
Table 1: Core Technological Components of Modern Genome Editing Systems
| Component | Traditional Cre-Lox | Enhanced PCE Systems |
|---|---|---|
| Recognition Site | Symmetrical LoxP sites | Asymmetric Lox variants |
| Recombinase Efficiency | Wild-type Cre | AI-engineered Cre (3.5× efficiency) |
| Reaction Reversibility | High (symmetrical sites) | 10-fold reduction |
| Sequence Footprint | Residual Lox sites (scarring) | Scarless editing via Re-pegRNA |
| Editing Scale | Kilobases | Kilobases to Megabases |
Comparative Performance Analysis: Quantitative Editing Metrics
Editing Efficiency and Scalability
Direct comparison of editing capabilities reveals dramatic differences between traditional Cre-Lox and PCE systems. While conventional Cre-Lox demonstrates effective recombination for genetic elements up to approximately 4 kb, its efficiency declines significantly with larger fragments and fails completely beyond 15 kb [1]. In stark contrast, PCE systems enable precise manipulations at unprecedented scales, achieving:
- Targeted integration of DNA fragments up to 18.8 kb
- Complete replacement of 5-kb DNA sequences
- Chromosomal inversions spanning 12 megabases
- Chromosomal deletions of 4 megabases
- Whole-chromosome translocations [21] [17]
This scalability breakthrough is further enhanced by substantially improved efficiency. The AI-guided engineering of Cre recombinase through the AiCErec system yielded variants with 250% higher recombination efficiency compared to wild-type Cre, directly addressing the activity limitations that have historically constrained large-scale DNA engineering [21] [7].
Editing Precision and Permanence
Editing fidelity represents another domain where PCE systems demonstrate marked improvement. Traditional Cre-Lox recombination leaves behind residual LoxP sites that create permanent genetic scars, potentially disrupting native gene function or regulation. Furthermore, the symmetrical nature of conventional Lox sites enables reversible reactions that can spontaneously undo genetic modifications, compromising editing stability [5].
PCE technologies address both limitations through integrated solutions. The Re-pegRNA scarless editing system harnesses prime editing technology to precisely restore the original genomic sequence after recombination, eliminating all foreign sequence footprints [17]. Meanwhile, the asymmetric Lox variants reduce reversible recombination by more than 10-fold, approaching the background level of negative controls and ensuring that edits remain permanent once introduced [6] [22].
Table 2: Performance Comparison of Genome Editing Technologies
| Performance Metric | Traditional Cre-Lox | PCE Systems |
|---|---|---|
| Maximum Efficient Scale | ~4 kb | >12 Mb |
| Recombination Efficiency | Baseline | 3.5× improvement |
| Editing Precision | Residual Lox sites | Scarless |
| Reaction Reversibility | High | 10-fold reduction |
| Double-Strand Breaks | No | No |
| Multiplexing Capacity | Limited | High (orthogonal sites) |
Experimental Methodologies: From Design to Validation
PCE Workflow: Integrated Engineering Approach
The experimental pipeline for Programmable Chromosome Engineering incorporates multiple advanced technologies in a coordinated workflow:
Key Experimental Applications and Validation
Researchers implemented this workflow to achieve groundbreaking genetic modifications in both plant and mammalian systems. In one notable demonstration, they engineered herbicide-resistant rice by creating a precise 315-kb chromosomal inversion—a feat that would be prohibitively difficult with conventional editing technologies [17] [22]. This involved:
- Target Identification: Selecting the specific 315-kb genomic region containing herbicide resistance genes.
- Asymmetric Lox Placement: Flanking the target region with engineered Lox variants in opposite orientations to enable inversion.
- AiCErec Delivery: Introducing the optimized recombinase to initiate recombination.
- Scar Removal: Utilizing Re-pegRNA to eliminate residual Lox sites after inversion.
- Phenotypic Validation: Confirming herbicide resistance in modified rice plants.
In mammalian systems, the team demonstrated the technology's therapeutic potential by achieving a 12-Mb inversion at human disease-related sites and performing scarless chromosome fusions, highlighting applications for modeling and potentially treating genetic disorders caused by large-scale chromosomal rearrangements [21].
Orthogonal Systems: Enabling Multiplexed Genome Engineering
A significant advancement in recombinase technology involves the development of orthogonal Lox systems that enable simultaneous, independent genetic modifications. Traditional Cre-Lox systems suffer from promiscuous recombination between all Lox sites present in a genome, preventing targeted multiplexing. Recent research has addressed this limitation by developing 63 symmetrical LoxP variants (LoxPsym) and testing 1,192 pairwise combinations to identify sites with minimal cross-reactivity [37].
