CRISPR-Cas Systems: A Comprehensive Guide to Types I-VI Classification, Mechanisms, and Biomedical Applications
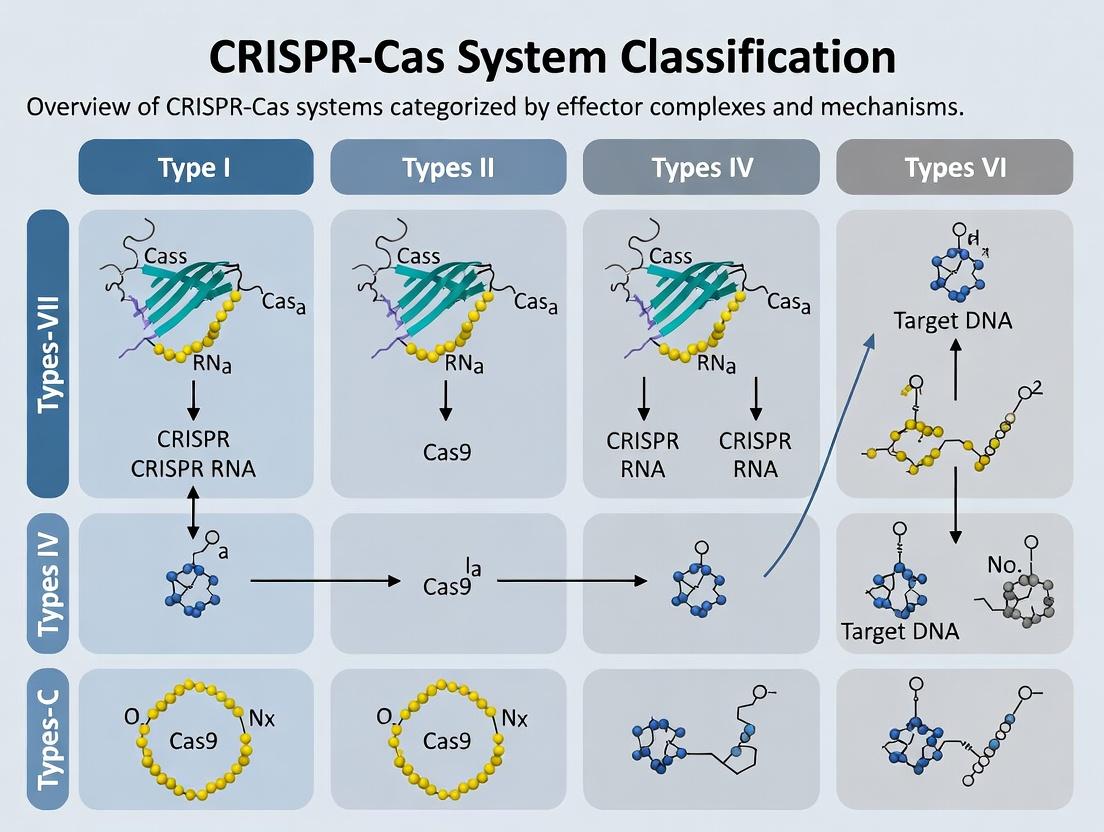
This article provides a detailed, up-to-date exploration of the classification and functional diversity of CRISPR-Cas systems, spanning Types I through VI.
CRISPR-Cas Systems: A Comprehensive Guide to Types I-VI Classification, Mechanisms, and Biomedical Applications
Abstract
This article provides a detailed, up-to-date exploration of the classification and functional diversity of CRISPR-Cas systems, spanning Types I through VI. It serves as a foundational reference for researchers, detailing the core molecular architectures and mechanisms of each type. The content progresses to examine current methodological applications in genetic engineering, diagnostics, and therapeutics, while addressing common experimental challenges and optimization strategies. Finally, it offers a comparative analysis of system selection criteria, validation protocols, and emerging tools. Designed for scientists and drug development professionals, this guide synthesizes the latest research to inform experimental design and future translational research.
Decoding CRISPR-Cas Diversity: Core Architectures and Mechanisms of Types I-VI
CRISPR-Cas systems constitute a heritable, adaptive immune mechanism that enables prokaryotes to defend against invasive genetic elements such as phages and plasmids. The core principle involves the acquisition of short DNA sequences from invaders into the host's CRISPR array, their transcription into guide RNAs (crRNAs), and the crRNA-guided targeting and cleavage of homologous foreign nucleic acids by Cas effector proteins. This immune response exhibits sequence specificity and memory, hallmarks of adaptive immunity. Research into the classification of these systems into Types I-VI (and beyond) has revealed a staggering diversity of molecular architectures and effector mechanisms, which has profound implications for understanding microbial ecology and evolution, as well as for biotechnological and therapeutic applications.
Core Mechanisms and Classification Framework
The adaptive immune function operates in three distinct stages:
- Adaptation: Cas1-Cas2 complexes mediate the acquisition of short protospacers from invading DNA and their integration as new spacers into the CRISPR array.
- Expression: The CRISPR array is transcribed and processed into mature crRNAs, each containing a spacer sequence guide.
- Interference: The crRNA assembles with one or more Cas proteins into an effector complex that surveils the cell, identifies complementary nucleic acids (protospacers), and neutralizes the threat.
The classification of CRISPR-Cas systems into Types (I-VI) is based on the identity of the cas gene repertoire, particularly the signature effector protein(s) responsible for interference. Recent research has expanded this classification to include additional types (e.g., Type VII, IX) and subtypes, based on continued genomic discovery.
Table 1: Core Features of Major CRISPR-Cas System Types (I-VI)
| Type | Signature Effector Complex | Target Nucleic Acid | Cleavage Mechanism | Protospacer Adjacent Motif (PAM) Requirement | Key Distinguishing Feature |
|---|---|---|---|---|---|
| I | Multi-subunit Cascade (Cas3) | dsDNA | Cas3 helicase-nuclease | Yes (upstream/downstream) | Large, multi-protein surveillance complex; Cas3 cleaves displaced strand. |
| II | Single effector Cas9 | dsDNA | RuvC & HNH nuclease domains | Yes (downstream) | Single, large effector protein; requires trans-activating crRNA (tracrRNA). |
| III | Multi-subunit Csm (III-A) / Cmr (III-B) | ssRNA, ss/dsDNA* | Cas7-like subunits cleave RNA | No (for RNA) | Transcription-dependent DNA targeting; collateral RNase activity. |
| IV | Multi-subunit (DinG family nuclease) | dsDNA (presumed) | Presumed DinG helicase/nuclease | Likely | Minimal, poorly characterized system; often on plasmids. |
| V | Single effector Cas12 (e.g., Cas12a) | dsDNA, ssDNA* | RuvC-like nuclease domain | Yes (T-rich) | Single effector; unleashes non-specific ssDNase collateral activity post-target recognition. |
| VI | Single effector Cas13 (e.g., Cas13a) | ssRNA | Two HEPN domains | No (flanking sequence effects) | Targets RNA; unleashes non-specific RNase collateral activity post-target recognition. |
*Type III can cleave DNA via transcription-coupled mechanisms. Type V effectors like Cas12 can cleave ssDNA non-specifically after activation.
Key Experimental Protocols in CRISPR-Cas Research
Protocol forIn VitroPAM Determination (Type II, V Systems)
Objective: Identify the Protospacer Adjacent Motif (PAM) sequence required for Cas protein targeting. Methodology:
- Library Construction: Generate a plasmid library containing a randomized NNNN sequence (PAM candidate) adjacent to a fixed protospacer sequence.
- In Vitro Cleavage: Incubate the plasmid library with purified Cas effector protein (e.g., Cas9, Cas12a) and a complementary crRNA.
- Selection & Sequencing: Isolate the linearized (cleaved) DNA products via gel electrophoresis or size-selection. Amplify and deep-sequence the PAM region from both the cleaved and uncleaved pools.
- Bioinformatics Analysis: Compare the sequence enrichment/depletion in the cleaved pool versus the input library to determine the PAM consensus sequence.
Protocol for Assessing Collateral Activity (Type VI, V Systems)
Objective: Measure the non-specific (collateral) nuclease activity of Cas13 (RNase) or Cas12 (DNase) upon target recognition. Methodology:
- Fluorescent Reporter Design: For Cas13, use an ssRNA reporter with a fluorophore-quencher pair. For Cas12, use an ssDNA reporter with a fluorophore-quencher pair.
- Reaction Setup: Combine the Cas effector, crRNA, and fluorescent reporter in a reaction buffer. Divide into two tubes.
- Activation: Add the specific target RNA (for Cas13) or DNA (for Cas12) to the experimental tube. Leave it out of the control tube.
- Measurement: Monitor real-time fluorescence (e.g., in a plate reader). A sharp increase in fluorescence in the experimental tube only indicates collateral cleavage of the reporter upon target-specific activation.
Protocol forIn VivoInterference Assay
Objective: Quantify the immune defense capability of a CRISPR-Cas system in its native host. Methodology:
- Strain Preparation: Use a prokaryotic strain harboring the CRISPR-Cas system of interest (immune) and an isogenic strain lacking it (non-immune).
- Challenge: Infect both cultures with a serial dilution of the target phage (or transform with a targeting plasmid) at a known multiplicity of infection (MOI).
- Efficiency of Plating (EOP): Plate the infected cultures on soft agar lawns. Count plaque-forming units (PFU) or plasmid transformants.
- Calculation: EOP = (PFU on immune host) / (PFU on non-immune host). A functional system yields an EOP reduction of several orders of magnitude (e.g., 10^-4 to 10^-6).
Table 2: Quantitative Comparison of Key CRISPR-Cas System Parameters
| Parameter | Type I (Cascade-Cas3) | Type II (Cas9) | Type III (Csm/Cmr) | Type V (Cas12a) | Type VI (Cas13a) |
|---|---|---|---|---|---|
| Typical crRNA Length (nt) | ~60-70 | ~100 (crRNA+tracrRNA) | ~40-50 | ~43-45 | ~64-66 |
| Interference Speed (bp/s)* | ~250-500 (Cas3) | ~1-10 (cleavage fast) | Not quantified | Fast cleavage, slow turnover | Fast cleavage, slow turnover |
| Interference Fidelity (% off-target) | <1% (high) | Varies (0.1-50%+; can be high) | High (requires transcription) | High (strict PAM) | Moderate (tolerant of mismatches) |
| System Size (kbp in locus) | ~15-20 | ~4-5 | ~20-25 | ~3-4 | ~3.5-4.5 |
| Primary Biotech Application | Large-scale genomic edits, screens | Gene knockout, activation, imaging | RNA knockdown, diagnostics | DNA editing, diagnostics (DETECTR) | RNA editing, diagnostics (SHERLOCK) |
*Helicase-dependent rates for DNA-targeting systems; represents translocation/search speed.
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for CRISPR-Cas Research
| Reagent / Material | Function in Research | Example Application |
|---|---|---|
| Purified Recombinant Cas Proteins | Core effector enzyme for in vitro assays, structural studies, and diagnostic development. | In vitro cleavage assays, PAM determination, creating ribonucleoprotein (RNP) complexes for editing. |
| crRNA/tracrRNA Oligonucleotides | Synthetic guide RNAs that program the sequence specificity of the Cas effector. | Designing targeted interference experiments; creating libraries for high-throughput screens. |
| In Vitro Transcription Kits | Generate large quantities of crRNA, tracrRNA, or target RNA/DNA for biochemical assays. | Producing substrates for Cas13 collateral activity assays or Cas9 cleavage kinetics. |
| Phage Genomic DNA Libraries | Diverse source of protospacers for studying adaptation or host-range evolution. | Performing in vivo spacer acquisition (adaptation) experiments. |
| Fluorescent-Quencher Reporter Probes | Detect nuclease (especially collateral) activity in real-time. | SHERLOCK (Cas13) and DETECTR (Cas12) diagnostic platforms; measuring enzyme kinetics. |
| Electrocompetent Prokaryotic Strains | Host cells for in vivo interference and adaptation assays, requiring efficient DNA transformation. | Measuring plasmid interference EOP; studying native CRISPR-Cas immune function. |
| Next-Generation Sequencing (NGS) Library Prep Kits | High-throughput analysis of spacer acquisition, phage escape mutants, and off-target effects. | Profiling the dynamics of CRISPR array evolution; genome-wide off-target assessment. |
| Single-Molecule Imaging Reagents (e.g., TIRF) | Visualize real-time binding, search, and cleavage kinetics of CRISPR effectors on DNA/RNA. | Studying the target search mechanism of Cascade or Cas9 at the molecular level. |
The Evolutionary Basis for Class 1 (Multi-Subunit) and Class 2 (Single-Effector) Division
Within the established classification of CRISPR-Cas systems (Types I-VI), a fundamental evolutionary and functional division exists between Class 1 and Class 2 systems. This division, based on the architecture of the effector module responsible for crRNA processing and target interference, has profound implications for mechanistic understanding and biotechnological application. Class 1 systems (Types I, III, and IV) utilize multi-subunit effector complexes, whereas Class 2 systems (Types II, V, and VI) employ single, large effector proteins. This guide details the evolutionary origins, comparative biochemistry, and experimental dissection of this core dichotomy, providing a framework for ongoing research and therapeutic development.
Evolutionary Origins and Phylogenetic Distribution
The evolution of CRISPR-Cas systems is marked by multiple horizontal gene transfer events and recombination. Phylogenetic analysis suggests Class 1 systems are evolutionarily older and more widespread in prokaryotes. Class 2 systems, particularly Type II and Type V, appear to have evolved more recently, likely derived from mobile genetic elements like Tn7-like transposons for Type V-K (Cas12) and ancestral endonucleases for Type VI (Cas13).
Table 1: Core Characteristics of Class 1 vs. Class 2 Systems
| Feature | Class 1 Systems (Types I, III, IV) | Class 2 Systems (Types II, V, VI) |
|---|---|---|
| Effector Architecture | Multi-protein complex (e.g., Cascade, Csm/Cmr complex) | Single effector protein (e.g., Cas9, Cas12, Cas13) |
| Prevalence in Prokaryotes | ~90% of CRISPR-containing bacteria; ~60% of CRISPR-containing archaea | ~10% of CRISPR-containing bacteria; rare in archaea |
| Ancestral Origin | Likely derived from ancestral prokaryotic defense modules | Likely evolved from mobile genetic elements (TnpB/IS605 family for Cas12; HEPN domains for Cas13) |
| crRNA Processing | Often requires separate processing complex (Cas6-like) | Typically intrinsic RNase activity within the effector (Cas9, Cas12) or host RNase III (Type II) |
| Target Interference | DNA (I, IV) or RNA/DNA (III) | DNA (II, V) or RNA (VI) |
| Genomic Locus Size | Large (>10 genes common) | Compact (1-4 genes common) |
Mechanistic and Structural Comparison
Class 1: Multi-Subunit Effector Complexes
Class 1 effectors are assemblies of multiple Cas protein subunits. For example, E. coli Type I-E Cascade comprises 11 subunits of 5 different Cas proteins (CasA,B,C,D,E). crRNA is stably integrated into the backbone of the complex. Target recognition involves concerted conformational changes across subunits, leading to recruitment of the dedicated nuclease Cas3 for DNA cleavage in Type I systems.
Class 2: Single-Effector Proteins
Class 2 effectors integrate all essential functions—crRNA binding, target recognition, and cleavage—into a single polypeptide. Cas9 (Type II) utilizes two separate nuclease domains (HNH, RuvC) to cut both strands of target DNA. Cas12 (Type V) employs a single RuvC domain for staggered DNA cleavage and exhibits trans-cleavage activity. Cas13 (Type VI) targets RNA and also displays robust trans-cleavage upon activation.
Table 2: Quantitative Biochemical Parameters of Representative Effectors
| Effector (Class, Type) | Molecular Weight (kDa) | Complex Subunits | Cleavage Rate (kobs, min⁻¹) | PAM/PFS Requirement | Key Reference (Example) |
|---|---|---|---|---|---|
| Cascade (Class 1, I-E) | ~405 kDa (complex) | 11 (CasA1B2C6D1E1) | Cas3 recruitment: ~1-2 min⁻¹ | 5'-protospacer adjacent motif (PAM) | Jackson et al., Nature, 2014 |
| Cas9 (Class 2, II-A) | ~160 kDa (single protein) | 1 (dimer possible) | DNA cleavage: ~0.1-1 min⁻¹ | 3'-NGG (SpCas9) | Jinek et al., Science, 2012 |
| Cas12a (Class 2, V-A) | ~130 kDa (single protein) | 1 | cis-cleavage: ~0.5 min⁻¹; trans-cleavage: >10³ min⁻¹ | 5'-TTTV | Chen et al., Science, 2018 |
| Cas13a (Class 2, VI-A) | ~130 kDa (single protein) | 1 | trans-cleavage: >10³ min⁻¹ | protospacer flanking site (PFS) | Abudayyeh et al., Nature, 2016 |
Key Experimental Protocols for Comparative Analysis
Protocol:In VitroReconstitution of a Class 1 Effector Complex
Objective: To purify individual subunits and assemble a functional Type I-E Cascade complex.
- Cloning: Express genes for CasA, CasB, CasC, CasD, and CasE from E. coli with N-terminal His-tags (or other affinity tags) in separate E. coli BL21(DE3) expression vectors.
- Protein Purification: Lyse cells, purify each subunit using immobilized metal affinity chromatography (IMAC), followed by size-exclusion chromatography (SEC) in buffer (20 mM HEPES pH 7.5, 150 mM KCl, 1 mM DTT, 5% glycerol).
- Complex Assembly: Mix subunits at molar ratios of A1:B2:C6:D1:E1. Incubate at 4°C for 1 hour with gentle agitation.
- Complex Purification: Load mixture onto a Superose 6 Increase SEC column. Collect the high molecular weight peak corresponding to the assembled complex.
- Validation: Analyze by negative-stain electron microscopy and gel-shift assay with cognate crRNA.
Protocol: MeasuringTrans-Cleavage Kinetics of Class 2 Effectors (Cas12/Cas13)
Objective: To quantify collateral cleavage activity, a hallmark of many Class 2 effectors.
- Sample Preparation: Purify effector protein (e.g., LbaCas12a). Synthesize crRNA and target DNA activator (dsDNA with correct PAM).
- Reaction Setup: In a 96-well plate, mix 50 nM Cas12a, 60 nM crRNA, and 100 nM activator DNA in reaction buffer (20 mM HEPES pH 7.5, 50 mM NaCl, 5 mM MgCl₂). Pre-incubate for 10 min at 37°C.
- Initiate Trans-Cleavage: Add a fluorescent reporter (e.g., 500 nM ssDNA probe labeled with 6-FAM and BHQ-1). Immediately transfer plate to a real-time PCR instrument or fluorimeter.
- Data Acquisition: Monitor fluorescence (excitation 485 nm, emission 520 nm) every 30 seconds for 1-2 hours.
- Analysis: Fit the resulting fluorescence vs. time curve to a first-order exponential equation to determine the observed rate constant (kobs).
Visualizing Functional Pathways and Evolution
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Research Reagent Solutions for CRISPR-Cas Studies
| Reagent / Material | Function / Application | Example Vendor / Cat. No. (Representative) |
|---|---|---|
| High-Fidelity DNA Polymerases | Cloning of Cas genes and CRISPR arrays without mutations. | NEB Q5, Thermo Fisher Phusion |
| Affinity Chromatography Resins | Purification of His-, MBP-, or GST-tagged Cas proteins and subunits. | Cytiva HisTrap HP, Ni-NTA Agarose (QIAGEN) |
| Size-Exclusion Chromatography Columns | Polishing protein preps and isolating assembled effector complexes. | Cytiva Superose 6 Increase, Bio-Gel P-6 (Bio-Rad) |
| Fluorescent Nucleic Acid Reporters | Quantifying nuclease activity (especially trans-cleavage). | IDT (FAM-Quencher probes), Biosearch Technologies (Black Hole Quenchers) |
| In Vitro Transcription Kits | Generating crRNA and target RNA for biochemical assays. | NEB HiScribe T7, Thermo Fisher MEGAscript |
| Electrophoretic Mobility Shift Assay (EMSA) Gels | Analyzing protein-nucleic acid complex formation (e.g., Cascade-crRNA). | Native PAGE gels (4-12%), Bio-Rad Mini-PROTEAN system |
| Real-Time PCR Instrument | Kinetic measurement of fluorescent cleavage reporter assays. | Bio-Rad CFX96, Applied Biosystems 7500 |
| Cas9 Nuclease (WT & variants) | Positive control for Class 2 effector biochemical studies. | NEB (M0386), IDT (Alt-R S.p. Cas9) |
Within the established classification of CRISPR-Cas adaptive immune systems (Types I-VI), Type I systems represent the most prevalent and diverse group. They are characterized by a multi-subunit effector complex for crRNA-guided target recognition and the signature Cas3 nuclease for target degradation. This whitepaper provides an in-depth technical analysis of the core machinery: the CRISPR-associated complex for antiviral defense (Cascade) and the Cas3 helicase-nuclease.
Structural and Functional Composition of the Cascade Complex
The Cascade complex serves as the surveillance module, responsible for crRNA maturation and double-stranded DNA (dsDNA) target recognition. Its composition varies among subtypes (I-A to I-G), with the well-studied E. coli Type I-E system serving as the archetype.
Table 1: Subunit Composition of Type I-E Cascade (E. coli)
| Subunit | Gene | Copies | Primary Function |
|---|---|---|---|
| Cas5e | cas5e |
1 | crRNA 5'-handle binding; contributes to backbone. |
| Cas6e | cas6e |
1 | crRNA processing: Cleaves pre-crRNA into mature spacers. Retained in complex. |
| Cas7e | cas7e |
6 | Forms the helical backbone; binds the spacer region of the crRNA. |
Cas8e (Cse1) |
cas8e |
1 | Large subunit; initiates dsDNA binding and R-loop formation; contains the PAM-interacting domain. |
Cas11e (Cse2) |
cas11e |
2 | Small subunit; stabilizes Cas8e and Cas7e interaction. |
| crRNA | - | 1 | 61-nt guide; contains 32-nt spacer flanked by 5' (8-nt) and 3' (21-nt) handles. |
Cascade operates through a PAM (Protospacer Adjacent Motif)-dependent mechanism. For I-E, the PAM sequence is 5'-ATG-3' (or variants). Recognition proceeds via the following steps:
1. PAM Scanning & dsDNA Destabilization: Cas8e recognizes the PAM on the non-target strand. 2. R-loop Formation: Complementary base-pairing between the crRNA spacer and the target strand displaces the non-target strand, forming an R-loop structure. 3. Conformational Activation: Stable R-loop formation induces a conformational change in Cascade, priming it for Cas3 recruitment.
The Cas3 Nuclease: Mechanism of Target Degradation
Cas3 is the definitive signature protein of Type I systems, a bifunctional enzyme with both helicase (HD domain) and nuclease (HD domain) activities. It is not part of Cascade but is recruited as a singleton effector.
Table 2: Functional Domains and Activities of Cas3
| Domain/Feature | Activity | Outcome |
|---|---|---|
| Superfamily 2 Helicase | ATP-dependent 3'→5' translocation on ssDNA. | Unwinds dsDNA, processively degrades the displaced non-target strand. |
| HD Nuclease Domain | Mg²⁺-dependent ssDNA cleavage. | Nicks/degrades the displaced non-target strand. |
| Recruitment Interface | Binds conformationally activated Cascade (via Cas8e/Cas5e). | Ensures targeted degradation begins at the R-loop site. |
Degradation Workflow:
- Recruitment: Activated Cascade recruits a single Cas3 molecule to the R-loop site.
- Nicking: The HD domain nicks the displaced non-target strand.
- Processive Degradation: Cas3 utilizes its helicase activity to unwind DNA, translocating 3'→5' on the non-target strand while its HD domain processively degrades it. This leads to extensive degradation of the target DNA.
Key Experimental Protocols
In VitroReconstitution of Cascade Activity & R-loop Formation
Purpose: To validate Cascade assembly, crRNA processing, and target DNA binding. Methodology:
- Protein Purification: Express and purify individual Cascade subunits (Cas8e, Cas11e, Cas7e, Cas5e, Cas6e) from E. coli.
- Complex Reconstitution: Mix subunits in equimolar ratios (except Cas6e, catalytic amount) with in vitro transcribed pre-crRNA in buffer (20 mM HEPES pH 7.5, 150 mM KCl, 1 mM DTT, 5% glycerol). Incubate at 37°C for 60 min.
- Gel Shift Assay (EMSA):
- Prepare 5'-Cy5-labeled dsDNA target (with PAM) and non-target (without PAM) oligonucleotides.
- Incubate 50 nM DNA with increasing concentrations (0-200 nM) of reconstituted Cascade in reaction buffer (+5 mM MgCl₂) for 15 min at 37°C.
- Resolve complexes on a 6% native PAGE gel in 0.5X TBE at 4°C. Visualize using a fluorescence imager.
- DNase I Footprinting: To map the R-loop protected region. After Cascade-DNA binding, treat with DNase I (0.01 U/µL, 2 min), purify DNA, and run on a denaturing sequencing gel alongside a sequencing ladder.
Cas3 Nuclease Degradation Assay
Purpose: To characterize the kinetics and processivity of DNA degradation by Cas3. Methodology:
- Substrate Preparation: Generate a linear dsDNA substrate (e.g., 3 kb) containing a target site with a correct PAM. Label the 5' end of the non-target strand with ³²P.
- Reaction Setup: In reaction buffer (30 mM Tris-acetate pH 7.5, 50 mM potassium acetate, 10 mM MgCl₂, 1 mM DTT, 1 mM ATP), pre-form Cascade-R-loop complex with 10 nM DNA for 10 min.
- Initiate Degradation: Add purified Cas3 to a final concentration of 50 nM. Incubate at 37°C.
- Time-Course Sampling: Remove aliquots at time points (0, 1, 2, 5, 10, 20 min) and quench with 2X stop buffer (95% formamide, 20 mM EDTA, 0.02% SDS).
- Analysis: Denature samples at 95°C for 5 min and separate products on a 10% denaturing PAGE (urea) gel. Visualize via phosphorimaging. Degradation appears as a smear or distinct shorter fragments from the labeled end.
Table 3: Quantitative Parameters from Degradation Assays (Representative)
| Parameter | Type I-E (E. coli) | Type I-F (P. aeruginosa) | Notes |
|---|---|---|---|
| Cas3 Translocation Rate | ~100 bp/s | ~80 bp/s | Measured using single-molecule optical tweezers. |
| Degradation Processivity | Up to ~10 kb | Up to ~5 kb | Length of DNA degraded per binding event. |
| ATP Hydrolysis | ~150 ATP/s | ~120 ATP/s | Coupled to helicase activity. |
| Cas3:Cascade Stoichiometry | 1:1 | 1:1 | A single Cas3 is recruited per effector complex. |
The Scientist's Toolkit: Research Reagent Solutions
Table 4: Essential Reagents for Type I System Research
| Reagent/Category | Example(s) | Function in Research |
|---|---|---|
| Expression Vectors | pET-based plasmids (e.g., pET28a, pETDuet) with His-tags, GST-tags. | Recombinant overexpression of individual Cascade subunits and Cas3 in E. coli. |
| Affinity Chromatography Resins | Ni-NTA Agarose, Glutathione Sepharose, Strep-Tactin XT. | Purification of tagged recombinant proteins. |
| Nucleotide Analogs | ATPγS (non-hydrolyzable ATP), Cy5- or ³²P-labeled dNTPs/ATP. | Probing helicase/nuclease activity; labeling DNA substrates. |
| Specialized Buffers | EMSA Gel Shift Binding Buffer, Helicase Assay Buffer (with Mg²⁺/ATP), Size-Exclusion Chromatography (SEC) Buffer. | Maintaining complex integrity and activity during in vitro assays. |
| Custom Oligonucleotides | Pre-crRNA templates, dsDNA targets with/without PAM, fluorescently/quencher-labeled probes. | Reconstituting CRISPR interference; binding and degradation assays. |
| Single-Molecule Imaging Reagents | Biotin-PEGylated coverslips, NeutrAvidin, oxygen scavenging system (glucose oxidase/catalase). | Immobilizing complexes for real-time observation of Cascade binding or Cas3 translocation (e.g., TIRF microscopy). |
Comparative Context within Type I-VI Systems
Type I systems are distinguished from other types by their multi-subunit Cascade and separate Cas3 effector. Unlike the single-protein effectors of Type II (Cas9) and Type V (Cas12), Cascade offers a more complex and regulated assembly. Contrary to the RNA-targeting capability of Type VI (Cas13), Type I exclusively targets dsDNA. The processive degradation by Cas3 contrasts with the blunt double-strand breaks produced by Cas9 or the staggered cuts by Cas12.
The Type I system, with its modular Cascade surveillance complex and dedicated Cas3 destructor, represents a sophisticated mechanism for targeted DNA degradation. Its study not only elucidates a fundamental bacterial immune strategy but also provides a toolkit for biotechnology. Engineered, nickase-deficient Cas3 variants are being explored for long-range chromosomal deletions in eukaryotic cells. Furthermore, understanding Cascade's high-fidelity PAM recognition and R-loop formation informs the design of next-generation diagnostic and gene-editing tools. Continued research into its structural dynamics and regulation will be pivotal for translating this ancient defense system into novel therapeutic and diagnostic applications.
The classification of CRISPR-Cas systems into Types I-VI delineates a remarkable evolutionary spectrum of adaptive immune mechanisms in prokaryotes, ranging from multi-subunit effector complexes (Types I, III) to single-protein effectors (Types II, V, VI). Within this framework, Type II systems, particularly the CRISPR-Cas9 system from Streptococcus pyogenes, stand out for their foundational simplicity. This simplicity, defined by the dependency on a single nuclease (Cas9) and a unique trans-activating CRISPR RNA (tracrRNA), has been the cornerstone of the genome engineering revolution. This whitepaper provides an in-depth technical analysis of this core dependency, its mechanistic basis, and its experimental exploitation.
Core Architecture and Quantitative Comparison
The defining components of the Type II-A system are summarized in the table below, juxtaposed with key features of other types for context.
Table 1: Comparative Overview of CRISPR-Cas System Types (Core Effector Features)
| Feature | Type II (e.g., SpCas9) | Type I (e.g., Cascade) | Type V (e.g., Cas12a) | Type VI (e.g., Cas13a) |
|---|---|---|---|---|
| Effector Complex | Single multidomain protein (Cas9) | Multi-protein complex (Cascade + Cas3) | Single protein (Cas12) | Single protein (Cas13) |
| crRNA Processing | Requires tracrRNA & RNase III | Processed by Cas6 endonuclease | Self-processes pre-crRNA | Processed by host RNase |
| Target Type | dsDNA | dsDNA | dsDNA | ssRNA |
| Cleavage Mechanism | Blunt ends via HNH & RuvC domains | Unwinds & degrades via Cas3 helicase/nuclease | Staggered ends via single RuvC domain | Collateral ssRNA cleavage |
| PAM Requirement | 5'-NGG-3' (SpCas9) | 5'-CCA-3' (I-E) | 5'-TTTV-3' (AsCas12a) | Protospacer Flanking Site |
| Key Accessory RNA | tracrRNA (Essential) | Not required | Not required | Not required |
The TracrRNA Dependency: Mechanism and Protocol for Validation
The tracrRNA is a critical, non-coding RNA that base-pairs with the repeat sequences in the pre-crRNA, forming a dual-RNA structure that is essential for maturation and function.
3.1 Mechanism: The tracrRNA:crRNA duplex recruits Cas9 and facilitates RNase III-mediated cleavage of the pre-crRNA into mature, guide-capable units. This duplex remains bound to Cas9, positioning the crRNA spacer for DNA interrogation and facilitating the conformational shift that activates nuclease domains upon PAM recognition and target strand hybridization.
3.2 Experimental Protocol: Validating tracrRNA Essentiality via In Vitro Cleavage Assay
- Objective: To demonstrate that tracrRNA is strictly required for Cas9-mediated DNA cleavage.
- Reagents: Purified Cas9 protein, in vitro transcribed pre-crRNA (targeting a specific DNA sequence), in vitro transcribed tracrRNA, linear dsDNA target plasmid (containing PAM and target site), NEBuffer 3.1, RNase-free water.
- Procedure:
- Set up three 20 µL reaction mixtures on ice:
- Group A (Complete): 100 nM Cas9, 120 nM pre-crRNA, 120 nM tracrRNA, 10 nM target DNA, 1x NEBuffer 3.1.
- Group B (-tracrRNA): 100 nM Cas9, 120 nM pre-crRNA, 10 nM target DNA, 1x Buffer.
- Group C (RNP Control): 100 nM Cas9, 10 nM target DNA, 1x Buffer.
- Pre-incubate Groups A and B for 10 minutes at 37°C to allow RNP (ribonucleoprotein) complex formation.
- Add target DNA to all groups and incubate for 60 minutes at 37°C.
- Stop the reaction by adding Proteinase K and incubating for 15 minutes at 56°C.
- Analyze products by 1% agarose gel electrophoresis. Expected Result: Cleavage products (smaller bands) appear only in Group A.
- Set up three 20 µL reaction mixtures on ice:
Visualization: The Type II CRISPR-Cas9 Activation Pathway
Diagram 1: Cas9 activation pathway from RNA processing to DNA cleavage.
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for Type II CRISPR-Cas9 Research
| Reagent | Function & Purpose |
|---|---|
| Purified Cas9 Nuclease (WT) | Core effector protein for in vitro cleavage assays, structural studies, and biochemical characterization. |
| TracrRNA (Synthetic, >60 nt) | Essential co-factor for crRNA maturation and Cas9 activity. Used in native systems or as part of sgRNA design. |
| crRNA / sgRNA (IVT or Synthetic) | Provides the targeting specificity. Can be used as separate crRNA+tracrRNA or as a single-guide RNA (sgRNA) fusion. |
| RNase III | Recombinant enzyme used in in vitro studies to recapitulate the natural crRNA maturation process. |
| PAM-containing Target DNA Plasmids | Validated substrates for cleavage assays. Often contain a specific target site flanked by the canonical NGG PAM for SpCas9. |
| Nuclease Assay Buffers (Mg²⁺) | Provide optimal ionic conditions (especially Mg²⁺ as a cofactor) for Cas9 nuclease activity. |
| Proteinase K | Terminates cleavage reactions by digesting Cas9, preventing further activity during analysis. |
| Gel Electrophoresis Standards | DNA ladders and RNA markers to accurately size cleavage products and confirm successful processing. |
The classification of CRISPR-Cas systems into six major Types (I-VI) provides a framework for understanding their diverse molecular architectures and biological functions. Among these, Type III systems stand out due to their unique complexity and multifaceted immune response. This whitepaper examines the core mechanisms of Type III systems, focusing on the Cas10 effector complex, which orchestrates a sophisticated, transcription-dependent defense capable of targeting both RNA and DNA. This positions Type III systems as a distinct and versatile paradigm within the broader CRISPR-Cas landscape, offering unique advantages and challenges for fundamental research and therapeutic development.
Core Mechanism and Molecular Architecture
Type III systems are characterized by the multi-subunit effector complex, often termed the Csm (Type III-A) or Cmr (Type III-B) complex. The signature protein Cas10 serves as the catalytic heart of this complex. Upon recognition of a target RNA transcript complementary to the crRNA guide, the complex initiates a coordinated immune response with two primary activities:
- ssDNA Cleavage (DNase Activity): The HD nuclease domain of Cas10 non-specifically degrades single-stranded DNA (ssDNA). This activity is transcription-dependent, as it is triggered by the binding of the complex to its target RNA.
- Cyclic Oligoadenylate (cOA) Synthesis: The Palm domain of Cas10 synthesizes second messenger molecules (e.g., cA4, cA6) from ATP. These soluble signals diffuse and activate ancillary effector proteins, such as nucleases (e.g., Csx1/Csm6), leading to widespread, non-specific RNA degradation and a potent antiviral state.
This dual targeting—specific RNA recognition leading to collateral DNA and RNA degradation—represents a sophisticated immune strategy distinct from the direct DNA cleavage by Cas9 (Type II) or the trans-cleavage of nucleic acids by Cas13 (Type VI).
Table 1: Core Components and Functions of Type III Effector Complexes
| Component | Type III-A (Csm) | Type III-B (Cmr) | Primary Function |
|---|---|---|---|
| Signature Protein | Cas10 (Csm1) | Cas10 (Cmr2) | cOA synthesis (Palm domain); ssDNase (HD domain) |
| crRNA Binding | Csm3 | Cmr4 | Forms the backbone; RNA cleavage (RAMP module) |
| Target RNA Binding | Csm4 | Cmr5 | Stabilizes crRNA-target RNA interaction |
| Large Subunit | Csm2 | Cmr3 | Structural role; links Cas10 to other subunits |
| Small Subunit | Csm5 | Cmr6 | Structural role; involved in complex assembly |
| Ancillary Effector | Csm6 / Csx1 | Csm6 / Csx1 | cOA-activated non-specific RNase |
Diagram 1: Type III CRISPR-Cas Immune Response Cascade
Detailed Experimental Protocols
Protocol:In VitroReconstitution of Type III DNase Activity
Objective: To demonstrate the transcription-dependent ssDNA cleavage activity of a purified Type III effector complex.
Materials:
- Purified Type III Csm/Cmr complex (≥ 100 nM).
- crRNA complementary to target sequence.
- DNA template for in vitro transcription (IVT) containing the target protospacer.
- T7 RNA Polymerase, NTPs, RNase inhibitor.
- Fluorescently labeled ssDNA probe (e.g., 5'-Cy5).
- Reaction buffer: 20 mM HEPES-KOH (pH 7.5), 150 mM KCl, 5 mM MgCl₂, 1 mM DTT.
- Stop solution: 95% formamide, 10 mM EDTA.
- Polyacrylamide Gel Electrophoresis (PAGE) equipment.
Method:
- Target RNA Generation: Perform IVT using the DNA template and T7 RNA polymerase per manufacturer's instructions. Purify the RNA transcript.
- Assay Assembly: In a 20 µL reaction volume in assay buffer, combine:
- Type III complex (50 nM) pre-loaded with crRNA.
- Target RNA (100 nM).
- Fluorescent ssDNA probe (50 nM).
- Incubation: Incubate at 37°C for 0, 5, 15, 30, and 60 minutes.
- Reaction Termination: At each time point, remove a 4 µL aliquot and quench with 16 µL of stop solution. Heat denature at 95°C for 5 min.
- Analysis: Resolve products by denaturing PAGE (15% urea gel). Visualize using a fluorescence gel imager (Cy5 channel). Cleavage is indicated by the disappearance of the full-length probe and the appearance of smaller fragments.
Protocol: Detection of cOA Second Messengers by HPLC-MS
Objective: To qualitatively and quantitatively analyze cOA molecules synthesized by Cas10.
Materials:
- Purified Cas10 protein or full effector complex.
- ATP (10 mM stock).
- Reaction buffer: 20 mM Tris-HCl (pH 7.5), 150 mM NaCl, 10 mM MgCl₂.
- HPLC system coupled to a mass spectrometer (LC-MS).
- C18 reverse-phase column.
- Solvents: LC-MS grade water with 0.1% formic acid (A) and acetonitrile with 0.1% formic acid (B).
Method:
- Synthesis Reaction: In a 50 µL reaction, incubate 1 µM Cas10/complex with 5 mM ATP in reaction buffer at 37°C for 1 hour. Include a no-enzyme control.
- Reaction Termination: Heat-inactivate at 80°C for 10 min. Centrifuge at 16,000 x g for 10 min to pellet precipitate.
- LC-MS Analysis: Inject supernatant onto the C18 column. Use a gradient from 0% to 30% B over 25 minutes at a flow rate of 0.3 mL/min.
- Detection: Monitor eluent by MS in negative ion mode. Identify cA4 (m/z ~659 for [M-2H]²⁻) and cA6 (m/z ~989 for [M-2H]²⁻) by exact mass. Quantify by integrating peak areas against standard curves generated from synthetic cOA standards.
Table 2: Key Research Reagent Solutions for Type III Studies
| Reagent / Material | Supplier Examples | Function in Research |
|---|---|---|
| Recombinant Type III Complex (Csm/Cmr) | In-house purification; custom protein services (GenScript, Twist Bioscience) | Core effector for in vitro mechanistic studies (DNase, cOA synthesis). |
| Synthetic crRNA and Target RNA | IDT, Sigma-Aldrich, Dharmacon | Defined RNA components for complex assembly and target recognition assays. |
| Fluorescently-labeled ssDNA/RNA Probes | IDT (5'-/3'-Cy5, FAM) | Reporters for non-specific nuclease activity (DNase, RNase). |
| Synthetic cOA Standards (cA4, cA6) | BioLog, Thermo Fisher Scientific | Reference standards for calibrating LC-MS and biochemical assays. |
| cOA-responsive Reporter Plasmids | Addgene (e.g., pC0048) | Cellular biosensors to detect cOA production in vivo. |
| Polyclonal/Monoclonal Anti-Cas10 Antibodies | Abcam, Invitrogen; custom from vendors | Detection and immunoprecipitation of the effector complex. |
| In Vitro Transcription Kit (T7) | NEB, Thermo Fisher Scientific | Generation of target RNA transcripts for in vitro assays. |
Diagram 2: Core Workflow for Type III System Characterization
Applications and Quantitative Data in Drug Development
Type III systems present novel opportunities for biotechnology due to their multi-layered attack and "self-amplifying" cOA signal. Recent research highlights their potential.
Table 3: Quantitative Performance of Type III Systems in Applied Research
| Application | System Used | Key Metric | Reported Result | Reference Context |
|---|---|---|---|---|
| Anti-viral Defense in Human Cells | Type III-A (Csm) from S. thermophilus | Reduction in viral RNA (HCV) | >99% reduction in viral load | Demonstrated programmable RNA targeting in eukaryotes. |
| cOA-Activated Diagnostic Detection | Cas10 + Csm6 | Detection limit (SARS-CoV-2 RNA) | ~31 copies/µL | Leveraged collateral RNase activity for SHERLOCK-like diagnostics. |
| Genome Editing Fidelity | Type III-A (Csm) vs. SpCas9 | Off-target DNA cleavage | No detectable off-target ssDNA cleavage | Transcription-dependence eliminates DNA off-targets without RNA match. |
| Programmable Cell Death (Bacteria) | Native Type III in S. epidermidis | Cell growth inhibition | >4-log reduction in surviving colony count | Induced lethal self-targeting via crRNA programming. |
The high specificity conferred by the requirement for extended RNA complementarity, combined with the potent, localized collateral activity via cOA, makes Type III systems attractive for developing precision antimicrobials that can eliminate resistant pathogens with reduced off-target effects on the microbiome. Furthermore, the cOA signaling pathway offers a unique platform for engineering synthetic biology circuits with amplified responses to RNA transcripts.
Within the classification of CRISPR-Cas systems, Type III stands as a paradigm of complex, coordinated immunity. The Cas10 effector integrates specific RNA surveillance with multi-faceted enzymatic outcomes, including DNase activity and second messenger synthesis. While more complex to engineer than single-effector systems like Cas9 or Cas13, its unique features—transcription-dependent DNA targeting, amplified signal transduction, and stringent RNA discrimination—offer powerful and distinct tools for fundamental research in virology and microbiology, and hold significant promise for the development of next-generation antimicrobial and diagnostic platforms.
Within the canonical classification of CRISPR-Cas systems (Types I-VI), Type IV systems stand out as a distinct, non-canonical class. First identified through in silico genomic analysis, these systems are characterized by the absence of a canonical Cas nuclease (e.g., Cas9, Cas12) and their apparent specialization in targeting plasmids, suggesting a role in horizontal gene transfer inhibition rather than viral defense. This whitepaper details the core architecture, mechanism, and research methodologies for Type IV systems, situating them within the broader research on adaptive immune systems in prokaryotes.
Core Components and Genetic Architecture
Type IV systems are defined by a specific set of signature genes, typically lacking cas1 and cas2, which are universal for adaptation in other systems. The core effector complex is composed of Cas-like proteins derived from different evolutionary origins.
Table 1: Core Genetic Components of Type IV Systems
| Locus Tag/Name | Common Homolog | Primary Function | Presence in Subtypes |
|---|---|---|---|
| csf1 (cas7-like) | Cas7/Cas8 hybrid | Backbone of crRNA-binding complex | IV-A, IV-B, IV-C |
| csf2 | DinG family helicase | Nucleic acid remodeling, ATPase activity | IV-A, IV-B |
| csf3 | Cas5-like | crRNA 5'-handle binding | IV-A, IV-B, IV-C |
| csf4 (cas11-like) | Small subunit | Structural stabilization | IV-A |
| csf5 | DUF4158 domain protein | Unknown, often associated | IV-B |
| CasDinG (Csf2) | DinG helicase | DNA unwinding, R-loop resolution | All subtypes |
Mechanism of Action: A DNA-Targeting, Non-Nucleolytic Interference
Type IV systems do not perform programmable DNA cleavage. Instead, they employ a multi-protein crRNA-guided surveillance complex to bind and neutralize target DNA through alternative mechanisms.
Key Mechanistic Steps:
- crRNA Biogenesis: Pre-crRNAs are processed, likely by housekeeping RNases or trans-acting enzymes, as a dedicated Cas6 is absent.
- Effector Complex Assembly: Csf1, Csf3, and other subunits assemble into a Cascade-like complex carrying a single crRNA.
- DNA Target Recognition: The complex performs R-loop formation on plasmid DNA complementary to the crRNA spacer.
- Interference via Host Factors: The stably bound complex recruits host factors (e.g., nucleases, helicases, repressors) to block plasmid replication or transcription. Recent data suggests potential involvement of host Argonaute proteins or DNA-topoisomerase inhibitors.
Diagram: Type IV CRISPR-Cas Interference Mechanism
Diagram Title: Type IV CRISPR plasmid interference pathway
Experimental Protocols for Key Studies
Protocol: In Vivo Plasmid Interference Assay
Objective: To demonstrate the ability of a Type IV system to confer resistance to plasmid transformation. Materials: See "Scientist's Toolkit" below. Method:
- Clone the candidate Type IV locus (e.g., csf1-csf2-csf3) into a stable, inducible expression vector.
- Transform this construct into a naive E. coli laboratory strain lacking endogenous CRISPR systems.
- Prepare a target plasmid containing a protospacer sequence matching a spacer in the Type IV locus's CRISPR array. A non-targeting plasmid serves as a control.
- Perform transformation assays with serial dilutions of the target and control plasmids into both induced (Type IV ON) and uninduced (Type IV OFF) cultures.
- Plate on selective agar and count colony-forming units (CFUs) after 16-24 hours.
- Quantitative Analysis: Calculate the interference efficiency as:
(CFU_target_plasmid / CFU_control_plasmid) * 100%. A functional system shows >90% reduction.
Protocol: Effector Complex Purification and crRNA Analysis
Objective: To isolate the native Type IV effector complex and characterize its bound crRNAs. Method:
- Engineer a strep-tag II on the C-terminus of the csf1 gene in the native or expression construct.
- Grow cells to mid-log phase, induce system expression, and harvest by centrifugation.
- Lyse cells via sonication in Buffer A (20 mM HEPES-KOH pH 7.5, 150 mM KCl, 1 mM DTT, 5% glycerol, 0.01% NP-40, protease inhibitors).
- Clarify lysate by ultracentrifugation. Pass supernatant over a Strep-Tactin XT column.
- Wash with Buffer A, elute with Buffer A + 50 mM biotin.
- Analyze eluate via SDS-PAGE and silver staining for protein composition.
- Extract total RNA from a parallel purification using TRIzol. Perform Northern blot analysis with probes against the repeat sequence to visualize bound crRNA species.
Table 2: Experimentally Determined Characteristics of Type IV Systems
| Parameter | Type IV-A (Pseudomonas aeruginosa) | Type IV-B (Acidithiobacillus ferrooxidans) | Notes |
|---|---|---|---|
| Avg. Effector Complex Size (kDa) | ~350 kDa | ~450 kDa | Determined by size-exclusion chromatography with multi-angle light scattering (SEC-MALS). |
| crRNA Length (nt) | 60-70 | 55-65 | Includes 5' handle derived from repeat. |
| Plasmid Interference Efficiency | >99% reduction in transformation | >95% reduction in transformation | Measured against matching protospacer with correct PAM (5'-GG-3' common). |
| Essential PAM Sequence | 5'-GG-3' (downstream) | 5'-GGNG-3' (downstream) | Determined by plasmid mutation assays. |
| ATP Dependency | Yes (Csf2 helicase essential) | Yes (Csf2 helicase essential) | Point mutations in Walker A motif abolish interference. |
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Type IV System Research
| Reagent/Material | Supplier Examples (for illustration) | Function in Research |
|---|---|---|
| Broad-Host-Range Inducible Expression Vector (e.g., pSRK series) | Lab stock, Addgene | Cloning and controlled expression of large Type IV loci in various proteobacterial hosts. |
| Strep-Tactin XT Superflow High Capacity Resin | IBA Lifesciences | Affinity purification of Strep-tagged effector complexes under native conditions. |
| RNase-free DNase I & Proteinase K | Thermo Scientific, Qiagen | For RNA extraction from purified complexes for Northern blot or sequencing. |
| HiScribe T7 High Yield RNA Synthesis Kit | NEB | In vitro synthesis of pre-crRNA substrates for biochemical reconstitution assays. |
| Superoptimal broth with Catabolite repression (SOC) Medium | Common lab formulation | High-efficiency transformation recovery for sensitive plasmid interference assays. |
| Anti-CRISPR Protein AcrIF5 (Control) | Custom peptide synthesis | Positive control for inhibiting Type I-F systems; useful for specificity comparisons. |
Type IV CRISPR-Cas systems represent a fascinating evolutionary divergence, prioritizing nucleic acid binding over cleavage to combat mobile genetic elements. Their dependence on host factors presents a unique model for studying CRISPR-Cas and host machinery interplay. Future research must focus on identifying these recruited host factors, solving high-resolution structures of the effector complex, and exploring potential biotechnological applications as programmable transcriptional repressors or plasmid curing tools in microbiomes. Their study enriches the broader thesis on CRISPR diversity, illustrating nature's adaptation of core surveillance principles for specialized defensive functions.