This systematic characterization yielded a set of 16 fully orthogonal LoxPsym variants that enable specific recombination only between matched sites, allowing researchers to perform multiple independent editing events within the same cell. The utility of this orthogonal system has been demonstrated across diverse biological contexts, including prokaryotes (E. coli), eukaryotes (S. cerevisiae), and plants (Z. mays), highlighting its broad applicability for complex genome engineering projects requiring multiple precise manipulations [37].
Table 3: Research Reagent Solutions for Precision Genome Engineering
| Reagent/Cell Line | Function | Application Context |
|---|---|---|
| HEK293T cells | Mammalian editing validation | PCE testing in human cells |
| AiCErec recombinase | High-efficiency DNA recombination | PCE and RePCE systems |
| Asymmetric Lox variants | Non-reversible recognition sites | Stable edit maintenance |
| Re-pegRNA | Scarless editing | Lox site removal after editing |
| Orthogonal LoxPsym set | Multiplexed genome engineering | Simultaneous independent edits |
| REDMAPCre system | Spatiotemporal control | Light-activated recombination |
Emerging Alternatives: CRISPR-Based and Optogenetic Systems
CRISPR-Assisted DNA Modification Technologies
While recombinase-based systems excel at large-scale DNA rearrangements without double-strand breaks, CRISPR-based technologies offer complementary capabilities for targeted insertion. CRISPR-associated transposase (CAST) systems represent a particularly promising alternative, combining the programmability of CRISPR guidance with the DNA integration capability of transposases. Type I-F CAST systems enable integration of donor sequences up to approximately 15.4 kb in prokaryotic hosts, while type V-K variants have accommodated inserts up to 30 kb [38].
However, current CAST systems show limited efficiency in mammalian cells, with type I-F achieving only ~1% editing efficiency in HEK293 cells with 1.3 kb donors, and type V-K variants reaching a maximum of ~3% efficiency with 3.2 kb therapeutic donors at the AAVS1 locus in HEK293 cells. These efficiencies remain substantially below those achieved with PCE systems, particularly for larger DNA fragments [38].
Spatiotemporal Control Systems
Beyond efficiency and scale improvements, researchers have developed advanced optogenetic systems for precise temporal and spatial control of recombination. The REDMAPCre system represents a significant advancement in this domain, enabling rapid (1-second illumination) recombination activation with an 85-fold increase in reporter expression over background levels [13].
This red-light-controlled split-Cre system based on ΔPhyA/FHY1 interaction addresses critical limitations of previous photoactivatable tools, including background activation, slow kinetics, and poor tissue penetration. The technology has been successfully implemented in transgenic mouse models, enabling non-invasive light-dependent recombination across multiple organs and facilitating functional genetic studies with high spatiotemporal resolution [13].
The comprehensive analysis of editing fidelity across systems reveals distinct advantage profiles that should guide technology selection for specific research applications:
For large-scale chromosomal rearrangements (kilobase to megabase scale) requiring high precision and permanence, PCE systems offer unparalleled capabilities. Their scarless editing, reduced reversibility, and dramatically expanded scale range make them ideal for synthetic biology applications, crop improvement programs, and disease modeling requiring extensive genomic reorganization.
For multiplexed genome engineering requiring multiple independent modifications, orthogonal LoxPsym systems provide the necessary specificity to prevent unintended cross-reactions between different editing events, making them valuable for metabolic engineering and complex genetic circuit installation.
For applications demanding precise spatiotemporal control, optogenetic systems like REDMAPCre enable unprecedented temporal resolution and tissue-specific editing, particularly valuable for developmental biology studies and functional genetic analysis in animal models.
The rapid evolution of genome engineering technologies continues to expand the boundaries of possible genetic modifications. While Cre-Lox remains a valuable tool for standard applications, PCE and related advanced systems offer transformative capabilities for research domains requiring maximal precision, scale, and permanence—ushering in a new era of chromosome-scale engineering with profound implications for basic research, therapeutic development, and agricultural biotechnology.
Table of Contents
- Introduction to Chromosome Engineering Technologies
- Fundamental Mechanisms of Cre-Lox and PCE Systems
- Comparative Analysis of Editing Capabilities
- Experimental Protocols for Key Applications
- Decision Framework for Research Applications
Chromosome engineering, the ability to make targeted, large-scale changes to DNA, is a cornerstone of modern genetics. It enables researchers to model human diseases, understand gene function, and develop new crop varieties. For decades, the Cre-Lox recombination system has been the dominant technology in this space, allowing for precise deletions, inversions, and translocations of DNA segments. [11] However, its utility has been constrained by limitations such as reversible reactions and residual "scar" sequences. [5]
A recent breakthrough, Programmable Chromosome Engineering (PCE), overcomes these historical barriers. Developed by a team at the Chinese Academy of Sciences, PCE represents a significant advance by enabling precise, scarless manipulation of DNA fragments ranging from kilobases to megabases in size. [39] This guide provides a objective comparison of the application spectrum of Cre-Lox versus the novel PCE systems, equipping researchers with the data needed to select the optimal tool for their experimental goals.