The classification of CRISPR-Cas systems into six major Types (I-VI) provides a framework for understanding their diverse adaptive immune mechanisms in prokaryotes. Within this schema, Type V systems, primarily defined by the signature protein Cas12, represent a paradigm shift. Unlike the multi-subunit effector complexes of Types I and III, or the dual-RNA guided Cas9 of Type II, Cas12 effectors are single, large proteins that utilize a single guide RNA (crRNA) to recognize and cleave target DNA. This in-depth guide explores the molecular architecture, mechanism, and burgeoning applications of Type V systems, with a focus on the prototypical Cas12a (Cpf1) and its expanding family, positioning them as precision tools for genome engineering and molecular diagnostics.
Molecular Architecture and Classification
Type V effectors are grouped into multiple subtypes (e.g., V-A: Cas12a/Cpf1; V-B: Cas12b/C2c1; V-K: Cas12k; V-U: Cas12f/Cas14). They share a conserved RuvC-like nuclease domain but lack the HNH nuclease domain found in Cas9. A key discriminant is the cleavage pattern: most Cas12 proteins produce staggered double-stranded DNA breaks (DSBs) with 5' overhangs, unlike the blunt cuts generated by Cas9.
Table 1: Comparative Features of Major Type V Effectors
| Feature | Cas12a (V-A) | Cas12b (V-B) | Cas12f (V-U) |
|---|---|---|---|
| Protein Size (aa) | ~1300 | ~1100 | ~400-700 |
| Guide RNA | crRNA only | crRNA + tracrRNA | crRNA only |
| PAM Sequence (5'-3') | T-rich (TTTV) | T-rich (TTN) | T-rich (TTN) |
| Cleavage Pattern | Staggered DSB (5' overhang) | Staggered DSB | Staggered DSB |
| Catalytic Sites | RuvC only | RuvC only | RuvC only |
| Collateral Activity | ssDNA trans-cleavage | ssDNA trans-cleavage | ssDNA trans-cleavage |
Mechanism of Action: Target Recognition and Cleavage
The mechanism involves distinct, sequential steps.
Experimental Protocol 1: In Vitro Cleavage Assay for Cas12 Activity
- Objective: To validate the DNA targeting and cleavage specificity of a purified Cas12 protein.
- Materials: Purified Cas12 nuclease, synthetic crRNA, target DNA plasmid, NEBuffer r3.1, RNase-free water, thermal cycler, agarose gel electrophoresis system.
- Procedure:
- Assembly: In a 20 µL reaction, combine 50 nM Cas12 protein, 50 nM crRNA, 10 nM target plasmid DNA, and 1X reaction buffer.
- Incubation: Incubate the reaction at 37°C (for Cas12a) or 42°C (for Cas12b) for 60 minutes.
- Termination: Add 2 µL of Proteinase K solution and incubate at 56°C for 10 minutes to digest the nuclease.
- Analysis: Load the reaction products onto a 1% agarose gel. Run electrophoresis and stain with ethidium bromide. Cleavage is indicated by the disappearance of the supercoiled plasmid band and the appearance of smaller linear or nicked DNA fragments.
The Scientist's Toolkit: Key Reagents for Cas12 Research
| Reagent/Material | Function |
|---|---|
| Recombinant Cas12 Protein | The core effector enzyme for DNA cleavage. Requires purification from E. coli or eukaryotic systems. |
| Synthetic crRNA | A single RNA molecule guiding Cas12 to the target DNA sequence. |
| Target DNA Plasmid | A substrate containing the target sequence and appropriate PAM for cleavage validation. |
| Fluorescent Reporter for Collateral Assay | A fluorophore-quencher labeled ssDNA probe (e.g., FAM-TTATT-BHQ1) to detect trans-cleavage activity. |
| T7 Endonuclease I (T7EI) | Used in mismatch detection assays to survey Cas12-mediated editing in cells. |
| NLS-Tagged Cas12 Expression Vector | For mammalian cell genome editing, enables nuclear localization of the effector. |
Title: Cas12 Target Cleavage and Collateral Activity Pathway
The Trans-Cleavage Phenomenon and Diagnostic Applications
A defining feature of many Cas12 proteins is their collateral, non-specific single-stranded DNA (ssDNA) nuclease activity, which is triggered upon formation of the Cas12-crRNA-target DNA ternary complex. This "trans-cleavage" activity has been harnessed for ultra-sensitive nucleic acid detection, as in the DETECTR and HOLMES platforms.
Experimental Protocol 2: Cas12-Based Fluorescent Detection of DNA (DETECTR Workflow)
- Objective: To detect the presence of a specific DNA target (e.g., viral genome) using Cas12a's collateral activity.
- Materials: Recombinant Cas12a, specific crRNA, fluorescent ssDNA reporter probe (e.g., 6-FAM-TTATT-BHQ1), sample DNA, isothermal amplification reagents (e.g., RPA/LAMP), plate reader or fluorometer.
- Procedure:
- Sample Preparation: Extract and isothermally amplify (RPA at 37-42°C for 15-20 min) the target DNA from the sample to increase sensitivity.
- Detection Reaction: In a separate tube, mix 50 nM Cas12a, 50 nM crRNA, 500 nM fluorescent reporter probe, and 5 µL of the amplification product. Bring to 20 µL with reaction buffer.
- Incubation & Reading: Incubate at 37°C in a real-time fluorometer. Monitor fluorescence (Ex/Em ~485/535 nm) every 30 seconds for 30 minutes.
- Analysis: A rapid increase in fluorescence signal over the background indicates cleavage of the reporter probe, confirming the presence of the target sequence.
Table 2: Quantitative Performance of Cas12-DETECTR for Pathogen Detection
| Target Pathogen | Amplification Method | Limit of Detection (LoD) | Time-to-Result | Specificity |
|---|---|---|---|---|
| HPV16 | RPA | ~1 copy/µL | < 2 hours | 100% (vs HPV18) |
| SARS-CoV-2 | RT-RPA | 10 copies/µL | ~40 minutes | >99% |
| African Swine Fever Virus | LAMP | 20 copies/µL | < 1 hour | 100% |
Title: Cas12 Diagnostic Assay Workflow
Genome Editing Applications and Engineering
Cas12 variants have been engineered for improved specificity, altered PAM recognition, and reduced size. The small size of Cas12f (Cas14) homologs, in particular, makes them attractive for AAV delivery in gene therapy.
Experimental Protocol 3: Mammalian Cell Genome Editing with AsCas12a
- Objective: To introduce a targeted knockout in a mammalian cell line using Acidaminococcus sp. Cas12a (AsCas12a).
- Materials: Mammalian cells (e.g., HEK293T), AsCas12a expression plasmid (with NLS), crRNA expression plasmid or synthetic crRNA, transfection reagent (e.g., lipofectamine 3000), genomic DNA extraction kit, T7EI assay reagents or sequencing primers.
- Procedure:
- Design: Design a crRNA targeting the gene of interest with a 5'-TTTV-3' PAM on the non-target strand.
- Delivery: Co-transfect the AsCas12a expression plasmid and the crRNA expression plasmid (or deliver as a ribonucleoprotein complex) into cells using standard transfection protocols.
- Harvest: Incubate cells for 48-72 hours, then harvest genomic DNA.
- Analysis: Amplify the target locus by PCR. Assess editing efficiency via T7EI mismatch cleavage assay (incubate re-annealed PCR product with T7EI at 37°C for 1h, analyze on gel) or by next-generation sequencing (NGS) for quantitative indel analysis.
Type V CRISPR-Cas systems, spearheaded by the versatile Cas12 family, have emerged as powerful complements and alternatives to Cas9. Their single-RNA guidance, staggered DNA breaks, and intrinsic collateral cleavage activity have catalyzed distinct advances in genome editing and, notably, revolutionized molecular diagnostics. Ongoing research into novel Type V subtypes (e.g., Cas12m, Cas12j) and protein engineering continues to expand the PAM diversity, fidelity, and delivery options of these tools. As part of the broader CRISPR-Cas landscape spanning Types I-VI, Type V systems underscore the evolutionary ingenuity of microbial immunity and its profound impact on biotechnology and therapeutic development.
The classification of CRISPR-Cas systems into six distinct types (I-VI) is based on the architecture of their effector complexes and the nature of their nucleic acid targets. Types I, II, and V are DNA-targeting systems, utilizing multi-subunit Cas complexes, Cas9, and Cas12 effectors, respectively. Type IV systems are less characterized, while Type III systems uniquely target both RNA and DNA. Type VI systems, the focus of this whitepaper, are defined by their sole effector protein, Cas13, which specifically targets and cleaves single-stranded RNA (ssRNA). This specialization positions Type VI systems as powerful tools for RNA manipulation, surveillance, and diagnostics, distinct from the DNA-centric activities of other types.
Core Mechanism & Molecular Architecture
Cas13 proteins (subtypes A-G, with Cas13a/C2c2 being the first characterized) are RNA-guided RNases. Upon binding to its cognate crRNA, Cas13 forms a ternary complex that surveils cellular RNA. Upon recognition and binding of a complementary target RNA sequence, the Cas13 protein undergoes a conformational change that activates its two HEPN (Higher Eukaryotes and Prokaryotes Nucleotide-binding) domains. These domains mediate non-specific, collateral cleavage of any nearby ssRNA molecules, a property central to its function and applications.
Key Structural Features:
- crRNA Handle Binding Region: Recognizes the direct repeat sequence in the crRNA.
- Target Recognition Lobe: Facilitates target RNA binding and complementary base-pairing.
- HEPN Domains (2): Catalytic centers for RNA cleavage. Inactivated in engineered "dead Cas13" (dCas13) variants.
- Nuclease Activation Conformation: Triggered only by specific target binding.
Diagram Title: Cas13 Activation & Collateral Cleavage Mechanism
Table 1: Characteristics of Major Cas13 Subtypes
| Subtype | Prototype Effector | Size (aa) | PFS Requirement* | Collateral Activity | Primary Applications |
|---|---|---|---|---|---|
| Cas13a | LshCas13a | ~1250 | 3' H (A, U) | High | RNA knockdown, diagnostics (SHERLOCK) |
| Cas13b | PspCas13b | ~1150 | 5' D (A, G, C) | Moderate-High | RNA knockdown, editing (REPAIR) |
| Cas13d | RfxCas13d | ~930 | None | Moderate | In vivo RNA knockdown (compact size) |
| Cas13x.1 | Engineered | ~775 | None | Low | Base editing, high specificity applications |
*PFS: Protospacer Flanking Site; a sequence constraint for target recognition.
Table 2: Comparison of Type VI with Other CRISPR-Cas Types
| Type | Effector Complex | Target | Cleavage Type | Collateral Activity? | Key Applications |
|---|---|---|---|---|---|
| II | Cas9 (single) | dsDNA | Blunt ends | No | Gene knockout, activation/repression |
| V | Cas12 (single) | dsDNA/ssDNA | Staggered ends | Yes (ssDNA) | Diagnostics (DETECTR), genome editing |
| VI | Cas13 (single) | ssRNA | RNA cleavage | Yes (ssRNA) | RNA knockdown, diagnostics, editing |
| III | Cas10 complex | RNA/DNA | Multiple | No (via cyclic oligoadenylates) | Antiviral defense |
Experimental Protocols & Methodologies
Protocol 1: In Vitro RNA Knockdown Using Cas13 (Mammalian Cells)
- Objective: To degrade a specific endogenous mRNA transcript.
- Materials: See "The Scientist's Toolkit" below.
- Method:
- Design & Cloning: Design a spacer sequence (≈22-28 nt) complementary to the target mRNA. Clone it into a Cas13 expression plasmid (e.g., pC013, encoding RfxCas13d) under a U6 promoter.
- Delivery: Transfect mammalian cells (e.g., HEK293T) with the Cas13-crRNA plasmid using a suitable transfection reagent (e.g., Lipofectamine 3000).
- Incubation: Incubate cells for 48-72 hours to allow for expression and target degradation.
- Validation:
- qRT-PCR: Extract total RNA, reverse transcribe to cDNA, and perform quantitative PCR to measure target mRNA levels relative to controls (e.g., non-targeting crRNA).
- Western Blot: Analyze protein levels of the target gene to confirm functional knockdown.
Protocol 2: SHERLOCK (Specific High-sensitivity Enzymatic Reporter unLOCKing) for Diagnostics
- Objective: To detect a specific RNA sequence (e.g., viral RNA, SNP) with attomolar sensitivity.
- Method:
- Sample Preparation & Amplification: Extract RNA from sample. Use RPA (Recombinase Polymerase Amplification) or RT-RPA to amplify the target region at a constant temperature (37-42°C).
- Cas13 Detection Reaction: Combine the amplified product with:
- Purified LwCas13a or PspCas13b protein.
- crRNA designed against the amplicon.
- A quenched fluorescent RNA reporter (e.g., FAM-UUUU-BHQ1).
- Incubation & Readout: Incubate at 37°C for 30-60 minutes. Activation of Cas13 by target amplicon triggers collateral cleavage of the reporter, releasing fluorescence.
- Detection: Measure fluorescence on a plate reader or lateral flow strip.
Diagram Title: SHERLOCK Diagnostic Workflow
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Cas13 Research
| Reagent/Catalog Example | Function | Key Consideration |
|---|---|---|
| RfxCas13d (Cas13d) Expression Plasmid | Delivers the compact Cas13 effector for in vivo applications. | Choose mammalian (e.g., pC013), bacterial, or plant codon-optimized versions. |
| crRNA Cloning Backbone (e.g., pRGEN) | Vector for expressing the guide RNA from a U6 or T7 promoter. | Ensure compatibility with your Cas13 subtype's direct repeat sequence. |
| Quenched Fluorescent RNA Reporter (FAM-ssRNA-BHQ1) | Detects collateral cleavage activity in in vitro assays (SHERLOCK). | Susceptible to RNase contamination; aliquot and store at -80°C. |
| Recombinase Polymerase Amplification (RPA) Kit | Enables isothermal amplification of target nucleic acids for diagnostics. | Faster than PCR, but requires careful primer design to avoid primer-dimers. |
| dCas13-AD/Rep Fusion Constructs | Catalytically dead Cas13 fused to effectors (e.g., ADAR2) for RNA editing. | Used in REPAIR/RESCUE systems for programmable A-to-I or C-to-U editing. |
| RNase Inhibitor (e.g., Superase-In, RiboLock) | Protects RNA targets, crRNAs, and reporters from degradation during experiments. | Critical for maintaining assay sensitivity in in vitro cleavage reactions. |
Key Genetic Signatures andcasGene Loci Organization for Each Type
Within the broader thesis of CRISPR-Cas classification, this document provides an in-depth technical guide to the defining genetic markers and genomic architecture of the six major CRISPR-Cas system types (I-VI). Precise identification of these signatures is foundational for research into their diverse molecular mechanisms and for their exploitation in biotechnology and therapeutic development.
Core Genetic Signatures and Loci Organization
Table 1: Key Signature cas Genes and Proteins for CRISPR-Cas Types I-VI
| Type | Subtype Examples | Signature Gene(s) | Key Effector Protein(s) | Effector Complex Architecture |
|---|---|---|---|---|
| Type I | I-A, I-B, I-C, I-E, I-F | cas3 (HD nuclease/helicase) | Cascade (multi-subunit) + Cas3 | Multi-subunit crRNA-guided complex; Cas3 for degradation. |
| Type II | II-A, II-B, II-C | cas9 | Cas9 | Single, multi-domain protein forming crRNA-guided DNA-binding nuclease. |
| Type III | III-A (Csm), III-B (Cmr) | cas10 (Palm-domain polymerase) | Csm or Cmr complex | Multi-subunit complex; DNase/RNase activity; cOA signaling. |
| Type IV | IV-A, IV-B, IV-C | csf1 (Cas8-like) | Multi-subunit effector | Multi-subunit crRNA-guided complex; lacks Cas1/2 adaptation module. |
| Type V | V-A (Cpf1/Cas12a), V-K (Cas12i) | cas12 | Cas12 (e.g., Cpf1) | Single RuvC-containing protein; creates staggered DNA cuts. |
| Type VI | VI-A (Cas13a), VI-B (Cas13b) | cas13 | Cas13 | Single HEPN-containing protein; RNA-guided RNase activity. |
Table 2: Characteristic Loci Organization and Additional Genetic Markers
| Type | Minimal Adaptation Module | Repeat Length (bp) | Protospacer Adjacent Motif (PAM) | Key Accessory Genes |
|---|---|---|---|---|
| Type I | cas1, cas2, cas4 (often) | ~28-37 | 5' or 3' DNA, subtype-specific | cas5, cas6, cas7, cas8 (subtype-specific). |
| Type II | cas1, cas2, cas9; tracrRNA gene | ~36 | 3' NGG (for SpCas9) | tracrRNA (non-coding), cas4, csn2 (in some). |
| Type III | cas1, cas2 | ~35-40 | 5' flanking RNA sequence | cas5, cas6, cas7, cas10, csm2/cmr5 etc. |
| Type IV | Absent (non-autonomous) | ~35 | Not well-defined | csf1 (cas8-like), csf2, csf3, cas7, cas5. |
| Type V | cas1, cas2, cas4 (often) | ~36-44 | 5' T-rich (for Cas12a) | tracrRNA absent in most; cas4 often adjacent. |
| Type VI | cas1, cas2 (often absent) | ~30 | 5' or 3' flanking RNA sequence | HEPN-nuclease domain intrinsic to Cas13. |
Experimental Protocols for Identification and Validation
Protocol:In SilicoIdentification of CRISPR-Cas Loci from Genomic Data
Objective: To bioinformatically identify and classify CRISPR-Cas systems from whole-genome sequence assemblies. Methodology:
- Sequence Acquisition: Obtain genomic DNA sequences (FASTA format) from public databases (NCBI, ENA).
- CRISPR Array Detection: Use CRISPR identification tools (e.g., CRISPRCasFinder, PILER-CR) to locate direct repeat-spacer arrays.
- cas Gene Identification: Search genomic regions flanking identified arrays (typically ± 10 kb) for cas homologs using HMMER against curated protein family databases (e.g., Pfam, TIGRFAMs) or BLASTP against custom cas gene databases.
- Loci Delineation & Typing: Cluster identified cas genes into putative operons based on proximity and orientation. Assign type and subtype by matching the gene constellation to established signatures (Table 1 & 2).
- Phylogenetic Validation: Perform multiple sequence alignment and phylogenetic analysis of key marker proteins (e.g., Cas3, Cas9, Cas10) to confirm classification.
Protocol: Functional Validation of a Type V-A (Cas12a) System
Objective: To experimentally confirm the interference activity and PAM requirement of a putative Cas12a locus. Methodology:
- Cloning: Synthesize and clone the identified cas12a gene and its cognate CRISPR array into an expression vector.
- Reporter Assay Construction: Clone a target DNA sequence with a putative 5' TTTN PAM upstream of a reporter gene (e.g., GFP, LacZ) into a target plasmid.
- Transformation: Co-transform both plasmids into a suitable bacterial host (e.g., E. coli BL21) that lacks endogenous CRISPR systems.
- Interference Assay: Measure reporter activity (fluorescence, β-galactosidase) and compare to a control with a mutated PAM or spacer. Quantify survival of target plasmid via plating on selective media.
- In Vitro Cleavage Assay: Purify the recombinant Cas12a protein and transcribe the crRNA. Incubate with linearized target DNA plasmid. Analyze cleavage products via gel electrophoresis to confirm staggered double-strand breaks.
Visualization of Classification Logic and Workflow
Title: Decision Tree for CRISPR-Cas Typing Based on Effector Signatures
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagent Solutions for CRISPR-Cas Characterization Experiments
| Reagent / Material | Supplier Examples | Function in Research |
|---|---|---|
| High-Fidelity DNA Polymerase (e.g., Q5, Phusion) | NEB, Thermo Fisher | Amplifies cas operons and target sequences for cloning with minimal error. |
| CRISPR Array Synthesis Oligos | IDT, Twist Bioscience | For constructing custom CRISPR arrays with desired spacers to test targeting. |
| Recombinant Cas Protein Expression Systems (e.g., pET vectors) | Novagen (MilliporeSigma) | Provides high-yield purification of cas effector proteins for in vitro assays. |
| T7 RNA Polymerase & NTPs | NEB, Thermo Fisher | For in vitro transcription of crRNAs essential for functional validation assays. |
| HEK293T or E. coli (e.g., BL21) Cells | ATCC, Invitrogen | Model organisms for in vivo interference and plasmid survival assays. |
| Fluorescent Reporter Plasmids (e.g., pGFP, pRFP) | Addgene, Clontech | Quantifies CRISPR interference efficiency via fluorescence loss. |
| Next-Generation Sequencing (NGS) Library Prep Kits | Illumina, Oxford Nanopore | For deep sequencing to analyze spacer acquisition (adaptation) and target cleavage efficiency. |
| Anti-Cas Antibodies (e.g., anti-FLAG, anti-Cas9) | Abcam, Sigma, Cell Signaling | Validates protein expression and localization via Western blot or immunofluorescence. |
From Bench to Bedside: Harnessing Specific CRISPR-Cas Types for Research and Therapy
Type II (Cas9) as the Workhorse for Genome Editing and Knockout Strategies
CRISPR-Cas systems, adaptive immune systems in prokaryotes, are broadly classified into two classes (1-6) based on effector module architecture. Class 2 systems (Types II, V, VI) utilize a single, large effector protein for nucleic acid targeting and cleavage. Among these, the Type II CRISPR-Cas9 system, derived primarily from Streptococcus pyogenes (SpCas9), has emerged as the predominant workhorse for programmable genome editing in eukaryotic cells due to its simplicity, high efficiency, and versatility. Its development has catalyzed a revolution in functional genomics, drug target validation, and therapeutic development.
Core Mechanism of the Cas9 System
The canonical SpCas9 system requires two RNA components: the CRISPR RNA (crRNA) and the trans-activating crRNA (tracrRNA), which can be synthetically fused into a single-guide RNA (sgRNA). The sgRNA directs the Cas9 nuclease to a complementary ~20-nucleotide genomic DNA sequence adjacent to a Protospacer Adjacent Motif (PAM; 5'-NGG-3' for SpCas9). Cas9 undergoes a conformational change upon PAM recognition and local DNA melting, leading to RNA-DNA hybridization. This positions the HNH nuclease domain to cleave the complementary (target) DNA strand and the RuvC-like domain to cleave the non-complementary strand, generating a blunt-ended double-strand break (DSB).
Diagram: Cas9-sgRNA Complex and DNA Cleavage Mechanism
Diagram Title: Cas9-sgRNA RNP binds PAM and cleaves target DNA.
Key Experimental Protocols for Genome Editing and Knockout
Protocol: Mammalian Cell Gene Knockout via NHEJ
This protocol generates frameshift mutations via error-prone Non-Homologous End Joining (NHEJ) repair.
- sgRNA Design: Design a 20-nt guide sequence targeting an early coding exon of the gene of interest. Verify specificity using resources like CRISPick or CHOPCHOP.
- Cloning: Clone the sgRNA sequence into a plasmid vector containing both the sgRNA scaffold and a Cas9 expression cassette (e.g., pSpCas9(BB)-2A-Puro, Addgene #62988).
- Delivery: Transfect the plasmid into target mammalian cells (e.g., HEK293T) using a suitable transfection reagent (Lipofectamine 3000, FuGENE HD).
- Selection & Expansion: Apply appropriate selection (e.g., puromycin) 24-48h post-transfection for 3-5 days. Expand surviving cells.
- Validation:
- Genomic DNA Extraction: Harvest cells, extract gDNA.
- PCR Amplification: PCR amplify the target locus (amplicon size: 400-800 bp).
- Analysis:
- T7 Endonuclease I (T7EI) or Surveyor Assay: Detect heteroduplex formation from indel mixtures.
- Sanger Sequencing & Deconvolution: Sequence PCR products and analyze trace files with tools like TIDE or ICE to quantify editing efficiency.
Protocol: Knock-in via HDR
This protocol uses a donor DNA template with homology arms to introduce precise edits via Homology-Directed Repair (HDR).
- sgRNA Design: Design a sgRNA with the cut site as close as possible to the intended edit.
- Donor Template Construction: Synthesize a single-stranded oligodeoxynucleotide (ssODN) or a double-stranded DNA plasmid containing the desired edit (e.g., point mutation, tag) flanked by homology arms (70-100 bp each for ssODNs).
- Delivery: Co-deliver the following into target cells:
- Cas9: mRNA or purified protein.
- sgRNA: in vitro transcribed RNA or synthetic crRNA:tracrRNA duplex.
- Donor Template: ssODN or plasmid. Use electroporation (e.g., Neon system) for primary cells or sensitive lines.
- Screening: Isolate single-cell clones by limiting dilution or FACS. Screen clones by junction PCR and Sanger sequencing to identify precise edits.
Data Presentation: Comparative Analysis of Cas9 Variants and Delivery Methods
Table 1: Key Cas9 Orthologs and Engineered Variants
| Nuclease | Source | PAM Sequence | Size (aa) | Primary Applications | Notes |
|---|---|---|---|---|---|
| SpCas9 | S. pyogenes | 5'-NGG-3' | 1368 | General editing, KO, KI | Standard workhorse; high activity. |
| SpCas9-NG | Engineered | 5'-NG-3' | ~1368 | Expanded targeting | Relaxed PAM broadens target range. |
| SaCas9 | S. aureus | 5'-NNGRRT-3' | 1053 | In vivo delivery | Smaller size fits into AAV vectors. |
| SpCas9-D10A | Engineered | 5'-NGG-3' | 1368 | Nickase, HDR, base editing | "Nickase" creates single-strand breaks; reduces off-targets. |
| SpCas9-HF1 | Engineered | 5'-NGG-3' | 1368 | High-fidelity editing | Reduced non-specific DNA contacts; lowers off-target effects. |
Table 2: Common Delivery Methods for CRISPR-Cas9 Components
| Method | Components Delivered | Typical Efficiency | Advantages | Disadvantages |
|---|---|---|---|---|
| Plasmid DNA | Cas9 + sgRNA expression cassettes | 20-70% (transfectable lines) | Simple, cost-effective, sustained expression. | Risk of random integration, immunogenicity, toxicity. |
| RNP (Ribonucleoprotein) | Purified Cas9 protein + sgRNA | 50-90% (various cells) | Fast action, reduced off-targets, no DNA integration. | Transient activity, requires protein purification, delivery can be challenging. |
| Lentivirus | Integrative or non-integrative vectors | >80% (hard-to-transfect) | High efficiency, stable expression/transduction. | DNA integration risk (for integrating vectors), size limitations, biosafety level 2+. |
| AAV | Cas9/sgRNA expression cassettes | Varies by serotype & tissue | Low immunogenicity, excellent in vivo delivery. | Very small cargo capacity (<4.7 kb), persistent expression. |
Diagram: Major DNA Repair Pathways Following Cas9 Cleavage
Diagram Title: DNA repair pathways determine editing outcomes post-Cas9 cut.
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Cas9-Based Editing Experiments
| Reagent / Material | Supplier Examples | Function in Experiment |
|---|---|---|
| SpCas9 Expression Plasmid (e.g., pSpCas9(BB)) | Addgene, Thermo Fisher | Provides DNA template for expression of the Cas9 nuclease in cells. |
| sgRNA Cloning Vector (e.g., pU6-sgRNA) | Addgene, Sigma-Aldrich | Backbone for inserting and expressing the target-specific guide RNA. |
| Synthetic sgRNA or crRNA:tracrRNA | IDT, Synthego | Chemically synthesized, ready-to-use guide RNAs; high purity, rapid delivery. |
| Recombinant SpCas9 Nuclease Protein | IDT, NEB | Purified Cas9 protein for forming RNPs; enables DNA-free, rapid editing. |
| HDR Donor Template (ssODN) | IDT, Genewiz | Single-stranded DNA oligo with homology arms for precise sequence insertion or correction. |
| Lipofectamine CRISPRMAX | Thermo Fisher | A lipid-based transfection reagent optimized for RNP or plasmid delivery. |
| T7 Endonuclease I | NEB, Thermo Fisher | Enzyme for mismatch cleavage assay to detect indels at target locus. |
| Guide-it Genotype Confirmation Kit | Takara Bio | A complete solution for PCR amplification and T7EI analysis of edited loci. |
| RNeasy / DNeasy Kits | Qiagen | For high-quality RNA/DNA extraction from edited cells for downstream analysis. |
| Next-Generation Sequencing Library Prep Kit (e.g., Illumina) | Illumina, Roche | For deep sequencing of target sites to comprehensively assess on- and off-target editing. |
Leveraging Type V (Cas12) for DNA Detection, Diagnostics, and Multiplex Editing
The systematic classification of CRISPR-Cas systems into Types I-VI is based on distinct effector module architectures and interference mechanisms. Within this framework, Type V systems, characterized by single effector proteins like Cas12, are distinguished by their RuvC-like nuclease domain and lack of an HNH domain. This unique architecture enables DNA cleavage and, notably, trans-cleavage activity upon target recognition, making it exceptionally versatile for applications beyond canonical gene editing. This guide details the technical exploitation of Cas12's dual cis and trans cleavage activities for diagnostics and multiplexed genome engineering.
Core Mechanisms and Quantitative Comparison
Table 1: Comparison of Key Type V (Cas12) Effectors
| Effector | PAM Sequence (5'->3') | Cleavage Pattern (Target Strand/ Non-Target Strand) | Trans-Cleavage Activity | Primary Applications |
|---|---|---|---|---|
| Cas12a (Cpfl) | TTTV (V = A, C, G) | Staggered cut (18-23bp downstream) / Blunt cut | Yes (ssDNA) | Editing, Diagnostics (e.g., DETECTR) |
| Cas12b | ATT, TTT, TCT | Staggered cut | Yes (ssDNA) | Editing, Diagnostics (thermostable) |
| Cas12c/d | T-rich | Putative blunt cuts | No (inactive RuvC) | (Potential transcriptional repression) |
| Cas12e (CasX) | TTCN | Blunt cut | No | Compact editing |
| Cas12f (Cas14) | Not required | ssDNA target cleavage | Yes (ssDNA) | ssDNA detection, Diagnostics |
| Cas12i/j | T-rich | Staggered cut | Yes (ssDNA) | Compact editing, Diagnostics |
Table 2: Performance Metrics of Cas12-Based Diagnostic Platforms
| Platform/Assay | Limit of Detection (LoD) | Time-to-Result | Readout Method | Key Advantage |
|---|---|---|---|---|
| DETECTR | ~aM to fM (1-10 copies/µL) | <60 min | Fluorescent (FAM-quencher) | Isothermal, rapid |
| HOLMES | aM levels | 60-90 min | Fluorescent or Colorimetric | One-pot reaction |
| HOLMESv2 | Sub-aM | ~60 min | Fluorescent | Integrated RT-LAMP for RNA viruses |
| CDetection (Cas12b) | fM levels | 30 min | Lateral Flow Strip | Thermostable, field-deployable |
Experimental Protocols
Protocol 1: DETECTR Assay for Viral DNA Detection (e.g., HPV)
- Principle: Target DNA binding activates Cas12a's collateral trans-cleavage of reporter molecules, generating fluorescence.
- Reagents: Recombinant LbCas12a, crRNA, ssDNA-FQ reporter, target DNA, NEBuffer r2.1.
- Procedure:
- Reaction Setup: Assemble 25 µL reaction: 1x NEBuffer r2.1, 50 nM LbCas12a, 60 nM crRNA, 100 nM ssDNA-FQ reporter, 2 µL of extracted DNA sample.
- Incubation: Run reaction at 37°C for 30-60 minutes in a real-time PCR machine or fluorometer.
- Detection: Monitor fluorescence (λex/λem: 485/535 nm) in real-time or as an endpoint measurement. Positive samples show exponential signal increase.
- Validation: Use synthetic target oligos as positive control and no-template as negative control.
Protocol 2: Multiplexed Gene Editing Using Orthogonal Cas12a Arrays
- Principle: Co-expression of multiple, distinct Cas12a nucleases with their cognate crRNAs enables simultaneous editing at multiple loci.
- Reagents: Plasmids encoding LbCas12a, AsCas12a, and MbCas12a; a crRNA expression array targeting distinct genomic loci with compatible PAMs; transfection reagent; target cells.
- Procedure:
- Construct Design: Clone a polycistronic crRNA array where each spacer (targeting a different locus) is flanked by direct repeats compatible with its intended Cas12a ortholog.
- Delivery: Co-transfect mammalian cells with the three Cas12a expression plasmids and the single crRNA array plasmid.
- Analysis: Harvest genomic DNA 72-96h post-transfection. Assess editing efficiency at each target locus via T7E1 assay or next-generation sequencing (NGS).
- Key Consideration: Ensure strict orthogonality of crRNA-Cas12a pairs to prevent cross-talk.
Diagrams
Diagram 1: Cas12 Activation and Dual Cleavage Mechanism (100 chars)
Diagram 2: Cas12 Diagnostic Assay Workflow (90 chars)
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Cas12 Research & Development
| Reagent/Category | Example Product/Supplier | Function & Brief Explanation |
|---|---|---|
| Recombinant Cas12 Proteins | LbCas12a (NEB, IDT), AsCas12a (Thermo), AapCas12b (MCLAB) | Purified effector proteins for in vitro cleavage, diagnostic assay development, and RNP complex formation. |
| crRNA Synthesis Kits | Custom Alt-R CRISPR crRNA (IDT), Synthetic crRNA (Synthego) | High-purity, chemical synthesis of target-specific guide RNAs with modified termini for enhanced stability. |
| Fluorescent-Quencher Reporters | ssDNA-FQ (FAM-TTATT-BHQ1, etc.) (Biosearch Tech) | Substrates for trans-cleavage activity. Cleavage separates fluorophore from quencher, generating signal. |
| Isothermal Amplification Kits | TwistAmp Basic (RPA) Kit (TwistDx), LAMP Kit (NEB) | For pre-amplifying target DNA/RNA prior to Cas12 detection, enabling ultra-high sensitivity (aM). |
| Lateral Flow Readout Systems | Milenia HybriDetect (TwistDx) | Strip-based visual detection of cleaved reporter fragments, enabling equipment-free diagnostics. |
| Cas12 Expression Plasmids | pY010 (Addgene #69976), MBPCas12a (Addgene #89344) | Mammalian expression vectors for stable or transient delivery of Cas12 and crRNA arrays for editing. |
| Orthogonal Cas12 Variants | LbCas12a, AsCas12a, MbCas12a kits (ToolGen) | Distinct Cas12 proteins with non-overlapping PAM requirements for multiplexed genome editing. |
| Nuclease Activity Buffers | NEBuffer r2.1/r3.1 (NEB), ToloBio Cas12a Buffer | Optimized reaction buffers providing ideal ionic conditions for maximal Cas12 cleavage efficiency. |
Utilizing Type VI (Cas13) for RNA Knockdown, Editing, and SHERLOCK Diagnostics
The systematic classification of CRISPR-Cas systems into six primary types (I-VI) delineates their evolutionary relationships and mechanistic diversity. Types I, III, and IV utilize multi-subunit effector complexes, while Types II, V, and VI are characterized by single, large effector proteins. Type VI systems, featuring the RNA-guided RNase Cas13, represent a distinct evolutionary lineage adapted exclusively for RNA targeting. Unlike DNA-targeting systems (Types II, V), Cas13's collateral RNase activity upon target recognition provides a unique foundation for both programmable RNA manipulation and ultrasensitive diagnostic applications, expanding the CRISPR toolkit beyond genomic editing.
Mechanism of Cas13: Target Binding and Collateral Cleavage
Cas13 proteins (subtypes A, B, C, D) possess two Higher Eukaryotes and Prokaryotes Nucleotide-binding (HEPN) domains that confer RNase activity. Upon binding to a specific RNA target via its CRISPR RNA (crRNA) guide, Cas13 undergoes a conformational change that activates its non-specific, collateral cleavage of nearby single-stranded RNA (ssRNA). This trans-cleavage is the cornerstone of both RNA knockdown and diagnostic applications. Editing applications, such as REPAIR, utilize an inactivated Cas13 (dCas13) fused to effector domains.
Diagram 1: Cas13 activation and collateral cleavage
Applications: Protocols and Methodologies
RNA Knockdown (in Mammalian Cells)
Protocol: Transient, programmable RNA knockdown using catalytically active Cas13 (e.g., Cas13a from Leptotrichia wadei, LwaCas13a).
- Guide Design: Design a 28-nt spacer sequence within the target mRNA, avoiding secondary structures. A 5' protospacer flanking site (PFS) preference (e.g., an 'A' for LwaCas13a) must be considered.
- Cloning: Clone the spacer into an expression vector containing a U6-driven crRNA scaffold. Co-transfect with a mammalian expression vector for the Cas13a protein (e.g., CMV promoter-driven, NLS-tagged).
- Delivery: Transfect HEK293T or relevant cell line using a lipid-based transfection reagent (e.g., Lipofectamine 3000). Use a 1:2 mass ratio of Cas13 plasmid to crRNA plasmid.
- Analysis: Harvest cells 48-72 hours post-transfection. Assess knockdown efficiency via:
- qRT-PCR: Quantify remaining target mRNA levels.
- Western Blot: Measure reduction in corresponding protein levels.
- Controls: Include a non-targeting crRNA control and measure off-target effects by RNA-seq.
RNA Editing (RESCUE & REPAIR Systems)
Protocol: C-to-U or A-to-I RNA editing using dCas13-ADAR fusions.
- Effector Construction: Clone a catalytically dead Cas13 (dCas13; HEPN mutations: R...H) fused to the deaminase domain of ADAR2 (for REPAIRv2, A-to-I) or an engineered ADAR2 variant (for RESCUE, C-to-U) into a mammalian expression vector.
- Guide Design: Design a crRNA that positions the target adenosine (A) or cytidine (C) within a specific editing window (typically ~50-nt 5' of the spacer). For improved specificity, incorporate mutations in the crRNA to recruit endogenous ADAR proteins.
- Cell Transfection: Co-transfect the dCas13-ADAR plasmid and the crRNA plasmid into cells as in 3.1.
- Validation: Harvest RNA 48-72 hours post-transfection.
- Perform RT-PCR on the target region.
- Sanger Sequencing or Next-Generation Sequencing (NGS): To quantify the percentage of edited transcripts.
- Confirm functional protein changes if applicable.
SHERLOCK Diagnostics
Protocol: Specific High-sensitivity Enzymatic Reporter unLOCKing for pathogen or variant detection.
- Sample Preparation: Extract RNA from sample (saliva, serum, etc.) using a commercial kit. Perform isothermal amplification (e.g., RPA - Recombinase Polymerase Amplification at 37-42°C for 15-30 min) to amplify target sequences, incorporating a T7 promoter.
- In Vitro Transcription (IVT): Use T7 RNA polymerase to transcribe the RPA amplicon to RNA (37°C, 30 min).
- Cas13 Detection Reaction:
- Prepare a reaction mix containing: LwaCas13a protein (50-100 nM), specific crRNA (50-100 nM), and a fluorescent quenched RNA reporter (e.g., FAM-UUUUU-BHQ1, 500 nM) in a buffer.
- Add the IVT RNA product to the mix.
- Incubate at 37°C in a real-time PCR machine or lateral flow strip.
- Readout:
- Fluorometer/Plate Reader: Measure fluorescence increase over time (kinetic readout).
- Lateral Flow: Use FAM- and biotin-labeled reporters. Cleavage prevents capture, yielding a test line. Results in 5-10 minutes.
- Quantification: Compare to a standard curve for absolute quantification.
Diagram 2: SHERLOCK diagnostic workflow
Table 1: Performance Metrics of Cas13 Applications
| Application | System/Variant | Key Metric | Reported Efficiency/Range | Notes |
|---|---|---|---|---|
| RNA Knockdown | LwaCas13a in HEK293T | mRNA Reduction | 50-95% | Efficiency varies by target site; off-target effects observed in early systems. |
| RNA Editing (A-to-I) | REPAIRv2 (dPspCas13b) | Editing Efficiency at Target Site | 20-40% (optimized sites) | Specificity improved >1000x over REPAIRv1. |
| RNA Editing (C-to-U) | RESCUE (dPspCas13b) | Editing Efficiency at Target Site | Up to ~30% | Enables C-to-U changes, expanding editome. |
| SHERLOCK Detection | SHERLOCKv2 (LwaCas13a) | Limit of Detection (LoD) | ~2 attomolar (aM) | Single molecule sensitivity; specific to single nucleotide variants. |
| SHERLOCK Multiplexing | SHERLOCKv2 | Simultaneous Targets | Up to 4 targets | Uses orthogonal Cas13 variants (Cas13a, Cas13b, Cas13c, Cas13d). |
Table 2: Comparison of Common Cas13 Orthologs
| Ortholog | Size (aa) | PFS Preference | Collateral Activity | Primary Use Case |
|---|---|---|---|---|
| LwaCas13a | ~1250 | 5' A, H (A/C/U) | High | Knockdown, SHERLOCK diagnostics |
| PspCas13b | ~1120 | 5' D (A/G/U) | Very High | Knockdown, Editing (REPAIR/RESCUE) |
| RfxCas13d (CasRx) | ~930 | None | High | Knockdown (compact size favored in vivo) |
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Cas13 Research
| Reagent / Material | Function / Purpose | Example Vendor/Product |
|---|---|---|
| Cas13 Expression Plasmids | Mammalian or bacterial expression vectors for active or dead Cas13 orthologs. | Addgene (LwaCas13a, dPspCas13b) |
| crRNA Cloning Vectors | Vectors with U6 or T7 promoters for expressing custom guide RNAs. | Addgene, IDT (gBlock synthesis) |
| Fluorescent Quenched RNA Reporters | ssRNA oligos with fluorophore-quencher pairs (e.g., FAM-UUUUU-BHQ1) for detecting collateral cleavage. | Integrated DNA Technologies (IDT) |
| Recombinase Polymerase Amplification (RPA) Kit | For isothermal nucleic acid amplification in SHERLOCK protocols. | TwistDx |
| T7 RNA Polymerase Kit | For in vitro transcription of RPA amplicons to generate Cas13 substrates. | NEB HiScribe |
| RNA Purification Kits | For clean extraction of RNA from cells or clinical samples prior to analysis or SHERLOCK. | Qiagen, Zymo Research |
| Nucleofection / Transfection Reagents | For efficient delivery of Cas13 RNP or plasmids into difficult cell lines. | Lonza Nucleofector, Lipofectamine |
| Next-Generation Sequencing (NGS) Library Prep Kits | For comprehensive assessment of editing efficiency and off-target transcriptome effects. | Illumina, Oxford Nanopore |
Emerging Applications of Type I and Type III Systems for Large-Scale Genomic Deletions and Antiviral Strategies
Within the CRISPR-Cas classification framework (Types I-VI), Type I (multi-subunit effector complexes) and Type III (RNA-targeting, multi-protein complexes) systems are distinguished by their unique mechanisms. Unlike the widely adopted Type II (Cas9) and Type V (Cas12) systems, Types I and III offer distinct advantages for large-scale genomic engineering and host-cell antiviral defense, respectively. This guide details their emerging applications and technical protocols.
Type I Systems: Cascade-Driven Large Deletions
Type I systems (e.g., I-A to I-F) utilize a CRISPR RNA (crRNA)-guided surveillance complex (Cascade) to recognize dsDNA targets, recruiting the helicase-nuclease Cas3 for processive degradation. Cas3's helicase activity unwinds DNA, while its nuclease activity creates single-stranded breaks, leading to extensive resection and kilobase-scale deletions.
Type III Systems: Transcription-Coupled Antiviral Defense
Type III systems (e.g., III-A, III-B) recognize target RNA transcripts, activating dual enzymatic activities. The Cas10 subunit synthesizes cyclic oligonucleotide second messengers (cOAs) that activate non-specific nucleases (e.g., Csm6, Csx1), while also exhibiting collateral RNase and DNase activities under certain conditions, providing a robust, amplification-based antiviral response.
Table 1: Quantitative Comparison of Type I and Type III Systems
| Feature | Type I System (e.g., I-C, I-E) | Type III System (e.g., III-A, III-B) |
|---|---|---|
| Target | Double-stranded DNA | Single-stranded RNA (primary) / ssDNA (collateral) |
| Effector Complex | Cascade (multi-protein) + Cas3 | Csm/Cmr RNP complex (multi-protein) |
| Signature Enzyme | Cas3 (helicase-nuclease) | Cas10 (Palm-domain polymerase, DNase/RNase) |
| Cleavage Outcome | Large deletions (1-100 kb) via processive resection | Transcription attenuation, RNA cleavage, collateral degradation |
| Second Messenger | No | Yes (cOA synthesis: cA4, cA6) |
| Typical Deletion Size | 7-100 kilobases | Not applicable for genomic DNA deletion |
| PAM Requirement | Yes (specific to subtype) | No (requires transcription bubble) |
| Key Application | Chromosomal engineering, mega-deletions | Antiviral defense, RNA knockdown, nucleic acid detection |
Experimental Protocols
Protocol for Type I-C Mediated Large Genomic Deletions inE. coli
Objective: Generate a defined 50-kb chromosomal deletion. Materials: See "The Scientist's Toolkit" (Section 5). Methodology:
- crRNA Array Design & Cloning: Design two 32-nt crRNAs targeting the 5' and 3' boundaries of the genomic region to be deleted. Clone spacers sequentially into plasmid pCASCADE under a constitutive promoter.
- Transformation: Electroporate the engineered pCASCADE plasmid and a helper plasmid expressing Cas3 (pCas3) into the target E. coli strain.
- Induction & Selection: Grow transformed cells in LB + antibiotics at 30°C to OD600 0.5. Add 0.2% L-arabinose to induce Cascade and Cas3 expression for 4 hours.
- Screening: Plate serial dilutions on non-selective media. Screen individual colonies by colony PCR using primers flanking the target deletion region. A successful deletion yields a ~1 kb product vs. ~51 kb for WT.
- Validation: Confirm deletion boundaries by Sanger sequencing of PCR products. Verify phenotype as expected.
Protocol for Assessing Type III-B Antiviral Activity via RNA Challenge
Objective: Quantify antiviral immunity provided by a Pyrococcus furiosus Type III-B system in response to phage RNA. Materials: See "The Scientist's Toolkit" (Section 5). Methodology:
- Reconstitution in vitro: Purify recombinant PfCmr complex and Csx1 nuclease. Assemble the Cmr-crRNA complex by incubating purified Cmr proteins with synthetic crRNA (37°C, 15 min).
- RNA Target Challenge: In a reaction buffer (20 mM Tris-HCl pH 7.5, 150 mM KCl, 5 mM MgCl2), combine 100 nM Cmr-crRNA complex with 500 nM target viral RNA transcript. Incubate at 55°C (optimal for P. furiosus) for 30 min.
- cOA Extraction & Detection: Add 0.5 U of heat-labile phosphatase to degrade NTPs. Heat-inactivate (75°C, 5 min). Analyze supernatant by LC-MS/MS to detect and quantify cyclic tetra-adenylate (cA4) production.
- Collateral RNase Assay: In a separate reaction, include a fluorophore-quencher labeled nonspecific RNA reporter. Measure fluorescence increase (ex/em 485/535 nm) over 60 min to quantify collateral Csx1 activation.
- Data Analysis: Correlate cA4 synthesis levels with collateral RNase activity and viral RNA degradation (by gel electrophoresis).
Visualizations
Diagram 1: Type I CRISPR for large genomic deletions
Diagram 2: Type III antiviral signaling pathway
The Scientist's Toolkit: Essential Research Reagents
Table 2: Key Reagent Solutions for Featured Experiments
| Reagent/Material | Function in Experiment | Example Source/Product |
|---|---|---|
| pCASCADE Plasmid | Expresses the Cascade crRNA array for target recognition. | Addgene #Multiple (subtype-specific) |
| pCas3 Expression Plasmid | Provides inducible or constitutive expression of the Cas3 nuclease-helicase. | Custom cloning or genomic locus. |
| L-Arabinose | Inducer for arabinose-promoter driven expression of CRISPR components in bacteria. | Sigma-Aldrich, A91906. |
| High-Fidelity DNA Polymerase | Accurate amplification of large genomic regions for deletion screening. | NEB Q5, Thermo Fisher Phusion. |
| Recombinant Cmr/Csm Complex | Purified Type III effector complex for in vitro reconstitution assays. | Lab purification from E. coli overexpression. |
| Cyclic Oligoadenylate (cA4/cA6) Standard | LC-MS/MS standard for quantifying second messenger production. | Biolog Life Science Institute, C-140. |
| Fluorophore-Quencher RNA Reporter (e.g., 6-FAM/ddQ) | Substrate for detecting collateral RNase activity. | Integrated DNA Technologies, Custom. |
| Heat-Labile Phosphatase | Degrades NTPs prior to cOA detection without degrading cOAs. | NEB, M0371S. |
CRISPR-Based Genetic Screens and Functional Genomics Using Different Effectors
CRISPR-Cas systems, classified into Types I-VI, offer a diverse toolkit for functional genomics. This whitepaper details the application of these systems in genetic screens, leveraging their distinct effector mechanisms—from DNA cleavage by Cas9 (Type II) to targeted transcriptional modulation or nucleic acid cleavage by Cas12, Cas13, and beyond. Framed within the broader thesis of CRISPR system classification, this guide provides a technical roadmap for designing and executing sophisticated genetic screens.
Effector Mechanisms Across CRISPR-Cas Types
The core function of each CRISPR-Cas type is defined by its effector complex, which dictates its application in genetic screens.
Table 1: CRISPR-Cas Effectors for Functional Genomics
| CRISPR-Cas Type | Effector Protein | Native Function | Primary Application in Screens | Target Molecule |
|---|---|---|---|---|
| Type II | Cas9 | dsDNA cleavage | Knockout, Activation/Repression | DNA |
| Type V (A) | Cas12a (Cpf1) | dsDNA cleavage | Knockout, Multiplexing | DNA |
| Type V (B) | Cas12b | dsDNA cleavage | Knockout (thermotolerant) | DNA |
| Type VI | Cas13a/d | ssRNA cleavage | RNA knockdown, imaging | RNA |
| Type I | Cascade-Cas3 | dsDNA degradation | Large deletions, knockouts | DNA |
| Type III | Cas10 complex | ssRNA/DNA cleavage | RNA targeting, immunity | RNA/DNA |
| Type II (Deriv.) | dCas9 | DNA binding | Epigenetic editing, imaging | DNA |
Experimental Protocols for Key Screen Modalities
Protocol: Genome-Wide CRISPR Knockout Screen Using Lentiviral sgRNA Libraries (Type II/V)
This protocol enables identification of genes essential for a cellular phenotype (e.g., drug resistance, cell growth).
Materials:
- Lentiviral sgRNA Library: (e.g., Brunello, 4 sgRNAs/gene, ~76,441 sgRNAs total).