Fundamental Mechanisms of Cre-Lox and PCE Systems
Understanding the core components and mechanisms of each system is crucial for appreciating their applications and limitations.
The Cre-Lox System: This system, derived from bacteriophage P1, consists of two parts: the Cre recombinase enzyme and its 34-base-pair DNA recognition site, loxP. [11] [40] [41] The outcome of recombination is determined by the orientation and location of the loxP sites. The system operates through a tetrameric complex that facilitates DNA cleavage and strand exchange. [11] [40]
- Core Components & Workflow: The diagram above illustrates the fundamental process of Cre-Lox recombination. The Cre recombinase recognizes and binds to loxP sites, forming a complex that catalyzes DNA recombination. The final genetic outcome—whether a deletion, inversion, or translocation—depends entirely on the orientation and genomic location of the loxP sites. [16] [40] [41]
The PCE Systems: This new technology integrates three key innovations to overcome Cre-Lox limitations:
- Asymmetric Lox Variants: Novel lox sites were designed to reduce reversible recombination by over 10-fold, ensuring edits are stable. [5]
- Engineered Recombinase (AiCErec): Using a protein-directed evolution system, the Cre recombinase was optimized for higher efficiency, yielding a variant with 3.5 times the recombination efficiency of the wild-type enzyme. [5]
- Scarless Editing Strategy: A method using specifically designed prime editing guide RNAs (Re-pegRNAs) was developed to precisely replace residual lox sites with the original genomic sequence after editing, leaving no trace. [5]
The integration of these technologies resulted in two programmable platforms: PCE and RePCE, which allow for flexible and precise manipulation of very large DNA segments. [5]
Comparative Analysis of Editing Capabilities
The following tables summarize the performance and operational characteristics of Cre-Lox and PCE systems based on published data.
Table 1: Documented Editing Outcomes and Efficiencies
| Editing Type | Cre-Lox System | PCE Systems | Key Citations |
|---|---|---|---|
| Deletion | Efficient for small segments (<4 kb); efficiency decreases with distance; up to 4 Mb demonstrated in ES cells. [1] [24] | 4 Mb deletion demonstrated in plant and animal cells. [39] | [1] [24] [39] |
| Insertion | Used with additional steps (e.g., recombinase-mediated cassette exchange). [11] | Targeted integration of an 18.8 kb DNA fragment. [39] [5] | [11] [39] |
| Inversion | Well-established; orientation of loxP sites dictates inversion. [11] [40] | 12 Mb chromosomal inversion; 315 kb inversion in rice as a functional proof-of-concept. [39] [5] | [11] [39] |
| Translocation | Possible when loxP sites are on different chromosomes. [16] [40] | Whole-chromosome translocation demonstrated. [39] | [16] [40] [39] |
| Replacement | Not a primary function. | Precise replacement of a 5 kb DNA sequence. [39] [5] | [39] |
Table 2: Key Technical and Operational Parameters
| Parameter | Cre-Lox System | PCE Systems | Key Citations |
|---|---|---|---|
| Typical Optimal Range | < 4 kb for high efficiency; up to megabases possible with lower efficiency. [1] [24] | Kilobase to megabase scale. [39] | [1] [24] [39] |
| Editing Precision | Leaves behind residual loxP "scar" sites. [11] | Scarless; residual sites can be replaced with the original sequence. [5] | [11] [5] |
| Reaction Reversibility | Inherently reversible, which can negate desired edits. [5] | Irreversible; asymmetric lox variants reduce reversibility to background levels. [5] | [5] |
| Recombinase Efficiency | Wild-type Cre efficiency is baseline. | AiCErec engineered variant shows 3.5x higher efficiency than wild-type Cre. [5] | [5] |
| Key Limitation | Reversibility, scar sequences, and lower efficiency for large edits. | Technology is novel; broader adoption and validation in diverse models are pending. | [1] [5] |
Experimental Protocols for Key Applications
Below are detailed methodologies for achieving specific chromosomal rearrangements with the Cre-Lox system, which has well-established protocols, and the innovative approach underpinning PCE.
Cre-Lox Protocol for Conditional Gene Knockout This is a foundational application for creating tissue-specific or inducible gene knockouts in animal models. [11] [16] [1]
- Generate Floxed Allele: Create a targeting construct where the critical exon(s) of the target gene are flanked ("floxed") by two loxP sites in the same orientation. The construct also includes a selectable marker (e.g., neomycin resistance), which may itself be floxed for later removal.
- ES Cell Targeting and Selection: Introduce the targeting construct into mouse embryonic stem (ES) cells. Select for successfully targeted clones using the resistance marker.
- Remove Selection Marker (Optional): Transiently express Cre recombinase in targeted ES cells to excise the floxed selection marker, leaving a single loxP site behind.
- Generate Founder Mice: Use targeted ES cells to generate chimeric mice and breed them to obtain germline transmission of the floxed allele.
- Cross with Cre Driver Line: Cross the floxed mouse line with a second transgenic line expressing Cre recombinase under a tissue-specific or inducible promoter (e.g., Cre-ERT activated by tamoxifen).
- Validate Recombination: In the offspring, confirm gene deletion via genotyping (PCR), Southern blot, or loss-of-protein assays (Western blot/immunohistochemistry) in the target tissue. [11] [1] [41]
PCE Workflow for Large-Scale, Scarless Engineering The PCE protocol is a integrated, multi-step process designed for precision and scale. [5]
- Design and Clone Asymmetric Lox Sites: Program the insertion positions and orientations for the novel, asymmetric lox variants (e.g., lox71/66) into your donor and target constructs using the PCE platform.
- Deliver PCE Components: Co-deliver the engineered AiCErec recombinase and the donor DNA containing the large DNA fragment flanked by the asymmetric lox sites into the plant or animal cells.
- Select for Integration: The AiCErec recombinase catalyzes efficient, forward-oriented integration of the large DNA fragment into the genome at the target site.
- Remove Residual Lox Sites (RePCE): To achieve a scarless edit, deliver a prime editor system equipped with a specifically designed Re-pegRNA. This guide RNA directs the editor to the residual lox site and provides the template to rewrite it precisely back to the original genomic sequence.
- Screen and Validate: Screen for successful editing events using PCR, sequencing, and functional assays. The final product is a seamless edit with no exogenous sequences. [5]
The Scientist's Toolkit: Essential Research Reagents
Successful implementation of these technologies requires a suite of specialized reagents and tools.
Table 3: Essential Research Reagents for Chromosome Engineering
| Reagent / Tool | Function | Example Use Case |
|---|---|---|
| Cre Recombinase | Enzyme that catalyzes site-specific recombination at loxP sites. [11] | Constitutively active form for permanent, ubiquitous gene deletion. |
| Inducible Cre (e.g., Cre-ERT2) | Cre fused to a modified estrogen receptor; activated by tamoxifen for temporal control. [16] | To induce gene knockout at a specific time point in adult animals, avoiding developmental effects. |
| Tissue-Specific Promoters | Genetic elements that drive Cre expression only in certain cell types. [11] [16] | To restrict genetic recombination to a specific organ or tissue (e.g., neurons, liver). |
| loxP Site | The 34 bp native recognition sequence for Cre recombinase. [11] [40] | The standard site for floxing alleles in traditional knockout models. |
| Mutant lox Sites (e.g., lox2272, lox511) | Variant lox sites that are not cross-compatible with loxP, enabling independent recombination events. [11] [40] | Used in DIO/FLEx switches to control gene expression in AAV vectors without interference. |
| Floxed Mouse Line | A transgenic mouse strain with a target gene segment flanked by loxP sites. [16] [1] | One half of the cross for generating conditional knockout models. |
| Cre Driver Mouse Line | A transgenic mouse strain expressing Cre in a defined pattern. [16] [1] | The other half of the cross; determines where and when the floxed gene is deleted. |
| AiCErec Recombinase | An engineered Cre variant with significantly higher recombination efficiency. [5] | Key component of the PCE system for efficient large-fragment integration. |
| Asymmetric Lox Variants | Novel lox sites engineered to minimize reversible recombination. [5] | Used in PCE systems to ensure stable, unidirectional integration of DNA fragments. |
| Re-pegRNA | A specially designed prime editing guide RNA for scarless removal of residual lox sites. [5] | Used in the RePCE system to restore the original genomic sequence after integration. |
Decision Framework for Research Applications
Choosing between Cre-Lox and PCE depends on the project's specific requirements. The following workflow can guide this decision:
For researchers using the Cre-Lox system, optimization is key. Recent systematic studies recommend using heterozygous floxed alleles and breeding Cre-driver mice between 8-20 weeks of age for highest efficiency. Critically, loxP spacing should be kept below 4 kb to ensure complete recombination and avoid mosaicism. [1] [3] The emergence of PCE systems does not render Cre-Lox obsolete; rather, it provides a powerful new option for ambitious engineering projects where scale and seamless precision are paramount.