- Target Cells: Dividing mammalian cells (e.g., HeLa, HEK293T).
- Packaging Plasmids: psPAX2, pMD2.G.
- Selection Agent: Puromycin.
- Reagents: Polybrene, Lenti-X Concentrator, DNA extraction kit, NGS library prep kit.
Method:
- Library Virus Production: Co-transfect HEK293T cells with sgRNA library plasmid, psPAX2, and pMD2.G using PEI transfection reagent. Harvest virus-containing supernatant at 48h and 72h post-transfection. Concentrate using Lenti-X.
- Cell Infection & Selection: Seed target cells in 6-well plates. Transduce with virus at an MOI of ~0.3 to ensure single sgRNA integration in the presence of 8 µg/mL Polybrene. After 24h, replace medium. At 48h post-transfection, begin selection with 1-3 µg/mL puromycin for 5-7 days.
- Screen Execution: Split selected cells into experimental (e.g., drug-treated) and control (DMSO) arms. Maintain cultures for 14-21 population doublings, keeping a minimum of 500 cells per sgRNA to maintain library representation.
- Genomic DNA Extraction & NGS: Harvest ≥ 1e7 cells per condition. Extract gDNA (Qiagen Maxi Prep). PCR-amplify integrated sgRNA sequences using 25µg gDNA per 100µL reaction with barcoded primers (22-28 cycles). Pool amplicons and sequence on an Illumina HiSeq (75bp single-end).
- Analysis: Align reads to library reference. Calculate sgRNA abundance fold-change and statistical significance (e.g., MAGeCK or BAGEL algorithm).
Protocol: CRISPR Interference (CRISPRi) Screen with dCas9-KRAB (Type II Derivative)
For transcriptional repression, ideal for studying essential genes without inducing double-strand breaks.
Materials:
- CRISPRi sgRNA Library: Targeting transcriptional start sites (TSS; -50 to +300 bp).
- Stable Cell Line: Expressing dCas9-KRAB fusion protein.
- Other reagents as in 3.1.
Method:
- Generate or obtain a cell line stably expressing dCas9-KRAB (e.g., via lentiviral transduction and blasticidin selection).
- Perform lentiviral transduction of the CRISPRi sgRNA library (e.g., Dolcetto library) into the dCas9-KRAB cell line as in Steps 1-2 of Protocol 3.1, but without puromycin selection if the dCas9 and sgRNA vectors share the same resistance marker. Use FACS to select for sgRNA+ cells if a fluorescent marker is present.
- Apply phenotypic pressure (e.g., differentiation inducer) and harvest cells at relevant time points.
- Process and analyze as in Steps 4-5 of Protocol 3.1.
Protocol: CRISPR Activation (CRISPRa) Screen Using dCas9-VPR
For transcriptional activation, identifying genes whose overexpression drives a phenotype.
Materials:
- CRISPRa sgRNA Library: Designed for enhancer regions or upstream of TSS (e.g., Calabrese library).
- Stable Cell Line: Expressing dCas9-VPR (VP64-p65-Rta) activator.
- Protocol is identical to 3.2, substituting the CRISPRa library and dCas9-VPR cell line.
Protocol: RNA-Targeting Screen with Cas13 (Type VI)
For knocking down RNA transcripts in the cytoplasm, assessing post-transcriptional effects.
Materials:
- Cas13 crRNA Library: Targeting exonic regions of transcripts.
- Stable Cell Line: Expressing catalytically active Cas13d (e.g., RfxCas13d) with nuclear localization signal removed.
- Other reagents as in 3.1.
Method:
- Generate a cell line stably expressing Cas13d.
- Transduce with lentiviral library of crRNA expression constructs. Select.
- Proceed with phenotypic assay, harvest, and NGS analysis of integrated crRNA cassettes.
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for CRISPR Screens
| Item | Function & Description | Example Vendor/Product |
|---|---|---|
| Genome-wide sgRNA Library | Pooled guide constructs for targeting each gene; cloned into lentiviral backbone. | Addgene (Brunello KO, Dolcetto i, Calabrese a) |
| dCas9 Effector Fusion Construct | Nuclease-dead Cas9 fused to transcriptional modulator (KRAB for repression, VPR for activation). | Addgene (pLV hU6-sgRNA hUbC-dCas9-KRAB-T2a-Puro) |
| Cas13d Expression Construct | Vector for stable expression of Type VI RNA-targeting effector. | Addgene (pRfxCas13d) |
| Lentiviral Packaging Plasmids | Essential for producing replication-incompetent viral particles. | Addgene (psPAX2, pMD2.G) |
| Polybrene (Hexadimethrine Bromide) | Cationic polymer that enhances viral transduction efficiency. | Sigma-Aldrich (H9268) |
| Lenti-X Concentrator | Simplifies lentivirus concentration via precipitation. | Takara Bio (631231) |
| Next-Generation Sequencing Kit | For preparing sgRNA amplicon libraries from genomic DNA. | Illumina (Nextera XT) |
| Screen Analysis Software | Computationally identifies enriched/depleted sgRNAs. | MAGeCK, BAGEL, CRISPRcloud |
Data Analysis & Key Metrics
Table 3: Quantitative Benchmarks for CRISPR Screen Success
| Parameter | Optimal Value or Benchmark | Rationale |
|---|---|---|
| Library Coverage (Cells/sgRNA) | >500-1000x at start | Prevents stochastic sgRNA dropout |
| Viral Transduction MOI | 0.3-0.5 | Minimizes cells with multiple integrations |
| Post-Selection sgRNA Representation | >95% of library recovered | Indicates successful library delivery |
| Phenotype Duration | 14-21 population doublings | Allows phenotype manifestation |
| NGS Read Depth | >200 reads/sgRNA | Enables robust statistical power |
| Key Hit Validation Rate (Typical) | 60-80% | Measures screen precision |
Visualization of Workflows and Mechanisms
Title: CRISPR Screen Experimental Workflow
Title: Effector Mechanisms and Outcomes
The strategic selection of CRISPR-Cas effector proteins, from Types I to VI, enables precise perturbation of DNA, RNA, and epigenetic states at scale. Integrating these tools into robust screening protocols, as detailed herein, allows researchers to systematically map gene function, identify drug targets, and validate mechanisms within the comprehensive framework of CRISPR system biology. Continuous evolution of these effectors promises even greater precision and novel screening modalities in functional genomics.
Therapeutic applications of CRISPR-Cas systems are fundamentally linked to the mechanistic diversity encapsulated within the classification of Types I-VI. This framework, based on signature genes, cas operon organization, and effector module structure, directly informs therapeutic strategy selection. Type II (single effector Cas9) and Type V (single effector Cas12) systems are predominantly harnessed for in vivo gene therapy due to their DNA targeting simplicity and high editing efficiency. Type VI systems (Cas13), which target RNA, offer distinct advantages for modulating gene expression without permanent genomic alteration and for antiviral applications. The multi-protein effector complexes of Types I, III, and IV, while less utilized currently, present future opportunities for sophisticated genomic targeting and immune modulation. This whitepaper details the translation of these molecular tools into three pivotal therapeutic arenas, grounded in the latest experimental data and protocols.
In Vivo Gene Therapy
Technical Overview: In vivo gene therapy involves the direct delivery of CRISPR components into a patient's body to correct genetic defects at their source. This approach is transformative for monogenic disorders. The choice of CRISPR system (primarily Cas9 or Cas12) and delivery vector (viral vs. non-viral) is critical.
Key Experimental Protocol: AAV-Mediated Delivery for Liver-Directed Therapy (e.g., for Hereditary Transthyretin Amyloidosis)
- CRISPR Construct Design: Clone a SaCas9 (or a compact Cas12 variant) expression cassette and a disease-specific sgRNA sequence into an AAV vector (serotype 8 or 9 for hepatotropism). A donor DNA template for HDR may be included for precise corrections.
- Vector Production & Purification: Produce the recombinant AAV (rAAV) using a triple-transfection method in HEK293 cells, followed by purification via iodixanol gradient ultracentrifugation and concentration.
- In Vivo Administration: Systemically administer the rAAV via tail-vein injection into a transgenic mouse model of the disease. A standard dose ranges from 1e11 to 5e12 vector genomes per gram of mouse body weight.
- Analysis: At predetermined timepoints (e.g., 4, 8, 12 weeks), assess:
- Editing Efficiency: Deep sequencing of the target locus from isolated hepatocyte genomic DNA.
- Protein/Functional Correction: ELISA or mass spectrometry for mutant protein reduction in serum; histology of target tissues.
- Safety: Surveyor or T7E1 assay for off-target editing at predicted loci; liver enzyme (ALT/AST) assays for toxicity.
Quantitative Data Summary: Recent Clinical & Preclinical Outcomes
Table 1: In Vivo Gene Therapy Clinical Trial & Preclinical Data
| Target Disease | CRISPR System | Delivery Method | Primary Outcome (Efficiency) | Key Safety Notes | Phase/Status |
|---|---|---|---|---|---|
| Hereditary Transthyretin Amyloidosis (ATTR) | SaCas9 | AAV9 (systemic) | >90% TTR protein reduction in serum (clinical) | Low incidence of mild liver enzyme elevation | Clinical I (Completed) |
| Leber Congenital Amaurosis 10 | SpCas9 (ABE) | AAV5 (subretinal) | 28% allelic editing in retinal cells (clinical) | No serious adverse events related to treatment | Clinical I/II (Ongoing) |
| Huntington's Disease (Mouse Model) | SpCas9 | AAV-PHP.eB (systemic) | ~45% reduction in mutant HTT aggregates in brain | Minimal off-target effects by GUIDE-seq | Preclinical |
| Duchenne Muscular Dystrophy (Mouse Model) | Cas12 (Cpf1) | AAV9 (systemic) | Exon skipping restored dystrophin in ~50% of cardiomyocytes | No evidence of cardiotoxicity | Preclinical |
Diagram: Workflow for AAV-Mediated In Vivo Gene Editing
Research Reagent Solutions: In Vivo Gene Editing
Table 2: Essential Reagents for In Vivo Gene Therapy Research
| Reagent/Material | Function & Rationale |
|---|---|
| AAV Serotype 8/9 Helper-Free System | Provides all components for producing recombinant AAV without helper virus contamination; serotypes dictate tissue tropism (e.g., liver, CNS, muscle). |
| HEK293T/AAV-293 Cells | Packaging cell line optimized for high-titer AAV production via triple-transfection. |
| Iodixanol Density Gradient Medium | Enables high-purity, high-recovery purification of AAV vectors away from cellular debris and empty capsids. |
| Next-Generation Sequencing Kit | For deep amplicon sequencing to quantify on-target editing efficiency and profile off-target events. |
| T7 Endonuclease I or Surveyor Nuclease | For initial, rapid detection of indel formation at the target site (mismatch cleavage assays). |
| Animal Model (Transgenic/Patient-Derived Xenograft) | Genetically accurate model for testing efficacy and safety of the in vivo editing approach. |
Cancer Immunotherapy
Technical Overview: CRISPR is revolutionizing cancer immunotherapy by engineering immune cells (e.g., T cells, NK cells) to enhance their tumor-targeting specificity, persistence, and potency. Key strategies include generating chimeric antigen receptor (CAR) T cells, disrupting immune checkpoint genes (e.g., PD-1), and creating allogeneic "off-the-shelf" cell products.
Key Experimental Protocol: Manufacturing CRISPR-Engineered CAR-T Cells
- Design & Preparation: Design sgRNAs targeting the endogenous T cell receptor alpha constant (TRAC) locus and/or PDCD1. Prepare Cas9 ribonucleoprotein (RNP) complexes by mixing purified SpCas9 protein with synthetic sgRNA.
- T Cell Isolation & Activation: Isolate primary human T cells from leukapheresis product using a Ficoll gradient and CD3/CD28 magnetic bead activation.
- Electroporation: At 48 hours post-activation, electroporate T cells with the Cas9 RNP complex using a nucleofector device. Simultaneously, deliver an AAV6 donor template encoding the CAR construct, targeted for integration into the TRAC locus.
- Expansion & Validation: Expand cells in IL-2 and IL-7/IL-15 containing media for 10-14 days. Validate by:
- Flow Cytometry: For surface CAR expression and knockout of TCR and PD-1.
- Functional Assays: In vitro cytotoxicity against antigen-positive tumor cells; cytokine (IFN-γ, IL-2) release.
- Genomic Safety: Targeted deep sequencing of the on-target site and top predicted off-target sites.
Quantitative Data Summary: Engineered Cell Therapy Trials
Table 3: CRISPR-Engineered Cancer Immunotherapy Data
| Therapy Target | Cell Type | CRISPR Edit(s) | Key Efficacy Metric | Notable Safety Findings | Clinical Stage |
|---|---|---|---|---|---|
| NY-ESO-1+ Solid Tumors | T Cells | TRAC & PDCD1 KO | Persistence of edited cells up to 9 months | No treatment-related severe AEs; edits stable | Phase I (Ongoing) |
| BCMA+ Multiple Myeloma | Allogeneic CAR-T | TRAC & B2M KO | 100% overall response rate in small cohort (clinical) | No graft-vs-host disease reported | Phase I (Reported) |
| CD19+ Leukemia/Lymphoma | CAR-T Cells | TRAC Knock-in | Superior expansion and tumor control vs. retroviral CAR in preclinical models | Reduced T cell exhaustion phenotype | Preclinical/IND-enabling |
| Mesothelin+ Solid Tumors | TCR-T Cells | Endogenous TCR KO & PD-1 KO | Enhanced anti-tumor activity in mouse xenograft model | Mitigated T cell dysfunction | Preclinical |
Diagram: CRISPR Engineering of Allogeneic CAR-T Cells
Antimicrobials
Technical Overview: CRISPR-Cas systems, particularly the RNA-targeting Type VI (Cas13) and DNA-targeting Types II (Cas9) and V (Cas12), are being deployed as sequence-specific antimicrobials. Strategies include: (a) Bacteriophage Delivery: Cas systems programmed to destroy antibiotic resistance genes or essential bacterial genomic sequences; (b) Diagnostic (DETECTR): Leveraging Cas12/Cas13 collateral cleavage for pathogen nucleic acid detection; (c) Antiviral: Using Cas13 to target and degrade RNA viral genomes within human cells.
Key Experimental Protocol: Cas13a-Based Antiviral Therapy in Human Cells
- Design & Cloning: Design crRNAs targeting conserved regions of the viral RNA genome (e.g., Influenza A virus segments). Clone a mammalian expression vector for catalytically active LwaCas13a with a nuclear localization signal, followed by an array of crRNA sequences under a U6 promoter.
- Cell Culture & Infection: Seed A549 (lung epithelial) cells in a 24-well plate. Co-transfect with the Cas13a-crRNA plasmid and a reporter plasmid expressing the viral RNA segment fused to a fluorescent tag.
- Challenge & Assessment: 24h post-transfection, infect cells with Influenza A virus (MOI=0.1). At 48h post-infection, harvest cells and supernatant.
- Analysis:
- Viral Load: Quantify viral titer via plaque assay or RT-qPCR of supernatant.
- Target Engagement: Measure fluorescence loss from the reporter construct via flow cytometry.
- Cell Viability: Assess via MTT or similar assay to confirm Cas13a treatment specificity.
Quantitative Data Summary: Antimicrobial CRISPR Applications
Table 4: CRISPR-Based Antimicrobial Efficacy Data
| Target Pathogen | CRISPR System | Delivery Method | Efficacy Metric | Specificity/Resistance Notes |
|---|---|---|---|---|
| Staphylococcus aureus (MRSA) | SaCas9 (Phage) | Engineered Lysogen | >99.9% killing in vitro; reduced biofilm load in vivo | Selective against MRSA, spares other flora |
| Influenza A Virus | LwaCas13a | Lentivirus (Human Cells) | ~100-1000 fold reduction in viral titer | Effective against multiple strains; no host RNA collateral damage |
| Mycobacterium tuberculosis | FnCas12a (DETECTR) | N/A (Diagnostic) | Sensitivity: 1 CFU/mL in sputum | Distinguishes drug-resistant strains by SNP detection |
| Pseudomonas aeruginosa | Cas3 (Type I) | Conjugative Plasmid | Re-sensitized to 3 different antibiotic classes | Eliminates plasmid-borne resistance genes |
Diagram: CRISPR-Cas Antimicrobial Strategies
The therapeutic translation of CRISPR-Cas systems is a direct function of their underlying classification. The precision of Cas9/Cas12 for DNA modification anchors in vivo gene therapy and cellular engineering for cancer. The RNA-guided RNA cleavage by Cas13 opens unique avenues for antiviral development and diagnostics. As research into the diverse mechanisms of Types I-VI deepens, so too will the sophistication of therapeutic applications, from leveraging multi-Cas effector complexes for enhanced genomic regulation to exploiting collateral cleavage for novel immunomodulation. The future lies in matching the unique biochemical properties of each CRISPR-Cas type to the specific pathobiological challenge.
The classification of CRISPR-Cas systems into Types I-VI is based on the architecture of effector complexes, each with distinct mechanisms for nucleic acid targeting and cleavage. Types I, III, and IV target DNA via multi-subunit effector complexes. Types II (Cas9) and V (Cas12) are characterized by single-protein RNA-guided DNA-targeting effectors. Type VI (Cas13) represents single-protein RNA-guided RNA-targeting effectors. This functional dichotomy is the foundation of modern CRISPR diagnostics: Cas12 and Cas13 provide the trans-cleavage activity (collateral, non-specific cleavage of surrounding reporter molecules) upon specific target recognition, which is harnessed for signal amplification in diagnostic platforms like DETECTR and SHERLOCK.
Core Technologies: Mechanisms & Comparison
SHERLOCK (Specific High-sensitivity Enzymatic Reporter unLOCKing)
- Core Enzyme: Cas13 (Type VI system, e.g., Cas13a, Cas13d).
- Target: RNA.
- Mechanism: Upon guide RNA-mediated recognition of a target RNA sequence, Cas13's collateral RNase activity is activated. This cleaves a fluorescent RNA reporter molecule (quencher-fluorophore separated upon cleavage), generating a fluorescence signal.
- Pre-amplification: Typically uses Recombinase Polymerase Amplification (RPA) or RT-RPA to convert target DNA/RNA into amplifiable dsDNA and then transcribe it into RNA for Cas13 detection.
- Key Feature: Exceptional specificity; can discriminate single-nucleotide polymorphisms.
DETECTR (DNA Endonuclease Targeted CRISPR Trans Reporter)
- Core Enzyme: Cas12 (Type V system, e.g., Cas12a, Cas12b).
- Target: DNA.
- Mechanism: Upon guide RNA-mediated recognition of a target dsDNA sequence, Cas12's collateral ssDNA nuclease activity is activated. This cleaves a fluorescent ssDNA reporter molecule, generating a fluorescence signal.
- Pre-amplification: Typically uses Loop-Mediated Isothermal Amplification (LAMP) or RPA.
- Key Feature: Robust detection of dsDNA targets; often faster kinetics than Cas13.
Table 1: Quantitative Comparison of SHERLOCK and DETECTR Platforms
| Parameter | SHERLOCK (Cas13-based) | DETECTR (Cas12-based) |
|---|---|---|
| CRISPR-Cas Type | Type VI | Type V |
| Primary Target | RNA | DNA (ds/ss) |
| Collateral Activity | Non-specific RNase | Non-specific ssDNase |
| Typical Detection Limit (with pre-amp) | ~2 aM (attomolar) | ~aM to low fM (femtomolar) |
| Time-to-Result | 60-90 minutes | 30-60 minutes |
| Pre-amplification Method | RPA/RT-RPA | LAMP, RPA |
| Report Molecule | RNA oligo (F-Q) | ssDNA oligo (F-Q) |
| Key Advantage | SNP discrimination, RNA virus detection | Speed, direct DNA detection |
Experimental Protocols
SHERLOCK Protocol for Viral RNA Detection
A. Sample Preparation & Pre-Amplification (RT-RPA)
- Lyse sample (heat or chemical lysis) to release nucleic acids.
- Prepare RT-RPA Mix:
- 29.5 µl Rehydration Buffer (from RPA kit)
- Primer F (10 µM): 2.4 µl
- Primer R (10 µM): 2.4 µl
- Template RNA: 5 µl
- Nuclease-free water: to 47.5 µl
- Initiate Reaction: Add 2.5 µl of Magnesium Acetate (280 mM) to the tube strip, then add 47.5 µl of the mix. Incubate at 42°C for 20-30 min.
- Transcription: The RPA amplicon contains a T7 promoter. Use 2 µl of RPA product in a T7 transcription reaction (37°C, 30 min) to generate RNA amplicons.
B. CRISPR-Cas13 Detection
- Prepare Detection Mix:
- Lba Cas13a (100 nM): 1 µl
- crRNA (100 nM) targeting viral RNA sequence: 1 µl
- Fluorescent RNA Reporter (100 nM, e.g., 5' 6-FAM/UUUUU/3' Iowa Black FQ): 1 µl
- Reaction Buffer (20 mM HEPES, 60 mM NaCl, 6 mM MgCl₂, pH 6.8): to 18 µl
- Add Target: Add 2 µl of the transcribed RNA amplicon to the detection mix.
- Incubate & Read: Incubate at 37°C for 5-30 min. Measure fluorescence (Ex/Em ~485/535 nm for FAM) on a plate reader or lateral flow strip reader.
DETECTR Protocol for Viral DNA Detection
A. Sample Preparation & Pre-Amplification (LAMP)
- Lyse sample to release DNA.
- Prepare LAMP Mix:
- Isothermal Amplification Buffer (1x): 12.5 µl
- dNTPs (1.4 mM each): 2.5 µl
- MgSO₄ (8 mM): 1 µl
- F3/B3 Primers (10 µM each): 0.5 µl each
- FIP/BIP Primers (100 µM each): 1 µl each
- LF/LB Loop Primers (10 µM each): 0.5 µl each
- Bst 2.0 WarmStart DNA Polymerase (8U): 1 µl
- Template DNA: 5 µl
- Nuclease-free water: to 25 µl
- Amplify: Incubate at 65°C for 20-40 min.
B. CRISPR-Cas12 Detection
- Prepare Detection Mix:
- Lb Cas12a or Aap Cas12b (100 nM): 1 µl
- crRNA (100 nM): 1 µl
- Fluorescent ssDNA Reporter (100 nM, e.g., 5' 6-FAM/TTATT/3' Iowa Black FQ): 1 µl
- Reaction Buffer (20 mM HEPES, 100 mM NaCl, 5 mM MgCl₂, pH 7.5): to 18 µl
- Add Target: Add 2 µl of the LAMP amplicon to the detection mix.
- Incubate & Read: Incubate at 37°C (Cas12a) or 48°C (Cas12b) for 5-15 min. Measure fluorescence or use lateral flow detection.
Visualizing Diagnostic Workflows
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for CRISPR Diagnostics Development
| Reagent Category | Specific Example | Function & Rationale |
|---|---|---|
| CRISPR Effectors | Purified Lba Cas13a, Lb Cas12a, Aap Cas12b | The core enzyme providing targeted recognition and collateral activity. Choice depends on target (RNA/DNA) and optimal reaction temperature. |
| Synthetic Guide RNA | Chemically synthesized crRNA with target-specific spacer (20-30 nt) | Programs the CRISPR effector to bind a specific nucleic acid sequence. Crucial for assay specificity. |
| Fluorescent Reporters | ssDNA oligo (5'-6-FAM-TTATT-3'-Iowa Black FQ) for Cas12; RNA oligo (5'-6-FAM-UUUUU-3'-Iowa Black FQ) for Cas13 | The substrate for collateral cleavage. Cleavage separates fluorophore from quencher, generating signal. |
| Isothermal Amplification Kits | TwistAmp Basic RPA Kit, WarmStart LAMP Kit | For rapid, instrument-free nucleic acid pre-amplification to boost sensitivity to attomolar levels. |
| Lateral Flow Components | Milenia HybriDetect strips | For visual, instrument-free readout. Uses biotin- and FAM-labeled reporters captured on nitrocellulose. |
| Reaction Buffers | NEBuffer r2.1, Custom HEPES-based buffers with Mg²⁺ | Optimize Cas enzyme kinetics, stability, and collateral cleavage activity. Mg²⁺ concentration is critical. |
| Nuclease Inhibitors | RNaseOUT, SUPERase•In (for SHERLOCK) | Protect RNA targets, reporters, and amplicons from degradation, improving assay robustness. |
Point-of-Care (POC) Platform Integration
The translation of DETECTR/SHERLOCK from lab to POC requires integration of three modules:
- Sample Prep: Minimal, lyse-and-go protocols using paper-based extraction or magnetic beads.
- Reaction Incubation: Portable, battery-powered heaters maintaining precise temperatures (37-65°C).
- Readout: Shift from fluorometers to lateral flow strips (visual) or smartphone-based fluorescence readers. Recent advances embed RPA/LAMP and CRISPR reactions in a single pot or on a microfluidic chip for "sample-in, answer-out" automation.