Chromosome engineering, the ability to make defined and programmable changes to the genome, is a cornerstone of modern genetic research. For decades, the Cre-loxP system has been an indispensable tool in this field, enabling precise chromosomal rearrangements such as deletions, duplications, and inversions in mouse models and other organisms [24]. Its reliability and well-established methodology have made it a gold standard. However, the advent of CRISPR-based technologies has revolutionized genetic manipulation, introducing new levels of precision and versatility. Among these, prime editing represents a significant leap forward, offering the ability to install precise base substitutions, insertions, and deletions without requiring double-strand DNA breaks (DSBs) [42] [43].
This guide provides an objective comparison of the Cre-lox system's performance alongside and in integration with modern CRISPR and prime editing technologies. We will summarize key quantitative data, detail experimental protocols, and provide a clear framework for researchers to select the appropriate tool for their chromosome engineering objectives, whether in basic research or therapeutic drug development.
The Cre-loxP System
The Cre-lox system is a site-specific recombination technology derived from bacteriophage P1. The core component, Cre recombinase, catalyzes recombination between two 34-base pair DNA sequences known as loxP sites [24]. The outcome of the recombination is determined by the relative orientation and location of the loxP sites:
- loxP sites in the same orientation on a chromosome → Deletion of the intervening sequence.
- loxP sites in opposite orientation on a chromosome → Inversion of the intervening sequence.
- loxP sites on different chromosomes → Translocation [24].
A key feature of this system is its ability to act over very large genetic distances, facilitating rearrangements encompassing significant portions of a chromosome [24].
Prime Editing
Prime editing is a "search-and-replace" precision gene editing technology. It uses a fusion protein known as a prime editor, which consists of a Cas9 nickase (H840A) fused to an engineered reverse transcriptase (RT) [42] [43]. The system is guided by a specialized prime editing guide RNA (pegRNA) that performs two critical functions:
- It directs the editor to the target genomic locus.
- It encodes the desired edit(s) within its extension sequence, which includes a primer binding site (PBS) and a reverse transcription template (RTT) [43].
The process avoids double-strand breaks, leading to higher precision and fewer indel byproducts compared to traditional CRISPR-Cas9 nuclease editing [43]. The system has evolved through several generations (PE1 to PE5), with improvements in the reverse transcriptase enzyme and the co-delivery of factors to modulate cellular DNA repair, thereby enhancing efficiency and product purity [43].
Hybrid Technology: PASTE (Prime Editing with Recombinases)
A groundbreaking development is the fusion of prime editing with recombinase systems, creating a hybrid technology known as PASTE (Programmable Addition via Site-specific Targeting Elements). This approach leverages the strengths of both systems:
- Prime editing is used to precisely and cleanly install a recombinase "landing site" (e.g., the AttB site for the Bxb1 integrase) at a specific genomic location without a DSB.
- The Bxb1 integrase then catalyzes the efficient, high-fidelity integration of a large DNA cargo (from the AttP-containing donor plasmid) into the newly installed AttB site [44].
This technology enables the integration of very large DNA sequences (up to 35 kilobases has been demonstrated) that are far beyond the effective payload capacity of standard prime editing alone [44].
Table 1: Core Components and Primary Applications
| Technology | Core Enzymatic Machinery | Guide Molecule | Primary Editing Outcomes |
|---|---|---|---|
| Cre-loxP | Cre recombinase | None (relies on pre-placed loxP sites) | Deletions, Inversions, Duplications, Translocations |
| Prime Editing | Cas9 nickase + Reverse Transcriptase | pegRNA | Precise point mutations, small insertions & deletions (typically < 100 bp) |
| PASTE | Prime Editor + Bxb1 Integrase | pegRNA + DNA donor with AttP | Large, targeted DNA integration (theoretically unlimited) |
Quantitative Performance Comparison
The choice between Cre-lox, prime editing, and hybrid systems is heavily influenced by their performance characteristics, which vary significantly based on the intended edit.
Efficiency and Scalability of Genomic Rearrangements
The Cre-lox system demonstrates a unique ability to engineer large chromosomal segments. Foundational studies showed that while the efficiency of Cre-mediated recombination decreases with increasing genetic distance, it is not a limiting factor for most experimental designs. Rearrangements encompassing up to three-quarters of mouse chromosome 11 (spanning several centimorgans, equivalent to tens of megabases) have been successfully engineered in mouse embryonic stem (ES) cells [24]. The main limitation is often cellular viability, as larger deletions can be lethal [24].
In contrast, prime editing excels at the other end of the size spectrum. Its efficiency is optimal for smaller edits. A systematic 2025 analysis of Cre-mediated recombination—which shares similar "floxing" logic with the distance between lox sites—provides insightful parallel data. This study found that recombination efficiency is highest when the distance between recognition sites is short [1].