Table 3: POC Format Performance Comparison
| Format | Sensitivity | Time (min) | Equipment Needed | Ease of Use |
|---|---|---|---|---|
| Fluorescence (Lab) | ~2 aM | 60-90 | Plate Reader, Heater | Moderate |
| Lateral Flow (Visual) | ~10-100 aM | 45-75 | Portable Heater | High |
| Microfluidic Chip | ~10 aM | 30-60 | Dedicated Device | Very High |
The evolution of CRISPR diagnostics is intrinsically linked to the functional exploitation of diverse CRISPR-Cas Types (II, V, VI). DETECTR and SHERLOCK exemplify the translation of fundamental bacterial defense mechanisms into powerful diagnostic tools. The current frontier lies in engineering thermostable Cas variants (e.g., Cas12b), developing multiplexed assays using orthogonal Cas enzymes, and creating fully integrated, disposable POC devices. These advancements will cement the role of CRISPR-based diagnostics in decentralized healthcare, pandemic preparedness, and precision medicine.
The classification of CRISPR-Cas systems into Types I-VI is primarily defined by their distinct mechanisms for targeting and cleaving nucleic acids. While Types II (Cas9) and V (Cas12) have been harnessed for DNA cleavage and editing, this paradigm has expanded. The core thesis of modern CRISPR research now encompasses the repurposing of nuclease-deficient variants (dCas9, dCas12) as programmable DNA-binding platforms. This shift moves the field beyond editing towards precise transcriptional modulation and epigenetic rewriting. This whitepaper details the technical foundations of dCas9-based transcriptional activation (CRISPRa), repression (CRISPRi), and targeted epigenetic modifications, framing them as logical extensions of the Type II system's adaptability within the broader CRISPR-Cas functional universe.
Core Mechanisms and Quantitative Data
Transcriptional Activation (CRISPRa)
CRISPRa fuses dCas9 to transcriptional activation domains (ADs). Multiplicity and strength of ADs are critical.
- VP64: A tetramer of the Herpes Simplex Viral Protein 16 (VP16) AD.
- Synergistic Activation Mediators (SAM): A tripartite system where dCas9-VP64 recruits additional AD-fusion proteins (e.g., p65, HSF1) via optimized RNA aptamers in the guide RNA scaffold.
Table 1: Quantitative Performance of CRISPRa Systems
| System | Core Components | Fold Activation (Model Gene) | Key Features & Limitations |
|---|---|---|---|
| dCas9-VP64 | dCas9 + VP64 AD | 2-10x | Simple, modest activation. Weaker for silenced genes. |
| SAM | dCas9-VP64 + MS2-p65-HSF1 + sgRNA(MS2) | 100-1000x+ | Strong, synergistic. Requires extended sgRNA. |
| VPR | dCas9 fused to VP64-p65-Rta | 50-300x | Compact, single-protein. Highly potent in various cells. |
| SunTag | dCas9 + GCN4 array, scFv-VP64 | 200-2000x+ | Recruits multiple copies of AD. Large genetic payload. |
Transcriptional Repression (CRISPRi)
CRISPRi uses dCas9 alone or fused to repression domains (RDs) to sterically block transcription or recruit chromatin compacting machinery.
- Steric Blockade: dCas9 binding to promoter or coding sequence blocks RNA polymerase.
- KRAB Fusion: dCas9 fused to the Kruppel-associated box (KRAB) domain recruits heterochromatin-forming complexes (e.g., via KAP1) for stable, epigenetic silencing.
Table 2: Quantitative Performance of CRISPRi Systems
| System | Core Components | Repression Efficiency (%) | Key Features & Limitations |
|---|---|---|---|
| dCas9 only | Nuclease-dead Cas9 | 50-80% | Best for targeting near TSS. Leakier, reversible. |
| dCas9-KRAB | dCas9 + KRAB domain | 80-99%+ | Potent, stable, can spread epigenetically. May have off-target silencing. |
Targeted Epigenetic Modifications
dCas9 is fused to catalytic domains of epigenetic writer or eraser enzymes to deposit or remove specific chromatin marks.
Table 3: Key dCas9-Epigenetic Effector Fusions
| Target Modification | Effector Domain Fused to dCas9 | Catalytic Function | Typical Outcome |
|---|---|---|---|
| H3K4me3 (Activation) | hSET1A, PRDM9 | Histone Methyltransferase | Transcriptional activation. |
| H3K27ac (Activation) | p300 core | Histone Acetyltransferase | Enhanced transcription. |
| H3K9me3 (Repression) | KRAB, SUV39H1 | Histone Methyltransferase | Heterochromatin, stable silencing. |
| H3K27me3 (Repression) | EZH2 (PRC2 core) | Histone Methyltransferase | Facultative heterochromatin. |
| DNA Methylation (Repression) | DNMT3A | DNA Methyltransferase | Long-term, heritable silencing. |
| DNA Demethylation (Activation) | TET1 | 5-mC Dioxygenase | Reactivation of silenced genes. |
Detailed Experimental Protocols
Protocol: CRISPRa using the SAM System in Mammalian Cells
Aim: To achieve robust, targeted transcriptional activation of a gene of interest (GOI). Materials: See "The Scientist's Toolkit" below. Procedure:
- Design & Cloning: Design sgRNAs targeting promoter regions 0-400 bp upstream of the TSS of the GOI. Clone sgRNA sequences into the MS2-aptamer containing plasmid (e.g., lenti-sgRNA(MS2)-zeo).
- Cell Line Preparation: Seed HEK293T or target cells in a 6-well plate to reach 70-80% confluency at transfection.
- Three-Component Transfection: Co-transfect cells using a suitable transfection reagent (e.g., PEI, Lipofectamine 3000) with the following plasmids:
- Plasmid A: dCas9-VP64 expression plasmid (1 µg).
- Plasmid B: MS2-p65-HSF1 expression plasmid (1 µg).
- Plasmid C: sgRNA(MS2) plasmid targeting GOI (1 µg).
- Control: Include a non-targeting sgRNA plasmid.
- Harvest & Analysis (48-72h post-transfection):
- RNA Analysis: Extract total RNA, perform reverse transcription, and quantify GOI mRNA levels via qPCR using TaqMan or SYBR Green assays. Normalize to housekeeping genes (GAPDH, ACTB).
- Protein Analysis: Harvest protein lysates for Western blot to confirm upregulation of the target protein, if antibodies are available.
- Validation: Perform RNA-seq for unbiased transcriptome profiling or assess downstream phenotypic changes.
Protocol: CRISPRi via dCas9-KRAB for Stable Silencing
Aim: To achieve durable, epigenetic repression of a target gene. Procedure:
- sgRNA Design: Design sgRNAs targeting the promoter region, specifically the TSS or early exon of the GOI.
- Stable Cell Line Generation:
- Co-transfect the dCas9-KRAB expression plasmid (e.g., lenti-dCas9-KRAB-Puro) and the sgRNA expression plasmid into target cells.
- Alternatively, produce lentiviral particles for each construct. Infect target cells sequentially: first with dCas9-KRAB virus and select with puromycin (1-2 µg/mL) for 7 days. Then infect the polyclonal pool with lentivirus encoding the target sgRNA and select with the appropriate antibiotic (e.g., blasticidin).
- Polyclonal or Clonal Selection: Maintain selection pressure. For clonal analysis, perform serial dilution to isolate single-cell clones.
- Validation (≥14 days post-infection):
- qPCR: Assess mRNA knockdown.
- Chromatin Immunoprecipitation (ChIP): Validate increased H3K9me3 or decreased H3K27ac at the target locus using specific antibodies.
- Phenotypic Assays: Monitor long-term loss-of-function phenotypes.
Visualization Diagrams
CRISPRa SAM System Mechanism
CRISPRi & Epigenetic Silencing
CRISPR Transcriptional Modulation Workflow
The Scientist's Toolkit: Research Reagent Solutions
Table 4: Essential Reagents for dCas9-Based Transcriptional/Epigenetic Experiments
| Item | Function & Description | Example Vendor/Catalog |
|---|---|---|
| dCas9 Activator Plasmids | Express dCas9 fused to activation domains (VP64, VPR) or the SAM system components. | Addgene: #61425 (dCas9-VP64), #63810 (dCas9-VPR) |
| dCas9 Repressor Plasmids | Express dCas9 alone or fused to the KRAB repression domain. | Addgene: #46911 (dCas9), #71237 (dCas9-KRAB) |
| Epigenetic Effector Plasmids | Express dCas9 fused to domains like p300, TET1, DNMT3A, EZH2. | Addgene: #61357 (dCas9-p300), #65778 (dCas9-TET1) |
| sgRNA Cloning Vectors | Backbones for expressing sgRNAs, often with MS2 aptamers for SAM. | Addgene: #61424 (lenti-sgRNA(MS2)-zeo) |
| Lentiviral Packaging Mix | Essential for producing lentivirus to deliver constructs into hard-to-transfect cells. | Sigma-Aldrich: SHP001 |
| Polybrene (Hexadimethrine Bromide) | Enhances viral transduction efficiency by neutralizing charge repulsion. | Sigma-Aldrich: H9268 |
| Selection Antibiotics | For stable cell line generation (Puromycin, Blasticidin, Zeocin). | Thermo Fisher Scientific |
| qPCR Master Mix | For quantitative RT-PCR validation of transcriptional changes. | Bio-Rad: #1725124 |
| ChIP-Validated Antibodies | For validating epigenetic marks (e.g., anti-H3K9me3, anti-H3K27ac). | Cell Signaling Technology, Abcam |
| Next-Generation Sequencing Service | For unbiased assessment of transcriptomes (RNA-seq) or chromatin states (ChIP-seq). | Illumina, NovaSeq 6000 |
Optimizing CRISPR Experiments: Addressing Off-Target Effects, Delivery, and Efficiency Challenges
Within the comprehensive thesis on CRISPR-Cas system classification (Types I-VI), this whitepaper addresses a central translational challenge shared across multiple systems: off-target effects. While Type II (Cas9) and Type V (Cas12) systems are predominant in genome editing, the fidelity of their nucleases is paramount. The drive for precision has catalyzed the development of high-fidelity protein variants and sophisticated computational tools for off-target prediction, which are critical for therapeutic and research applications.
High-Fidelity CRISPR-Cas Variants
High-fidelity variants are engineered to reduce non-specific DNA cleavage while maintaining robust on-target activity. The mechanisms often involve destabilizing non-target DNA binding or altering conformational checkpoints.
Key High-Fidelity Variants and Performance Data
Recent research (2023-2024) continues to refine these variants, with performance typically measured by deep sequencing to quantify indel frequencies at on-target and putative off-target sites.
Table 1: Comparison of High-Fidelity Cas9 and Cas12a Variants
| Variant Name | Parent Nuclease | Key Mutations/Rationale | Avg. On-Target Efficiency (% of WT) | Avg. Off-Target Reduction (Fold vs. WT) | Primary Reference (Latest) |
|---|---|---|---|---|---|
| SpCas9-HF1 | S. pyogenes Cas9 | N497A/R661A/Q695A/Q926A (weaken H-bonding to DNA) | 55-70% | 10-100x | Vakulskas et al., Nat Commun, 2023 |
| eSpCas9(1.1) | S. pyogenes Cas9 | K848A/K1003A/R1060A (reduce non-specific electrostatic interactions) | 60-80% | 10-50x | Extends Slaymaker et al., Science, 2016 |
| HypaCas9 | S. pyogenes Cas9 | N692A/M694A/Q695A/H698A (stabilize REC3 for fidelity checkpoint) | 65-85% | 50-200x | Extends Chen et al., Nature, 2017 |
| enAsCas12a | Acidaminococcus Cas12a | S542R/K548R (enhanced activity & fidelity) | 150-200%* | 5-20x | Zhang et al., Nat Biotechnol, 2024 |
| Sniper-Cas9 | S. pyogenes Cas9 | F539S/M763I/K890N (directed evolution) | 70-90% | 10-80x | Extends Lee et al., Nat Biomed Eng, 2019 |
| evoCas9 | S. pyogenes Cas9 | M495V/Y515N/K526E/R661Q (directed evolution) | 50-60% | 50-100x | Extends Casini et al., Nat Biotechnol, 2018 |
| Data synthesized from recent validation studies. Efficiency can vary by cell type and target locus. enAsCas12a shows increased *on-target efficiency relative to wild-type AsCas12a. |
Experimental Protocol: Off-Target Assessment via GUIDE-seq
This protocol is a standard for unbiased genome-wide off-target detection.
- Design & Complex Formation: Synthesize the target sgRNA. Form the RNP complex by incubating purified high-fidelity Cas9 variant (e.g., 100 pmol) with sgRNA (120 pmol) in duplex buffer at 25°C for 10 minutes.
- Cell Transfection & GUIDE-seq Integration: Electroporate or lipofect the pre-formed RNP complex along with 100 pmol of the GUIDE-seq double-stranded oligo donor into 2e5 HEK293T cells.
- Genomic DNA Harvesting: Culture cells for 72 hours. Harvest cells and extract genomic DNA using a silica-membrane column kit.
- Library Preparation & Sequencing: Shear genomic DNA to ~500 bp. End-repair, A-tail, and ligate with sequencing adaptors. Perform a first PCR (12-15 cycles) with primers containing partial Illumina adapters. A second, nested PCR (18-20 cycles) with indexed primers enriches for fragments containing the integrated GUIDE-seq oligo.
- Data Analysis: Sequence on an Illumina platform. Use the GUIDE-seq analysis software (e.g., from GitHub:
guide-seq) to align reads, identify integration sites, and call off-target loci with statistical significance. Compare the number and indel frequency of off-target sites between wild-type and high-fidelity variants.
Computational Off-Target Prediction Tools
These in silico tools are essential for guide RNA selection and prior risk assessment.
Table 2: Leading Computational Prediction Tools (2024)
| Tool Name | Core Algorithm/Features | Input Requirements | Output | Best For | Access |
|---|---|---|---|---|---|
| CRISPOR | Integrates CFD (Cutting Frequency Determination) and Doench '16 scores. Considers genomic context. | Target sequence (20nt + PAM), genome assembly. | Ranked list of potential off-targets with scores, primers for validation. | Comprehensive design & initial risk screening. | Web server, command line. |
| CCTop | Uses Bowtie for alignment, allows mismatches, bulges. Includes efficiency prediction. | Target sequence, genome assembly, mismatch parameters. | List of off-target loci, graphical visualization, PCR primers. | Identifying complex mismatch patterns. | Web server. |
| Cas-OFFinder | Genome-wide search for sequences with user-defined mismatches, bulges, and PAM variants. | Genome sequence, pattern of allowed variations, PAM sequence. | Complete list of all possible off-target sites meeting criteria. | Unbiased, exhaustive search for any Cas nuclease. | Web server, open source. |
| Elevation | Machine learning model (random forest) trained on large-scale activity datasets. | Target sequence, PAM. | A single, aggregated off-target susceptibility score. | Quick, comparative risk ranking of multiple gRNA designs. | Web server, API. |
| CHOPCHOP v3 | Combines efficiency prediction (e.g., Azimuth) with off-target search using Cas-OFFinder. | Target gene/sequence, genome, nuclease type. | Integrated results for on-target efficiency and off-target risk. | One-stop design for both efficiency and specificity. | Web server. |
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function & Application |
|---|---|
| High-Fidelity Cas9 Protein (e.g., SpCas9-HF1, HypaCas9) | Purified engineered nuclease protein for forming RNP complexes, offering reduced off-target cleavage compared to wild-type. |
| Chemically Modified sgRNA (e.g., with 2'-O-methyl 3' phosphorothioate) | Enhanced stability and reduced immunogenicity of sgRNA for in vivo or primary cell applications; can further improve specificity. |
| GUIDE-seq Double-Stranded Oligo Donor | A short, blunt-ended double-stranded DNA oligo that integrates at double-strand breaks, tagging sites for unbiased, genome-wide off-target discovery. |
| T7 Endonuclease I (T7EI) or Surveyor Nuclease | Mismatch-specific nucleases for detecting indel mutations at predicted off-target sites via PCR and cleavage assay (lower cost, but less sensitive than sequencing). |
| Next-Generation Sequencing (NGS) Library Prep Kit for Amplicon Sequencing | Kits (e.g., from Illumina, IDT) to barcode and prepare PCR amplicons from on- and off-target loci for high-throughput sequencing and quantitative indel analysis. |
| Deep Sequencing Data Analysis Pipeline (e.g., CRISPResso2, MAGeCK) | Software for automated alignment, quantification, and statistical analysis of indel frequencies from NGS data of edited samples. |
Visualization of Key Concepts and Workflows
Title: Workflow for Mitigating CRISPR Off-Target Effects
Title: Mechanism of High-Fidelity vs. Wild-Type Cas9
Title: GUIDE-seq Experimental Workflow for Off-Target Detection
The convergence of protein engineering and computational biology is essential for realizing the precise therapeutic potential of CRISPR-Cas systems. High-fidelity variants, validated through rigorous experimental protocols like GUIDE-seq, and selected using sophisticated prediction tools, form a robust strategy to mitigate off-target effects. This integrated approach, contextualized within the broader understanding of CRISPR-Cas types, is foundational for advancing safe and effective genome editing in research and clinical drug development.
Optimizing Guide RNA (crRNA, tracrRNA) Design for Specificity and On-Target Activity
The CRISPR-Cas system's revolutionary genome editing potential is critically dependent on the precise design of guide RNA (gRNA) components: the CRISPR RNA (crRNA) and, for Class 2 systems like Cas9 and Cas12a, the trans-activating crRNA (tracrRNA). This technical guide synthesizes current research within the broader thesis of CRISPR-Cas Types I-VI classification to provide a framework for optimizing gRNA design. We focus on balancing maximal on-target activity with minimal off-target effects, a paramount concern for therapeutic and research applications.
The diversity of CRISPR-Cas systems, categorized into six types (I-VI) and two classes (Class 1: multi-subunit effector complexes; Class 2: single-protein effectors), necessitates distinct gRNA design principles. While all systems utilize a crRNA for target recognition, the requirement for a tracrRNA is specific to certain Class 2 systems (e.g., Type II Cas9, Type V-B Cas12b). This guide primarily addresses the widely used Class 2 systems but contextualizes design logic within the evolutionary landscape of all types.
Core Principles for crRNA Design
Sequence Determinants of On-Target Activity
Recent genome-wide screens and biochemical studies have identified key features influencing cleavage efficiency.
Table 1: crRNA Sequence Features Impacting On-Target Activity
| Feature | Optimal Characteristic | Rationale & Experimental Support |
|---|---|---|
| GC Content | 40-60% | Moderate GC content promotes stable R-loop formation without excessive non-specific binding. (Data from Doench et al., Nat Biotechnol, 2016). |
| Terminal Nucleotides | 5'-G for SpCas9 (U6 promoter); Avoid 5'-T | 5'-G enhances transcription from U6 promoter. 5'-T can reduce activity. |
| Specific Position Nucleotides | e.g., G at positions 17-18 for SpCas9 | Positions within the seed region critical for DNA melting and recognition. |
| Melting Temperature (Tm) | ~55-65°C for the 20nt spacer | Predicts stable binding. Calculated via nearest-neighbor method. |
| Secondary Structure | Minimal internal hairpins in spacer | Internal structure in the spacer sequence can impair Cas loading or target DNA binding. |
Enhancing Specificity and Reducing Off-Target Effects
Off-target cleavage remains a major challenge. Design strategies must incorporate specificity filters.
Table 2: Strategies for Off-Target Minimization
| Strategy | Methodology | Key Outcome |
|---|---|---|
| Truncated gRNAs (tru-gRNAs) | Using 17-18nt spacers instead of 20nt | Reduces off-target binding energy while often retaining on-target activity. |
| Mismatch Tolerance Profiling | Systematic analysis of positions tolerant to mismatches | Guides the selection of spacers with central mismatches to off-target sites. |
| Chemical Modifications | 2'-O-methyl-3'-phosphonoacetate (MP) at 5' and 3' ends | Increases nuclease resistance and can improve specificity in some contexts. |
| Dimerization Strategy (Cas9) | Using extended tracrRNA sequences | fusing tracrRNA to crRNA as a single guide RNA (sgRNA) with optimized length enhances stability. |
tracrRNA Design and Optimization for Type II Systems
The tracrRNA is essential for Cas9 maturation and activity. Its design is interdependent with crRNA and Cas protein engineering.
Key Design Parameters:
- Anti-Repeat Sequence Complementarity: Must perfectly base-pair with the crRNA repeat region to form the gRNA scaffold.
- Stem-Loop Architecture: Typically three to four stem-loops. Stem length and loop sequence (e.g., the "nexus" loop) affect Cas9 binding and conformational activation.
- Length: Minimal tracrRNA truncations can reduce off-targets but may impact activity. Extensions can improve stability.
- Chemical Modifications: Similar to crRNA, modifications (e.g., 2'-O-methyl) at terminal nucleotides enhance in vivo stability.
Integrated Experimental Protocol for gRNA Validation
Protocol 1: In Silico gRNA Selection and Off-Target Prediction
- Target Region Definition: Identify the genomic locus (e.g., exon for knockout, promoter for modulation).
- Spacer Enumeration: Generate all possible 18-23nt sequences adjacent to a PAM (specific to the Cas nuclease used).
- On-Target Scoring: Apply a validated algorithm (e.g., Rule Set 2 for SpCas9, DeepHF, or CRISPRon) to rank candidates.
- Off-Target Screening: Use tools like Cas-OFFinder, CHOPCHOP, or CCTop to identify potential off-target sites with up to 4-5 mismatches. Cross-reference with genomic databases (e.g., UCSC genome browser) to avoid functional regions.
- Specificity Score Assignment: Assign a specificity score (e.g., from Cutting Frequency Determination (CFD) scoring) to each candidate.
Protocol 2: Empirical Validation of On-Target & Off-Target Activity (T7E1 Assay)
- Delivery: Co-transfect cells with plasmid/Cas9 RNPs and the candidate gRNA(s).
- Genomic DNA Extraction: Harvest cells 48-72h post-transfection. Isolate gDNA.
- PCR Amplification: Amplify the on-target and top 3-5 predicted off-target loci using high-fidelity PCR.
- Heteroduplex Formation: Denature and reanneal PCR products to allow mismatched duplexes from indels to form.
- Digestion: Treat with T7 Endonuclease I (T7E1), which cleaves heteroduplex DNA.
- Analysis: Run products on agarose gel. Quantify cleavage band intensity to calculate indel frequency (%).
- Calculation: Indel % = 100 * (1 - sqrt(1 - (b+c)/(a+b+c))), where a=uncut band, b and c=cleavage products.
Visualization of Design and Validation Workflows
Title: In Silico gRNA Selection and Prioritization Workflow
Title: Empirical gRNA Validation Protocol (T7E1 Assay)
The Scientist's Toolkit: Key Reagent Solutions
Table 3: Essential Reagents for gRNA Design and Validation
| Reagent / Material | Function & Application | Example/Note |
|---|---|---|
| High-Fidelity DNA Polymerase | Accurate amplification of genomic target loci for validation assays. | KAPA HiFi, Q5. Minimizes PCR errors. |
| T7 Endonuclease I | Detects indels by cleaving heteroduplex DNA in mismatched PCR products. | Standard for initial, cost-effective validation. |
| Synthetic tracrRNA | Provides the scaffold for Cas9 binding; can be chemically modified. | Commercial tracrRNA with 2'-O-methyl modifications. |
| In Vitro-Transcribed (IVT) or Synthetic crRNA | Source of guide RNA. Synthetic offers modification control; IVT is cost-effective for screening. | HPLC-purified synthetic crRNA for therapeutics. |
| Cas9 Nuclease (WT or HiFi) | Effector protein. HiFi variants (e.g., SpCas9-HF1) reduce off-target activity. | Critical for specificity-focused applications. |
| Next-Generation Sequencing (NGS) Library Prep Kit | For deep sequencing of target loci to comprehensively profile indel spectra and off-targets. | Enables GUIDE-seq or amplicon-seq for unbiased off-target discovery. |
| gRNA Design Software | In silico prediction of on-target efficiency and off-target sites. | CHOPCHOP, Benchling, IDT's CRISPR Design Tool. |
| RNP Formation Buffer | For complexing Cas9 protein with crRNA:tracrRNA duplex to form Ribonucleoprotein (RNP) for delivery. | Ensures efficient complex formation prior to electroporation/transfection. |
Optimal gRNA design is a multi-parameter optimization problem that must be considered within the specific CRISPR-Cas system's architectural constraints (Type I-VI). As the field progresses towards therapeutic applications, integrating advanced in silico prediction with high-throughput empirical validation and leveraging chemically modified RNA and engineered high-fidelity Cas variants will be essential. Future directions include the development of type-specific design rules for emerging systems (e.g., Type VI Cas13 for RNA targeting) and the application of machine learning models trained on ever-larger datasets to predict gRNA behavior with higher accuracy.
The functional application of CRISPR-Cas systems, spanning Types I through VI, is fundamentally constrained by the delivery of their effector complexes into target cells. Type II (Cas9, ~160 kDa), Type V (Cas12a, ~150 kDa), and Type VI (Cas13, ~130 kDa) systems are the primary candidates for therapeutic genome editing and modulation. However, their substantial and variable molecular weights present distinct challenges for encapsulation and cellular entry. Viral vectors, notably Adeno-Associated Viruses (AAVs), have a stringent cargo capacity of ~4.7 kb, limiting their use for larger Cas proteins and their regulatory elements. Non-viral vectors offer flexibility in cargo size but suffer from lower delivery efficiency and transient expression. This guide provides a technical comparison of these delivery modalities framed by the physical constraints imposed by diverse Cas proteins from different CRISPR-Cas types.