Table 2: Performance Metrics by Edit Size and Type
| Edit Type | Technology | Typical Size Range | Key Performance Metric | Reported Efficiency / Limitation |
|---|---|---|---|---|
| Large Deletion/Inversion | Cre-loxP | Several Mb (e.g., 3/4 of a chromosome) | Success in ES cells | Achievable, efficiency decreases with distance [24] |
| Small Deletion | Cre-loxP | N/A (distance between loxP sites) | Recombination Efficiency | Up to 11% for a 2 cM (~4 Mb) deletion in ES cells [24] |
| Small Edit | Prime Editing | < 100 bp | Editing Efficiency | Highly dependent on pegRNA design and cell type; optimized systems (PE4/PE5) can achieve high efficiency [43] |
| Large Integration | PASTE | > 5 kb, up to ~35 kb | Integration Efficiency | Highly efficient integration of large cargos (e.g., entire transgenes) [44] |
| Conditional Allele (in vivo) | Cre-loxP | N/A (distance between loxP sites) | Completeness & Mosaicism | Highly dependent on inter-loxP distance: ~54% complete recombination at 0.8 kb; fails at ≥15 kb [1] |
Precision and Specificity
- Cre-loxP: The system is highly specific to loxP sites, with minimal off-target effects. However, this specificity requires pre-placement of loxP sites in the genome, which is a time-consuming and resource-intensive process. A potential issue is mosaicism in vivo, where recombination does not occur in all cells of a tissue [1] [45].
- Prime Editing: Prime editing is renowned for its precision. It does not require DSBs, resulting in a very high edit-to-indel ratio and minimal off-target activity [43]. Cas9-dependent off-target editing is much less frequent with prime editors than with Cas9 nuclease, as editing requires three distinct DNA hybridization events (spacer, PBS, and 3' homology) [43].
- PASTE: While the integration step is efficient, a noted limitation is that the process is not entirely "scarless." Residual portions of the AttB/AttP recognition sites may remain in the final edited genome [44].
Experimental Protocols and Workflows
Cre-loxP Chromosome Engineering in ES Cells
This classic protocol for generating engineered mouse models involves multiple steps in embryonic stem (ES) cells [24]:
- Targeting Vector Construction: Two different targeting vectors are designed. One contains a loxP site and the 5' half of a selectable marker (e.g., 5' Hprt). The other contains a second loxP site and the 3' half of the marker (3' Hprt).
- Sequential ES Cell Targeting: ES cells are sequentially transfected with the two targeting vectors to create a cell line where the two loxP sites are correctly positioned in the genome.
- Cre-Mediated Recombination: The engineered ES cells are transfected with a Cre recombinase expression plasmid (e.g., pOG231).
- Selection and Analysis: Cells that have undergone the desired recombination event are selected using medium (e.g., HAT medium) that selects for the reconstituted full-length Hprt gene. Surviving clones are picked and validated by PCR and Southern blot analysis [24].
Cre-loxP Chromosome Engineering Workflow
Prime Editing in Mammalian Cells
The following protocol is adapted from best practices for conducting prime editing experiments in mammalian cells [43]:
- pegRNA Design: Design the pegRNA to include:
- A spacer sequence (typically 20 nt) to target the desired locus.
- A reverse transcription template (RTT, 10-25 nt) that encodes the desired edit.
- A primer binding site (PBS, 10-15 nt) complementary to the nicked genomic DNA strand.
- Prime Editor Selection: Choose an appropriate prime editor system (e.g., PE2, PE3, PE4, or PE5). PE4 and PE5, which transiently inhibit mismatch repair, often provide the best balance of high efficiency and low indel rates [43].
- Delivery: Co-deliver the prime editor (as mRNA or plasmid) and the pegRNA (as a synthesized RNA or expressed from a plasmid) into the target cells using methods such as electroporation or lipid nanoparticles.
- Analysis: After allowing time for editing and expression, harvest genomic DNA and analyze editing efficiency using next-generation sequencing (NGS) or high-resolution melt analysis.
Prime Editing Experimental Workflow
The Scientist's Toolkit: Essential Research Reagents
Successful implementation of these technologies requires a suite of reliable reagents. The table below details key solutions for setting up and conducting experiments.
Table 3: Essential Research Reagents and Resources
| Reagent / Resource | Function / Description | Example Use Cases |
|---|---|---|
| Conditional loxP Alleles | Genetically modified alleles where a critical gene segment is flanked by loxP sites. | Tissue-specific or inducible gene knockout studies in model organisms [24] [45]. |
| Inducible Cre-ERT² Lines | Transgenic lines expressing a Cre recombinase fused to a modified estrogen receptor, activated by tamoxifen. | Temporal control of recombination for studying gene function at specific developmental stages [45]. |
| pegRNA Synthesis Kits | Commercial kits for in vitro transcription or assembly of long, complex pegRNAs. | Generating the essential guide molecule for prime editing experiments [42] [43]. |
| Engineered Prime Editor Plasmids | Optimized plasmids (e.g., PEmax) for high-level expression of prime editor components in mammalian cells. | Maximizing prime editing efficiency in difficult-to-transfect cell types [43]. |
| Bxb1 Integrase & AttP Donor | The recombinase enzyme and a donor plasmid containing the AttP site and the cargo for integration. | Performing large DNA integrations using the PASTE technology [44]. |
| MLH1dn Protein/Plasmid | A dominant-negative version of the MLH1 protein to transiently inhibit mismatch repair. | Used with the PE4/PE5 systems to boost prime editing efficiency by preventing edit reversal [43]. |
The landscape of chromosome engineering is no longer defined by a single dominant technology but is instead a rich ecosystem where classic and modern tools complement each other. The Cre-loxP system remains unparalleled for large-scale chromosomal rearrangements and sophisticated genetic models relying on conditional and inducible mutagenesis. Its well-characterized behavior and extensive historical use in thousands of genetically modified mouse lines ensure its continued relevance.
Prime editing has established a new benchmark for precision, enabling a wide range of small-scale corrections and mutations with minimal undesirable byproducts. Its primary limitation—the inefficient integration of large DNA sequences—is being directly addressed by the development of hybrid systems. PASTE and similar technologies that couple prime editing with recombinases represent the cutting edge, effectively breaking the payload barrier and opening the door to the therapeutic correction of large genetic defects.
For the researcher designing a chromosome engineering project, the strategic choice is clear:
- For massive chromosomal rearrangements or advanced in vivo animal models, Cre-lox is the established and effective choice.
- For correcting point mutations or installing small, precise edits in any cell type, prime editing is the superior technology.
- For targeted integration of large transgenes (e.g., for CAR-T cell engineering), hybrid systems like PASTE are the most promising and efficient path forward.
The future lies not in the replacement of one technology by another, but in their intelligent integration, leveraging the unique strengths of each to answer complex biological questions and develop next-generation genetic medicines.
Conclusion
The advent of Programmable Chromosome Engineering (PCE) systems marks a paradigm shift in genome engineering, effectively overcoming the long-standing limitations of the Cre-Lox system. By integrating AI-driven protein engineering, novel recombination sites, and scarless editing strategies, PCE and RePCE deliver unprecedented efficiency, precision, and scale—enabling manipulations of up to megabases and whole-chromosome translocations. This technological leap has immediate, transformative implications, accelerating the development of resilient crops and opening new avenues for modeling and treating complex genetic disorders in biomedicine. For researchers and drug development professionals, these systems represent the new vanguard of genetic manipulation tools, poised to unlock the next generation of advances in synthetic biology and therapeutic development.
References
- 1. Systematic optimization and prediction of cre recombinase for ... [https://genomebiology.biomedcentral.com/articles/10.1186/s13059-025-03560-3]
- 2. Cre-Lox and Other Site-Specific Recombinases [https://www.addgene.org/collections/cre-lox/]
- 3. Large-Scale Genome-Wide Optimization and Prediction of ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11212873/]
- 4. Iterative recombinase technologies for efficient and precise ... [https://www.sciencedirect.com/science/article/abs/pii/S0092867425008001]
- 5. Programmable Chromosome Engineering systems enable ... [https://www.news-medical.net/news/20250804/Programmable-Chromosome-Engineering-systems-enable-precise-DNA-manipulations.aspx]
- 6. Scientists just cracked the code to editing entire ... [https://www.sciencedaily.com/releases/2025/08/250805041612.htm]
- 7. Rewriting the Code: New Tool Accelerates Gene Editing - QPS [https://www.qps.com/2025/09/03/rewriting-the-code-new-tool-accelerates-gene-editing/]
- 8. One-step Cre-loxP organism creation by TAx9 [https://www.nature.com/articles/s42003-025-07759-9]