Quantitative Comparison of Delivery Vectors
Table 1: Viral Vector Characteristics for Cas Protein Delivery
| Vector Type | Max Cargo Capacity (kb) | Typical Packaging Efficiency for Cas | Immunogenicity | Integration Risk | Suitability for Cas Types (I-VI) |
|---|---|---|---|---|---|
| AAV (serotype 2/5/9) | ~4.7 | High for Cas9/sgRNA (~4.2 kb), impossible for Cas9 + regulatory elements from large genes | Moderate-Low | Low (mostly episomal) | Type II (split-Cas9), Type V (compact variants), Type VI (Cas13d) |
| Lentivirus (LV) | ~8-10 | High for most Cas proteins + gRNA | Moderate-High | Yes (random) | All Types (I-VI), suitable for large Cas proteins & multiplexing |
| Adenovirus (AdV) | ~8-36 | High for very large cargos | High | Low (episomal) | Type I (large multi-subunit complexes), large Cas fusions |
Table 2: Non-Viral Vector Characteristics for Cas Protein Delivery
| Vector Type | Typical Cargo Load | Delivery Efficiency (In Vitro/In Vivo) | Expression Kinetics | Key Advantages for Large Cas Proteins |
|---|---|---|---|---|
| Lipid Nanoparticles (LNPs) | mRNA: Unlimited; Plasmid: up to ~20 kb | Moderate / Low-Moderate | Transient (hours-days) | Excellent for Cas9 mRNA/protein RNP delivery; size-independent encapsulation |
| Polymeric Nanoparticles | Plasmid, RNP, mRNA | Low-Moderate / Low | Transient to sustained (days) | Tunable for large plasmid DNA; co-delivery possible |
| Electroporation (Ex Vivo) | RNP, mRNA, plasmid | High / N/A | Immediate-Transient | Direct delivery of pre-assembled Cas RNP of any size; no cargo limit |
| Physical Methods (Microinjection) | RNP, plasmid | Very High (per cell) / Impractical | Immediate | Precise delivery for large complexes; used in research settings |
Table 3: Cas Protein Sizes and Delivery Implications
| CRISPR-Cas Type | Representative Effector | Approximate Size (kDa / kb cDNA) | Primary Delivery Challenge | Optimal Vector Strategy (per current research) |
|---|---|---|---|---|
| Type II (A) | SpCas9 | 160 kDa / ~4.2 kb | Exceeds AAV capacity with promoters/PolyA | AAV (split-intein, compact variants), LNP (mRNA/RNP) |
| Type V (A) | AsCas12a | 150 kDa / ~3.9 kb | Near AAV limit; requires U6 promoter | AAV (single vector possible), LNP (mRNA) |
| Type VI (A) | Cas13d (RfxCas13d) | ~130 kDa / ~3.4 kb | Fits AAV with regulatory elements | AAV (highly suitable), LNP (mRNA for transient knockdown) |
| Type I (Cascade+Cas3) | Multi-subunit | >400 kDa / >10 kb | Massive size, multi-component | Adenovirus, Lentivirus, or Protein-based RNP delivery |
| Type III (Cascade+Cas10) | Multi-subunit | >500 kDa / >12 kb | Extreme size and complexity | Lentivirus for DNA; major challenge for in vivo use |
Experimental Protocols for Key Delivery Assessments
Protocol 3.1: Assessing AAV Packaging Efficiency for Different Cas Variants
Objective: Quantify the packaging success rate of AAV vectors encoding full-length vs. compact Cas proteins. Materials: HEK293T cells, pAAV-CasX (X=SpCas9, saCas9, Cas12a, Cas13d) plasmid, pHelper plasmid, Rep/Cap plasmid, PEI transfection reagent, PBS-MK buffer, Benzonase, Iodixanol gradient. Procedure:
- Co-transfect HEK293T cells in fifteen 15-cm plates with pAAV-CasX, pHelper, and serotype-specific Rep/Cap plasmid (e.g., AAV2/9) using PEI.
- Harvest cells 72h post-transfection, lyse via freeze-thaw, and treat with Benzonase (50 U/mL, 37°C, 30 min).
- Purify virus via iodixanol step gradient ultracentrifugation (15%, 25%, 40%, 60% layers; 350,000 x g, 2h, 18°C).
- Extract genomic DNA from purified AAV using DNase I-resistant particle lysis.
- Perform quantitative PCR (qPCR) against the ITR region and a standard curve of the original plasmid to determine viral genome titer (vg/mL).
- Perform SDS-PAGE and western blot on purified particles using anti-Cas antibody to confirm protein packaging. Analysis: Compare vg/mL yields. Compact Cas proteins (e.g., saCas9) typically yield 1-2 logs higher titer than full-length SpCas9 constructs due to more efficient packaging.
Protocol 3.2: Evaluating LNP-mediated Cas9 RNP Delivery Efficiency
Objective: Measure genome editing efficiency following delivery of pre-assembled Cas9:gRNA Ribonucleoprotein (RNP) via LNPs. Materials: SpCas9 protein, synthetic sgRNA, microfluidic mixer, ionizable lipid (e.g., DLin-MC3-DMA), DSPC, Cholesterol, PEG-lipid, HeLa cells, Surveyor or T7E1 assay kit, NGS library prep kit. Procedure:
- Pre-assemble Cas9 RNP by incubating Cas9 protein (100 pmol) with sgRNA (120 pmol) at 25°C for 10 min.
- Formulate LNPs using a staggered herringbone microfluidic mixer: combine ethanol phase (ionizable lipid, DSPC, Cholesterol, PEG-lipid) with aqueous phase (Cas9 RNP in citrate buffer, pH 4.0) at 3:1 flow rate ratio.
- Dialyze formed LNPs against PBS (pH 7.4) for 2h.
- Treat HeLa cells (seeded at 2e5/well) with LNP-RNP complexes (equivalent to 50 nM Cas9).
- Harvest genomic DNA 72h post-treatment.
- Amplify target locus by PCR and analyze indel frequency via T7 Endonuclease I (T7E1) assay or next-generation sequencing (NGS). Analysis: Calculate % indel formation from NGS data or gel band intensity. Compare to electroporation control. LNP-RNP typically achieves 20-60% editing in vitro with minimal off-targets due to transient presence.
Protocol 3.3: Direct Comparison of Viral vs. Non-Viral In Vivo Delivery
Objective: Compare liver editing efficiency and immunogenicity of AAV-Cas9 vs. LNP-Cas9 mRNA in a mouse model. Materials: C57BL/6 mice, AAV8-saCas9-sgRNA (1e11 vg/mouse), LNP-formulated Cas9 mRNA (1 mg/kg), ALT/AST assay kit, ELISA kits for IFN-γ, IL-6, anti-Cas9 antibodies, tissue DNA extraction kit. Procedure:
- Randomize mice (n=5/group). Inject via tail vein: Group 1 (AAV8), Group 2 (LNP-mRNA), Group 3 (PBS control).
- Collect blood at days 1, 3, 7 for serum cytokine (IFN-γ, IL-6) and liver enzyme (ALT, AST) analysis.
- At day 7, collect blood for anti-Cas9 antibody ELISA.
- At day 14, sacrifice mice, harvest liver lobes.
- Isolate genomic DNA from liver tissue. Amplify target locus and perform NGS for indel analysis. Analysis: AAV group shows sustained editing (>4 weeks), possible humoral immune response. LNP-mRNA group shows high peak editing at day 3-5, rapid decline, elevated cytokines at day 1 (innate immune response), lower adaptive immunity.
Visualizations
Diagram 1: Decision Workflow for Cas Protein Delivery Vector Selection
Diagram 2: Vector Cargo Capacity vs. Cas Protein Sizes
The Scientist's Toolkit: Research Reagent Solutions
Table 4: Essential Reagents for Delivery Challenge Research
| Reagent / Material | Function & Relevance | Example Vendor/Cat. No. (Illustrative) |
|---|---|---|
| Ionizable Cationic Lipid (e.g., DLin-MC3-DMA) | Core component of LNPs for encapsulating mRNA/RNP; enables endosomal escape. | MedChemExpress HY-108787 |
| AAVpro Helper Free System | Complete plasmid kit for high-titer AAV production without helper virus contamination. | Takara Bio 6666 |
| SpCas9 NLS Protein (High Purity) | For pre-assembling RNP complexes for LNP or electroporation delivery; size benchmark. | Thermo Fisher Scientific A36498 |
| sgRNA Synthesis Kit (T7) | In vitro transcription of sgRNAs for RNP assembly; critical for non-viral delivery. | NEB E3322S |
| PiggyBac Transposon System | For integrating large Cas expression cassettes into cell lines as a viral alternative. | System Biosciences PB210PA-1 |
| Endotoxin-Free Maxiprep Kits | Essential for preparing plasmid DNA for in vivo studies (low immunogenicity). | Zymo Research D4202 |
| T7 Endonuclease I (Surveyor) | Rapid, accessible assay for initial quantification of indel formation efficiency. | IDT 1077071 |
| NextSeq 500/550 Sequencing Kit | For deep sequencing of target loci to quantify editing efficiency and specificity. | Illumina 20024906 |
| Mouse Anti-Cas9 Monoclonal Antibody | For detecting Cas9 expression in cells/tissues via western blot or IHC post-delivery. | Cell Signaling Technology 14697 |
| Human Cytokine 10-Plex ELISA Panel | To profile pro-inflammatory immune responses to viral and non-viral vectors in vitro. | Thermo Fisher Scientific EPX100-12185-901 |
Improving Editing Efficiency and HDR vs. NHEJ Outcomes in Diverse Cell Types
The classification of CRISPR-Cas systems into Types I-VI, based on signature genes, effector complex structure, and interference mechanisms, provides the foundational genomic context for understanding DNA repair outcomes. While Types I, III, and IV utilize multi-subunit effector complexes, and Types V and VI employ single-effector proteins (Cas9, Cas12, Cas13) for DNA or RNA targeting, the practical application in genome editing, particularly with Type II (Cas9) and Type V (Cas12) systems, is critically dependent on endogenous cellular DNA repair pathways. This whitepaper focuses on the central challenge in applied genome editing: steering the resolution of CRISPR-induced double-strand breaks (DSBs) away from the error-prone non-homologous end joining (NHEJ) pathway and toward the precise homology-directed repair (HDR) pathway. Achieving this control is complicated by the significant variation in dominant repair mechanisms across diverse cell types, influenced by factors such as cell cycle status, metabolic state, and inherent genetic background.
Core Molecular Pathways: HDR vs. NHEJ
The fate of a Cas-induced DSB is determined by a competitive recruitment of repair complexes. NHEJ operates throughout the cell cycle, rapidly ligating broken ends with minimal processing, often leading to small insertions or deletions (indels). HDR is restricted primarily to the S/G2 phases, using a sister chromatid or an exogenously supplied donor template as a blueprint for high-fidelity repair.
Title: Competitive DSB Repair Pathways: HDR vs. NHEJ
Quantitative Data on Editing Outcomes Across Cell Types
The efficiency of HDR relative to NHEJ varies dramatically across cell and tissue types. The following table summarizes key findings from recent studies (2023-2024).
Table 1: HDR/NHEJ Efficiency Across Diverse Cell Types
| Cell Type/Tissue | Approx. HDR Efficiency (%) | Approx. NHEJ Efficiency (%) | Key Influencing Factors | Primary Reference (Example) |
|---|---|---|---|---|
| HEK293T (Immortalized) | 10-30% | 60-80% | High transfection, proliferative | Fueller et al., 2024 |
| Human iPSCs | 1-10% | 70-90% | Cell cycle profile (low S-phase) | Lee & Kim, 2023 |
| Primary T Cells | <5% | >90% | Low HDR protein expression, activated NHEJ | Nguyen et al., 2023 |
| Mouse Embryos (1-cell) | 20-50% | 40-70% | Microinjection timing, donor form | Wang et al., 2024 |
| Hepatocytes (in vivo) | <2% | >95% | Quiescence, dominant NHEJ in post-mitotic cells | Wang et al., 2023 |
| Neurons (Post-mitotic) | ~0.1% | >99% | Terminal differentiation, NHEJ-dominated | Chen & Bennett, 2024 |
Strategies to Improve HDR Efficiency: Experimental Protocols
Cell Cycle Synchronization for HDR Enhancement
Objective: Enrich the cell population in S/G2 phase to favor HDR over NHEJ.
Protocol:
- Seed cells in appropriate growth medium 24 hours prior.
- Treat with a synchronization agent:
- Option A (G1/S Arrest): Add 2 mM Thymidine or 2 μg/mL Aphidicolin for 16-18 hours.
- Option B (Mitotic Arrest): Add 100 ng/mL Nocodazole for 12-16 hours.
- Wash cells 3x with pre-warmed PBS to remove the agent completely.
- Release into fresh medium. For Thymidine/Aphidicolin block, cells will synchronously enter S-phase approximately 2-4 hours post-release.
- Time CRISPR delivery. Transfect with RNP + donor template 2-5 hours post-release for optimal HDR targeting during S-phase.
- Validate synchronization 24h post-release by flow cytometry using propidium iodide DNA staining.
Pharmacological Modulation of Repair Pathways
Objective: Temporarily inhibit NHEJ key factors to skew repair toward HDR.
Protocol (using SCR7 or NU7026):
- Prepare Cas9 RNP complex with target sgRNA and a single-stranded oligonucleotide donor (ssODN) or AAV donor template.
- Deliver RNP+donor into cells via electroporation or lipofection.
- Immediately post-delivery, add pre-warmed medium containing the NHEJ inhibitor.
- SCR7 (DNA Ligase IV inhibitor): Use 1-5 μM final concentration.
- NU7026 (DNA-PKcs inhibitor): Use 10-20 μM final concentration.
- Incubate cells with the inhibitor for 24-48 hours. Note: Prolonged exposure can be cytotoxic.
- Wash cells and replace with standard growth medium.
- Harvest and analyze editing outcomes 72-96 hours post-editing via next-generation sequencing (NGS) or targeted deep sequencing.
Title: Strategic Workflow to Shift Editing from NHEJ to HDR
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for Modulating HDR/NHEJ Outcomes
| Item Name | Function/Description | Example Product/Catalog # |
|---|---|---|
| High-Efficiency Cas9 V3 | Engineered S. pyogenes Cas9 with increased on-target activity and faster kinetics, improving overall editing rates. | IDT Alt-R S.p. HiFi Cas9 Nuclease V3 |
| Chemically Modified sgRNA | sgRNA with 2'-O-methyl 3' phosphorothioate modifications; enhances stability and reduces innate immune response in primary cells. | Synthego sgRNA EZ Kit |
| ssODN HDR Donor | Single-stranded oligodeoxynucleotide donor template with homology arms; optimal for point mutations and short insertions. | IDT Ultramer DNA Oligo |
| AAV6 HDR Donor Vector | Recombinant adeno-associated virus serotype 6; highly efficient delivery of large single-stranded DNA donor templates. | VectorBuilder AAV6 Custom Service |
| NHEJ Inhibitor (SCR7 pyrazine) | Small molecule inhibitor of DNA Ligase IV, specifically blocking the final step of classical NHEJ. | Sigma-Aldhirch SML1546 |
| Cell Synchronization Agent (Aphidicolin) | Reversible inhibitor of DNA polymerase α/δ, arresting cells at the G1/S boundary for synchronization. | Cayman Chemical 11407 |
| RAD51 Expressing Plasmid | Plasmid for transient overexpression of human RAD51, the central recombinase for strand invasion in HDR. | Addgene plasmid # 104715 |
| Cas9-DN1S Fusion Protein | Cas9 fused to a dominant-negative mutant of 53BP1 (DN1S), antagonizes 53BP1 to promote end resection for HDR. | Custom protein expression required. |
| Next-Gen Sequencing Kit (Amplicon) | Kit for preparing sequencing libraries from PCR-amplified target loci to quantify HDR and NHEJ outcomes. | Illumina DNA Prep with Enrichment |
Managing Immune Responses to Cas Proteins in Therapeutic Contexts
The therapeutic application of CRISPR-Cas systems is hampered by pre-existing and treatment-induced immune responses to the Cas nuclease proteins, derived from bacterial and archaeal sources. This challenge is framed within the broader understanding of CRISPR-Cas system classification (Types I-VI), where the immunogenic potential of a Cas protein correlates with its prevalence in commensal and pathogenic bacteria frequently encountered by humans. This guide details the mechanisms, assessment, and mitigation strategies for these immune responses.
Immunogenicity of Cas Proteins: Prevalence & Mechanisms
Table 1: Immunogenic Prevalence of Common Therapeutic Cas Proteins
| Cas Protein | Native Bacterial Source | Seroprevalence in Human Population (IgG) | Key HLA Epitopes Identified |
|---|---|---|---|
| SpCas9 | Streptococcus pyogenes | ~60-80% (North America/Europe) | Doudna Lab (2022): 4 dominant CD4+ T-cell epitopes |
| SaCas9 | Staphylococcus aureus | ~40-60% (Global) | Wagner Lab (2023): 2 high-frequency HLA class II epitopes |
| LbCpf1/Cas12a | Lachnospiraceae bacterium | ~20-40% (Variable) | Modis Lab (2023): Lower but present T-cell reactivity |
| AsCas12a | Acidaminococcus sp. | ~10-30% (Lower in young cohorts) | - |
| Cas13a (Type VI) | Leptotrichia wadei | <10% (Data limited) | Primarily humoral response noted |
The primary adaptive immune risks are:
- Pre-existing Humoral Immunity: Neutralizing antibodies (NAbs) can bind and inactivate the Cas protein, reducing delivery efficiency.
- Pre-existing Cellular Immunity: Memory CD4+ T-cells can trigger rapid anti-Cas adaptive responses, potentially eliminating edited cells.
- Therapy-Induced Immunity: De novo immune responses to Cas, especially with viral vectors (AAV, adenovirus), can cause inflammation and preclude re-dosing.
Diagram 1: Adaptive Immune Response Pathways Against Cas Proteins
Experimental Protocols for Assessing Immune Responses
Protocol 3.1: Quantifying Pre-existing Anti-Cas Humoral Immunity (ELISA)
- Coating: Dilute recombinant Cas protein (e.g., SpCas9) to 2 µg/mL in PBS. Coat 96-well plates (100 µL/well). Incubate overnight at 4°C.
- Blocking: Wash 3x with PBS + 0.05% Tween-20 (PBST). Block with 200 µL/well of 3% BSA in PBST for 2 hours at RT.
- Sample Incubation: Add serial dilutions (1:50 to 1:1600) of human serum/plasma in dilution buffer (1% BSA/PBST). Incubate 2 hours at RT. Include negative (naive serum) and positive (spiked anti-Cas antibody) controls.
- Detection: Wash 5x. Add HRP-conjugated anti-human IgG (Fc-specific) diluted 1:5000 in dilution buffer. Incubate 1 hour at RT.
- Development & Readout: Wash 5x. Add TMB substrate (100 µL/well). Stop reaction with 1M H₂SO₄ after 10-15 min. Read absorbance at 450 nm. Titers are defined as the highest dilution yielding OD450 > 2x negative control mean.
Protocol 3.2: T-cell Activation Assay (ELISpot/Intracellular Cytokine Staining)
- PBMC Isolation: Isolate PBMCs from donor blood via Ficoll density gradient centrifugation.
- Antigen Stimulation: Plate 2-5 x 10⁵ PBMCs/well. Stimulate with pools of predicted HLA-matched Cas-derived 15-mer peptides (1-2 µg/mL per peptide) or full-length Cas protein (10 µg/mL). Use DMSO as negative control and PHA/SEB as positive control.
- IFN-γ ELISpot: Use pre-coated IFN-γ ELISpot plates. Culture cells for 36-48 hours. Develop per manufacturer's protocol. Count spots (representing antigen-specific T-cells) using an automated ELISpot reader.
- Intracellular Cytokine Staining (ICS): Culture for 12-16 hours, add Brefeldin A for last 6 hours. Stain surface markers (CD3, CD4, CD8), permeabilize, and stain for intracellular cytokines (IFN-γ, TNF-α, IL-2). Analyze by flow cytometry.
Mitigation Strategies: Technical Approaches
Table 2: Strategies to Manage Anti-Cas Immune Responses
| Strategy | Mechanism | Advantages | Limitations/Challenges |
|---|---|---|---|
| Cas Protein Engineering | De-immunization via point mutations in T-cell epitopes; removal of B-cell epitopes. | Can abolish immunogenicity while retaining function; permanent solution. | Requires extensive epitope mapping; risk of functional impairment. |
| Ex Vivo Delivery & Editing | Cells edited outside the body, washed, and infused. | Avoids systemic exposure to Cas protein; limits immune activation. | Only applicable for certain cell types (e.g., HSPCs, T-cells). |
| Transient Delivery (mRNA/LNP) | Short-lived Cas expression reduces antigen presentation. | Limits duration of immune exposure; low integration risk. | Initial immune trigger still possible; potential reactogenicity. |
| Immunosuppression | Co-administer corticosteroids (e.g., dexamethasone) or mTOR inhibitors. | Broadly suppresses adaptive response; clinically established. | Non-specific; side effects; may interfere with therapeutic cell function. |
| Cas Ortholog Selection | Use Cas proteins from bacteria with low human seroprevalence (e.g., Cas12f, Cas13). | Leverages natural low immunogenicity. | Often smaller, with different PAM requirements; less characterized. |
| Tolerogenic Nanoparticles | Deliver Cas in particles that promote antigen-specific tolerance (e.g., with rapamycin). | Induces antigen-specific immune tolerance. | Early-stage research; dosing and durability uncertain. |
| Serotype Switching (AAV) | Use different AAV capsids for re-dosing to evade NAbs. | Circumvents humoral immunity to vector. | Does not address immunity to Cas payload itself. |
Diagram 2: Decision Workflow for Managing Cas Immune Responses
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for Immune Response Studies
| Reagent/Category | Example Product/Source | Function in Experiment |
|---|---|---|
| Recombinant Cas Proteins | Commercial (e.g., Aldevron, Sino Biological) or in-house purified. | Coating antigen for ELISA; stimulant for T-cell assays. |
| Human Serum/PBMC Panels | Commercial biobanks (e.g., PrecisionMed, AllCells) or clinical cohorts. | Source for assessing pre-existing immunity across diverse HLA backgrounds. |
| HLA-Matched Peptide Pools | Custom synthesis (e.g., JPT Peptide Technologies, GenScript). | Overlapping 15-mers spanning Cas protein for T-cell epitope mapping. |
| ELISA Kits/Components | Anti-human IgG HRP (Jackson ImmunoResearch), TMB Substrate (Thermo Fisher). | Detection and quantification of Cas-specific antibodies. |
| ELISpot Kits | Human IFN-γ ELISpot PLUS (Mabtech) or Human IFN-γ/Granzyme B (Cellular Technology Limited). | Ex vivo quantification of antigen-specific T-cell responses. |
| Flow Cytometry Antibodies | Anti-CD3/CD4/CD8, anti-IFN-γ/TNF-α/IL-2 (BioLegend, BD Biosciences). | Phenotyping and intracellular cytokine staining for ICS. |
| De-immunized Cas Variants | Published variants (e.g., HypaCas9, evoCas9) or proprietary engineered proteins. | Testing the efficacy of protein engineering strategies. |
| Delivery Vehicles (Control) | Research-grade LNPs (e.g., PreciGenome), AAV vectors (e.g., Vigene). | Assessing immunogenicity of different delivery modalities. |
| Immunosuppressants (in vitro) | Cyclosporin A, Tacrolimus, Dexamethasone (Sigma-Millipore). | In vitro validation of pharmacologic mitigation strategies. |
Troubleshooting Low Activity in Type I, III, and IV Systems
Abstract Within the broader thesis of CRISPR-Cas system classification (Types I-VI), achieving robust activity in prokaryotic immune systems is crucial for both understanding microbial immunity and developing novel biotechnological tools. This guide provides a structured, technical approach for diagnosing and resolving low interference or adaptation activity in Type I, Type III, and Type IV CRISPR-Cas systems, which present unique operational complexities compared to the streamlined Type II and Type V systems.
1. Introduction: The Activity Challenge While Types II (Cas9) and V (Cas12) dominate applications, Types I, III, and IV offer unique advantages—such as multi-protein complex targeting (I), transcription-dependent RNA cleavage (III), and plasmid-specific immunity (IV). However, their multi-subunit nature and complex regulatory requirements often lead to experimentally observed low activity. This document frames troubleshooting within the thesis that system classification dictates a defined set of biochemical prerequisites that must be fully recapitulated in heterologous or native experimental setups.
2. Diagnostic Framework & Quantitative Benchmarks Systematic diagnosis begins by quantifying activity against established benchmarks. Key metrics are summarized below.
Table 1: Diagnostic Metrics for Low Activity in Type I, III, and IV Systems
| System Type | Primary Activity Assay | Typical High-Efficiency Benchmark (in vivo) | Common Low-Active Range | Key Quantitative Indicator |
|---|---|---|---|---|
| Type I (e.g., Cascade-Cas3) | Plasmid Clearance / Phage Survival | >99% clearance of target plasmid | <70% clearance | qPCR of plasmid copy number post-transformation. |
| Type III (e.g., Csm/Cmr) | RNA Degradation & ssDNA Cleavage | >90% reduction in target mRNA levels | <50% reduction | RT-qPCR of target transcript; dA activation assay. |
| Type IV (e.g., DinG/Csf) | Conjugative Plasmid Interference | >1000-fold reduction in conjugation efficiency | <10-fold reduction | Conjugation count (CFU/mL) vs. control. |
3. Systematic Troubleshooting by System
3.1 Type I Systems: Cascade Assembly & Cas3 Recruitment Low activity often stems from incomplete ribonucleoprotein complex (Cascade) formation or failed Cas3 helicase-nuclease recruitment.