- 9. Perspective: The current state of Cre driver mouse lines in ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10087282/]
- 10. Cre-lox and Other Recombinase Systems: An Overview [https://resources.jax.org/jax-blog/cre-lox-and-other-recombinase-systems-an-overview]
- 11. Cre-Lox recombination [https://en.wikipedia.org/wiki/Cre-Lox_recombination]
- 12. Mouse Cre-LoxP system: general principles to determine ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6333611/]
- 13. A rapid and efficient red-light-activated Cre recombinase ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12342929/]
- 14. Potential Pitfalls When Using the Cre-LoxP System in ... [https://www.oncoscience.us/news/pr/potential-pitfalls-when-using-the-cre-loxp-system-in-cancer-research/]
- 15. Establishment of Cre/LoxP-mediated multifunctional reporter ... [https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0325444]
- 16. Cre-lox & Other Recombinase Systems [https://www.jax.org/news-and-insights/jax-blog/2025/july/recombinase-systems]
- 17. Scientists Achieve Megabase-Scale Precision Genome ... [https://english.cas.cn/Special_Reports/2025q3/202510/t20251027_1094862.shtml]
- 18. Caixia Gao Team Achieves Megabase-Scale Precision ... [http://gaolab.genetics.ac.cn/en/new/202508/t20250806_650417.html]
- 19. sciencedirect.com/topics/neuroscience/ chromosome - engineering [https://www.sciencedirect.com/topics/neuroscience/chromosome-engineering]
- 20. Chinese scientists create tool to precisely edit millions of DNA segments [https://interestingengineering.com/science/china-tool-precisely-edit-millions-dna-segments]
- 21. Iterative recombinase technologies for efficient and precise genome... [https://pubmed.ncbi.nlm.nih.gov/40763736/]
- 22. Chinese Researchers Develop Breakthrough Megabase ... [https://www.seedworld.com/europe/2025/08/06/chinese-researchers-develop-breakthrough-megabase-scale-genome-editing-technologies/]
- 23. Site-specific recombination of asymmetric lox sites ... [https://www.sciencedirect.com/science/article/abs/pii/S096808960501206X]
- 24. Engineering Mouse Chromosomes with Cre-loxP [https://pmc.ncbi.nlm.nih.gov/articles/PMC85158/]
- 25. Cre-lox-Based System for Multiple Gene Deletions and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC1828656/]
- 26. Breakthrough in Genome Editing: Scientists Attain ... [https://bioengineer.org/breakthrough-in-genome-editing-scientists-attain-megabase-scale-precision-in-eukaryotic-cells/]
- 27. Prime Editing for Crop Improvement: A Systematic Review ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12385346/]
- 28. A long journey towards genome editing technologies in ... [https://www.frontiersin.org/journals/genome-editing/articles/10.3389/fgeed.2025.1663352/full]
- 29. 4 essential steps to verify your Cre-lox model [https://www.jax.org/news-and-insights/jax-blog/2014/april/four-essential-steps-to-verify-your-cre-lox-model]
- 30. Rewriting the Genome: The Dawn of Megabase-Scale ... [https://www.ailurus.bio/post/rewriting-the-genome-the-dawn-of-megabase-scale-programmable-rearrangement]
- 31. Cre/loxP Recombination System: Applications, Best ... [https://www.taconic.com/resources/creloxp-recombination-system-applications-best-practices-breeding-strategies]
- 32. Generation of Marker-Free Transgenic Rice Resistant to ... [https://www.frontiersin.org/journals/plant-science/articles/10.3389/fpls.2021.644437/full]
- 33. Lox'd in translation: contradictions in the nomenclature ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8116776/]
- 34. Unidirectional Cre-mediated genetic inversion in mice ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC275577/]
- 35. Evaluation of a library of loxP variants with a wide range of ... [https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0276657]
- 36. Scientists Achieve Megabase-Scale Precision Genome Editing in... [http://english.genetics.cas.cn/news_/researchnews/202508/t20250804_1048980.html]
- 37. Orthogonal LoxPsym sites allow multiplexed site-specific ... [https://www.nature.com/articles/s41467-024-44996-8]
- 38. Advances in large-scale DNA engineering with the ... [https://www.nature.com/articles/s12276-025-01530-0]
- 39. Chinese Scientists Unveil Major Breakthrough in Large ... [https://english.cas.cn/newsroom/cas_media/202508/t20250806_1049222.shtml]
- 40. Cre-Lox Recombination – Introduction | abm Inc. [https://info.abmgood.com/cre-lox-recombination-introduction]
- 41. Plasmids 101: Cre-lox [https://blog.addgene.org/plasmids-101-cre-lox]
- 42. Prime Editing as a Precision Gene Editing Tool [https://www.synthego.com/guide/crispr-methods/prime-editing/]
- 43. Designing and executing prime editing experiments in ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9799714/]
- 44. Large DNA Integrations With Prime Editing and ... [https://crisprmedicinenews.com/news/large-dna-integrations-with-prime-editing-and-recombinases/]
- 45. A comparative analysis of microglial inducible Cre lines [https://www.sciencedirect.com/science/article/pii/S2211124723010422]