Experimental Protocol: Cascade Assembly Gel-Shift Assay (EMSA)
- Materials: Purified Cas proteins (Cas5, Cas6, Cas7, Cas8, Cas11), fluorescein-labeled crRNA, non-target DNA (50 bp), target DNA (50 bp with PAM).
- Method: Pre-incubate proteins and crRNA (1:1.2 molar ratio) in assembly buffer (20 mM HEPES, 150 mM KCl, 1 mM DTT, 5% glycerol, pH 7.5) at 25°C for 30 min.
- Add increasing amounts of target/non-target DNA. Incubate 15 min.
- Resolve complexes on a 6% native PAGE gel in 0.5x TBE at 4°C, 100V for 60-90 min.
- Visualize using a fluorescence imager. A complete shift indicates proper assembly and DNA binding.
Key Reagent Solutions:
- Assembly Buffer with DTT & Glycerol: Maintains reducing environment and complex stability.
- Fluorescein-labeled crRNA: Enables sensitive, non-radioactive detection.
- Native PAGE Gel System: Preserves high-molecular-weight complexes.
3.2 Type III Systems: Dual RNA Targeting & Cyclic Oligoadenylate (cOA) Signaling Activity requires transcription of the target and proper cOA second messenger synthesis by the Cas10 subunit.
Experimental Protocol: cOA Extraction & Detection by LC-MS
- Materials: Active Csm/Cmr complex, ATP, ATP-α-S (non-hydrolyzable control), RNase inhibitor, stop solution (8 mM EDTA, 1.2% trifluoroacetic acid).
- Method: Incubate complex with target RNA (500 nM) and 1 mM ATP in reaction buffer at 37°C for 1 hour.
- Halt reaction with stop solution. Centrifuge at 16,000 x g for 10 min.
- Filter supernatant (0.22 µm) and analyze by LC-MS (negative ion mode).
- Detect cA4, cA6, or other cyclic oligomers specific to the subtype. Absence indicates Cas10 catalytic deficiency.
3.3 Type IV Systems: Plasmid Targeting & Complex Localization Function is tightly coupled to the presence of a targeting plasmid and often requires host factors.
Experimental Protocol: Fluorescence Reporter Plasmid Clearance Assay
- Materials: Recipient strain with chromosomal Type IV system, donor strain with mobilizable plasmid (RP4 oriT) expressing GFP and a spacer-matching sequence.
- Method: Perform biparental conjugation on solid media for 2 hours. Resuspend cells and plate dilutions on selective media to count transconjugants (GFP+ colonies).
- In parallel, use flow cytometry to analyze the recipient cell population for GFP fluorescence 4-6 hours post-conjugation initiation.
- Compare transconjugant counts and GFP-negative population percentage to a spacer-less control plasmid.
4. The Scientist's Toolkit: Essential Research Reagents
Table 2: Key Reagent Solutions for Troubleshooting
| Reagent / Material | Primary Function | Application |
|---|---|---|
| Nuclease-Deficient Mutant Complexes | Traps intermediate states for EMSA or structural studies. | All Types (I, III, IV) |
| ATP-γ-S (non-hydrolyzable) | Distinguishes between ATP binding and hydrolysis requirements. | Type I (Cas3), Type III (Cas10) |
| In vitro Transcription Kit (T7) | Generates high-yield, pure target RNA for Type III assays. | Type III |
| Mobilizable Plasmid with oriT | Enables controlled conjugation for Type IV interference assays. | Type IV |
| Anti-His / FLAG / HA Magnetic Beads | For rapid pull-down of tagged complexes to check subunit stoichiometry. | All Types |
| Size-Exclusion Chromatography Standards | Calibrates columns for analyzing native complex assembly. | All Types |
| cOA Standard (e.g., cA4) | LC-MS calibration for absolute quantification of second messenger. | Type III |
5. Visualizing Pathways & Workflows
Title: Diagnostic Decision Tree for Low CRISPR Activity
Title: Type I Cascade-Cas3 Activation Pathway
Title: Type III Dual RNA & cOA Signaling Cascade
6. Conclusion Resolving low activity in multi-subunit CRISPR-Cas systems demands a methodical, biochemistry-centered approach aligned with their classified mechanisms. By quantitatively benchmarking against standard assays, systematically diagnosing assembly and function, and utilizing targeted reagent kits, researchers can robustly activate these complex molecular machines, advancing both fundamental understanding within the Types I-VI classification framework and their potential translational applications.
Expression and Purification Challenges for Large, Multi-Subunit Complexes
The systematic classification and functional characterization of CRISPR-Cas systems (Types I-VI) represent a cornerstone of modern prokaryotic immunity research and biotechnological application. A central, yet formidable, challenge in this field is the robust production of the large, multi-subunit ribonucleoprotein complexes that constitute the functional effectors of these systems. This whitepaper provides an in-depth technical guide to overcoming the expression and purification hurdles for these complex macromolecular assemblies, enabling their structural and biochemical interrogation.
Key Challenges in Multi-Subunit Complex Production
Producing homogeneous, stable samples of complexes such as the Cascade surveillance complex (Type I), Cas9 (Type II), or the multi-protein Csm/Cmr complexes (Types III/VI) presents interconnected obstacles:
- Stoichiometric Imbalance: Differential expression yields of individual subunits in heterologous hosts (typically E. coli) lead to incomplete assemblies and vast excesses of some proteins.
- Insolubility and Aggregation: Individual subunits, especially large proteins like Cas proteins, often misfold and form inclusion bodies when overexpressed alone.
- Complex Instability: Fully assembled complexes can be prone to dissociation during lengthy purification procedures, leading to heterogeneity.
- RNA Component Integration: For many systems (e.g., Types I and III), a mature crRNA must be incorporated, requiring in vivo or in vitro transcription and processing steps.
Strategic Methodologies for Expression and Assembly
Co-expression Strategies
Simultaneous expression of all genes from a single polycistronic vector or multiple compatible vectors promotes in vivo assembly.
Protocol: Multi-Vector Co-expression in E. coli BL21(DE3)
- Vector Construction: Clone genes for each subunit into compatible expression vectors (e.g., pETDuet and pCDFDuet series) with different antibiotic resistance and replication origins. Include N- or C-terminal affinity tags (His6, Strep-II, GST) on one or two subunits.
- Transformation: Co-transform all plasmids into the expression host. Plate on LB agar containing all relevant antibiotics.
- Expression Test: Inoculate a single colony into 5 mL TB media with antibiotics. Grow at 37°C to OD600 ~0.6. Induce with 0.2-0.5 mM IPTG. Shift temperature to 18°C and express for 16-20 hours.
- Lysis and Check: Pellet cells, lyse via sonication in buffer (e.g., 50 mM Tris pH 8.0, 300 mM NaCl, 5% glycerol, 1 mM DTT), and analyze supernatant and pellet by SDS-PAGE to assess solubility and relative expression levels.
In VitroReconstitution
Subunits are expressed and purified individually before being mixed in precise stoichiometric ratios.
Protocol: Stepwise In Vitro Assembly of a Type I Cascade Complex
- Individual Purification: Express and purify subunits (Cas5, Cas6, Cas7, Cas8, Cas11) using standard IMAC and size-exclusion chromatography (SEC).
- crRNA Generation: In vitro transcribe the pre-crRNA from a DNA template using T7 RNA polymerase. Purify via denaturing PAGE or anion-exchange chromatography. Process to mature crRNA using purified Cas6 enzyme.
- Assembly Reaction: Combine subunits and crRNA in a molar ratio (e.g., 1:1:6:1:6:1 for Cas8:Cas11:Cas7:Cas5:Cas6:crRNA) in assembly buffer (20 mM HEPES pH 7.5, 150 mM KCl, 5 mM MgCl2, 1 mM DTT).
- Complex Isolation: Incubate on ice for 30 min, then at 4°C for 1-2 hours. Remove aggregates by centrifugation. Purify the assembled complex using SEC (e.g., Superose 6 Increase 10/300 GL).
Table 1: Comparison of Expression & Purification Strategies for CRISPR Complexes
| CRISPR Type | Example Complex | Subunit Count | Preferred Strategy | Typical Yield (mg/L culture) | Final Purity (%) | Key Stability Factor |
|---|---|---|---|---|---|---|
| Type I | Cascade (E. coli) | 11 proteins + crRNA | Co-expression | 0.5 - 2.0 | >95 | 5 mM MgCl2, 150-300 mM KCl |
| Type II | Cas9-sgRNA | 1 protein + RNA | Individual + in vitro mix | 5 - 20 | >98 | Low salt (≤150 mM NaCl) |
| Type III | Csm/Cmr complex | 4-5 proteins + crRNA | Co-expression | 0.2 - 1.5 | >90 | 200 mM NaCl, 2 mM MgCl2 |
| Type V | Cas12a-crRNA | 1 protein + RNA | Individual + in vitro mix | 3 - 15 | >98 | 5% Glycerol, 1 mM DTT |
| Type VI | Cas13-crRNA | 1 protein + RNA | Individual + in vitro mix | 1 - 8 | >95 | 1-2 mM EDTA, Reducing Agent |
Table 2: Common Chromatography Resins for Complex Purification
| Resin Type | Example Product | Target | Purpose in Purification | Elution Condition |
|---|---|---|---|---|
| IMAC | Ni-NTA Agarose | His6-tag | Initial capture, binds tagged subunit | 250-500 mM Imidazole |
| Anion Exchange | Q Sepharose HP | Negative surface charge | Polishing, removes nucleic acids | NaCl gradient (0-1 M) |
| Cation Exchange | SP Sepharose HP | Positive surface charge | Polishing, separates isoforms | NaCl gradient (0-1 M) |
| Gel Filtration | Superose 6 Increase | Hydrodynamic radius | Final step, isolates monodisperse complex | Isocratic (size-based) |
| Affinity | Strep-Tactin XT | Strep-II-tag | Highly specific capture | 50 mM Biotin |
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function | Example Use Case |
|---|---|---|
| pETDuet-1 Vector | Allows co-expression of two target genes from a single plasmid. | Cloning cas8 and cas5 genes in tandem for Type I complex. |
| BirA Biotin Ligase | Site-specifically biotinylates a 15-aa AviTag for streptavidin capture. | Adding an AviTag to Cas11 for single-particle EM studies. |
| PreScission Protease | Cleaves affinity tags with high specificity at low temperature (4°C). | Removing a GST tag post-purification without destabilizing complex. |
| HiLoad Superdex 200 pg | Preparative-grade SEC resin for high-resolution separation of large complexes (10-600 kDa). | Final polishing step for a 400 kDa Type III Csm complex. |
| RNase Inhibitor (Murine) | Inhibits a broad spectrum of RNases to protect crRNA integrity during purification. | Added to all lysis and purification buffers for RNA-containing complexes. |
| T7 RNA Polymerase (HiScribe) | High-yield in vitro transcription of long, structured crRNA precursors. | Generating pre-crRNA for in vitro assembly of Cascade. |
| HaloTag Ligand Resin | Covalent, high-affinity binding to HaloTag fusion proteins. | Capturing a low-abundance subunit to pull the entire complex from lysate. |
Experimental Workflows and Pathways
Diagram Title: Multi-Subunit Complex Purification Strategy Decision Tree
Diagram Title: Tandem Chromatography Purification Workflow
Assay Validation and Negative Control Strategies for Functional Experiments
Functional experiments are the cornerstone of elucidating the mechanisms of CRISPR-Cas systems, spanning Types I through VI. These experiments, which range from interference assays for Type II (Cas9) to cyclic oligonucleotide signaling in Type III or RNA targeting in Type VI (Cas13), demand rigorous validation. The precise classification and understanding of each system's unique biochemistry and cell biology hinge on the integrity of functional readouts. This guide details the core principles of assay validation and negative control design specifically for CRISPR-Cas functional genomics and biochemistry studies, ensuring that observed phenotypes—be it DNA cleavage, RNA degradation, or collateral activity—are accurately attributed to the intended Cas protein activity.
Foundational Principles of Assay Validation
Validation ensures an experimental method reliably measures the biological activity it intends to. For CRISPR-Cas studies, this is critical to distinguish true on-target effects from off-target artifacts or system-specific bystander effects.
Key Validation Parameters:
- Specificity: The ability to measure only the intended target cleavage or binding event. Crucial for differentiating Type I (multi-Cas complex) from Type V (single effector) activities.
- Sensitivity/Limit of Detection (LoD): The lowest concentration of active Cas:gRNA complex that can be reliably distinguished from background. Vital for quantifying collateral RNase activity in Type VI systems.
- Precision (Repeatability & Reproducibility): The closeness of agreement between repeated measurements under defined conditions.
- Accuracy: The closeness of the measured value to the true value of Cas-mediated editing efficiency.
- Robustness/Ruggedness: The capacity of the assay to remain unaffected by small, deliberate variations in protocol parameters (e.g., Mg²⁺ concentration for Cas12a nuclease activity).
- Dynamic Range & Linearity: The range over which the response (e.g., fluorescence from cleaved reporter) is linearly proportional to the amount of active effector complex.
Table 1: Key Validation Parameters for Common CRISPR-Cas Functional Assays
| Assay Type | Target Cas System | Typical Readout | Key Validation Parameter | Acceptable Benchmark |
|---|---|---|---|---|
| In Vitro Cleavage | Types II, V (DNA) | Gel electrophoresis, FRET | Specificity, LoD | >95% target cleavage; No non-specific banding. |
| Fluorescent Reporter Assay (e.g., HTRF) | Types II, V, VI | Time-resolved fluorescence | Sensitivity, Dynamic Range | LoD < 1 nM; Z'-factor > 0.5. |
| Cell-based Survival (Interference) | Type I, II | Colony formation | Robustness, Precision | CV < 20% across replicates. |
| Next-Gen Sequencing (NGS) | All Types | Indel or depletion frequency | Accuracy, Specificity | High correlation (R² > 0.95) with orthogonal method. |
The Critical Role of Negative Controls
Negative controls are designed to generate a null or baseline signal, isolating the specific contribution of the CRISPR-Cas component under investigation.
Hierarchy of Negative Controls:
- Target-Null Control: Cells or reactions lacking the target DNA/RNA sequence.
- Effector-Null Control: Reactions with catalytically inactive dCas protein (e.g., dCas9, dCas12).
- Guide-Null Control: Reactions with a non-targeting gRNA/scaffold.
- Component-Omission Controls: Systematic omission of individual components (e.g., gRNA, effector protein, co-factors like divalent cations).
- Delivery Control: For cell assays, control for transfection/transduction vehicle alone.
Detailed Experimental Protocols
Protocol 4.1:In Vitro DNA Cleavage Assay Validation for Type V-A (Cas12a)
Purpose: Validate the specific cis and trans cleavage activity of purified Cas12a:crRNA complex. Materials: Purified Cas12a, target and non-target dsDNA, crRNA, NEBuffer r2.1, SYBR Safe dye. Procedure:
- Prepare reaction master mix: 1x NEBuffer r2.1, 50 nM Cas12a, 75 nM crRNA. Incubate 10 min at 25°C for RNP formation.
- Aliquot the RNP mix. Add 10 nM target dsDNA to "Test" reactions. Add 10 nM non-target dsDNA of similar length to "Specificity Control." Add nuclease-free water to "Effector-Null Control."
- Initiate reactions by transferring to 37°C. Remove aliquots at t=0, 5, 15, 30, 60 min.
- Quench reactions with 2x DNA loading dye containing 50 mM EDTA.
- Resolve products on a 2% agarose gel with SYBR Safe, image, and quantify band intensities using ImageJ.
- Validation Analysis: Calculate cleavage percentage. Specificity is confirmed if non-target DNA shows <5% cleavage while target shows >80% within 30 min.
Protocol 4.2:Cell-Based Interference Assay for Type I-E System
Purpose: Quantify plasmid clearance efficiency in E. coli. Materials: E. coli strain expressing Cas proteins, targeting and non-targeting CRISPR array plasmid, transformation-competent cells, selective agar plates. Procedure:
- Transform the CRISPR array plasmid (encoding spacers) into the Cas-expressing strain. Include a non-targeting spacer control.
- Grow transformed colonies overnight in selective media.
- Perform a second transformation with a target plasmid carrying a protospacer with matching PAM and a non-target control plasmid.
- Plate serial dilutions on double-selective plates. Incubate overnight.
- Count colony-forming units (CFU). Interference efficiency = [1 - (CFU target plasmid / CFU non-target plasmid)] x 100%.
- Validation: A valid assay shows >10⁴-fold reduction in CFU for targeting vs. non-targeting controls.
Visualizations
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for CRISPR-Cas Functional Assay Validation
| Reagent Category | Specific Example | Function in Validation/Control |
|---|---|---|
| Catalytically Inactive Effectors | dCas9, dCas12 (D908A for Cas12a), dCas13 (dead HEPN mutants) | Serves as the definitive effector-null control to isolate DNA binding from cleavage or collateral activity. |
| Validated gRNA Controls | Non-targeting scramble gRNA; gRNA with mismatched spacer. | Guide-null controls to identify off-target effects and assay background. |
| Fluorescent Reporter Probes | ssDNA-FQ reporters for Cas12; ssRNA-FQ reporters for Cas13. | Enables real-time, quantitative kinetic measurements for determining LoD and dynamic range. |
| Synthetic Target Oligos | dsDNA with canonical PAM; RNA targets for Cas13. | Provides a defined, pure substrate for sensitivity and specificity testing, free from genomic complexity. |
| Standardized Cell Lines | E. coli BL21(DE3) with Type I-E cascade; HEK293T Cas9-expressing. | Ensures reproducibility and robustness in cell-based interference or editing assays. |
| NGS Validation Kits | Amplicon sequencing libraries with unique molecular identifiers (UMIs). | Gold standard for accuracy assessment, quantifying true editing frequencies and detecting rare off-target events. |
Choosing Your CRISPR Tool: A Comparative Analysis of Types I-VI for Specific Research Goals
CRISPR-Cas systems, classified into six major types (I-VI) and numerous subtypes, constitute the adaptive immune systems of bacteria and archaea. A comprehensive understanding of their molecular mechanisms is central to their classification and biotechnological application. This whitepaper provides a direct technical comparison of three defining characteristics across major Cas effector proteins: Protospacer Adjacent Motif (PAM) requirements, cleavage patterns of target nucleic acids, and effector complex sizes. This analysis is framed within the broader thesis that the evolutionary divergence of CRISPR-Cas types is fundamentally reflected in these quantifiable mechanistic and structural parameters, which directly dictate their suitability for specific research and therapeutic applications.
Core Mechanism Comparison
PAM Requirements: Recognition and Diversity
The PAM is a short sequence adjacent to the target DNA (or, in some cases, RNA) that is essential for self vs. non-self discrimination. Its recognition and sequence specificity vary drastically between CRISPR-Cas types.
Cleavage Patterns: Enzymatic Outcomes
The enzymatic activity and cleavage patterns of Cas effectors define their utility. Systems differ in the types of nucleic acids targeted (DNA, RNA, or both), the number of catalytic sites, and the resulting DNA ends.
Protein and Complex Sizes: Structural Implications
The molecular weight and quaternary structure of the effector complexes influence delivery, especially in therapeutic contexts like gene therapy.
Quantitative Data Comparison Tables
Table 1: PAM Requirements and Cleavage Patterns of Major Cas Effectors
| CRISPR-Cas System | Representative Effector | Canonical PAM Sequence (5'→3')* | Target Nucleic Acid | Cleavage Pattern | Blunt/Sticky End |
|---|---|---|---|---|---|
| Type II | Cas9 (SpCas9) | NGG | dsDNA | Generates DSB via HNH (complementary strand) and RuvC (non-complementary strand) domains. | Blunt (or 5' overhang for some variants) |
| Type V-A | Cas12a (Cpfl) | TTTV | dsDNA | Generates DSB via a single RuvC domain. Creates staggered cuts. | 5' overhang (4-8 nt) |
| Type V-K | Cas12f (Cas14, AsCas12f) | TTTV | dsDNA | Generates DSBs. Ultra-small size. | Blunt or 5' overhang |
| Type VI-A | Cas13a (LshCas13a) | Non-specific; requires protospacer flanking site (PFS) for some variants. | ssRNA | Collateral cleavage of non-target ssRNA upon activation. | N/A (RNA degradation) |
| Type VI-D | Cas13d | Minimal PFS requirement | ssRNA | Collateral cleavage of non-target ssRNA. Compact size. | N/A (RNA degradation) |
| Type I-E | Cascade + Cas3 | AAC (for E. coli) | dsDNA | Cascade complex recognizes target, recruits Cas3 helicase-nuclease for processive degradation. | N/A (processive degradation) |
| Type III-A/B | Cas10 Complex | No DNA PAM; requires transcript from target DNA. | ssRNA (primary), ssDNA (collateral) | Cleaves target RNA and non-specifically cleaves ssDNA upon activation. | N/A |
*N = any nucleotide; V = A, C, or G; PAM is located on the non-target strand.
Table 2: Protein and Complex Sizes
| CRISPR-Cas System | Representative Effector | Effector Protein Size (kDa) | Functional Complex Size & Composition | Key Structural Note |
|---|---|---|---|---|
| Type II | SpCas9 | ~160 kDa | Monomeric (~160 kDa) | Multi-domain single protein. |
| Type V-A | Cas12a | ~130-150 kDa | Monomeric (~130-150 kDa) | Single RuvC domain for cleavage. |
| Type V-K | Cas12f | ~40-70 kDa | Homo-dimeric (~80-140 kDa) | Ultra-small, functions as a dimer. |
| Type VI-A | Cas13a | ~120-130 kDa | Monomeric (~120-130 kDa) | Two HEPN domains for RNA cleavage. |
| Type VI-D | Cas13d | ~90-100 kDa | Monomeric (~90-100 kDa) | Compact RNA-targeting effector. |
| Type I-E | Cascade + Cas3 | Cascade: ~405 kDa; Cas3: ~110 kDa | Cascade (11 subunits: Cas5e, Cas6e, Cas7e, Cas8e, Cse1, Cse2) + Cas3 | Multi-subunit surveillance complex + separate effector. |
| Type III-A/B | Csm/Cmr Complex | Variable | Csm Complex (e.g., 5 subunits: Cas10, Csm2-5, crRNA) ~350 kDa | Multi-subunit complex with integrated cleavage activities. |
Detailed Experimental Protocols
Protocol:In VitroPAM Determination (PAM-SCAN)
This method identifies preferred PAM sequences for a Cas effector.
Materials:
- Purified Cas effector protein.
- Plasmid library containing a randomized PAM region (e.g., NNNN) adjacent to a constant protospacer.
- In vitro cleavage buffer (e.g., 20 mM HEPES pH 7.5, 150 mM KCl, 10 mM MgCl2, 1 mM DTT).
- DNase I or Proteinase K.
- PCR reagents and primers flanking the PAM region.
- High-throughput sequencing platform.
Methodology:
- Cleavage Reaction: Incubate the plasmid library with the Cas effector and crRNA in cleavage buffer at 37°C for 1 hour.
- Reaction Termination: Add Proteinase K or DNase I to digest the protein and stop the reaction.
- Recovery of Uncleaved Plasmids: Transform the reaction mixture into competent E. coli. Only circular, uncut plasmids will confer antibiotic resistance.
- Colony Selection & Sequencing: Isolate plasmid DNA from surviving colonies. Amplify the PAM region by PCR and subject the product to high-throughput sequencing.
- Data Analysis: Align sequenced inserts to the constant protospacer. The sequence immediately adjacent represents the PAM that prevented cleavage. The consensus PAM is derived from the enrichment of sequences in the output library compared to the initial input library.
Protocol:In VitroCleavage Assay for Pattern Analysis
This assay characterizes the cleavage site and overhang structure.
Materials:
- Purified Cas effector protein and cognate crRNA.
- Linear dsDNA substrate (100-500 bp) containing the target protospacer and PAM, 5'-end labeled with [γ-32P] ATP on one or both strands.
- Cleavage buffer (optimized for the effector, e.g., with Mg2+ for Cas9/Cas12).
- Denaturing Urea-PAGE gel (8-10%) or native TBE gel.
- Phosphorimager or autoradiography system.
Methodology:
- Complex Formation: Pre-incubate Cas protein with crRNA (1:2 molar ratio) at 25°C for 10 minutes to form the ribonucleoprotein (RNP) complex.
- Cleavage Reaction: Add the labeled DNA substrate to the RNP in cleavage buffer. Incubate at 37°C. Take aliquots at time points (e.g., 0, 1, 5, 15, 30 min).
- Reaction Stop: Add stop solution (e.g., 95% formamide, 20 mM EDTA, dyes).
- Electrophoresis:
- For precise cut site mapping: Heat samples to 95°C and run on a denaturing urea-PAGE gel alongside a Sanger sequencing ladder of the same substrate.
- For overhang analysis: Run samples on a high-resolution native TBE gel. Cleavage products with overhangs will migrate differently than blunt-end markers of the same length.
- Visualization & Analysis: Expose gel to a phosphor screen. The size of the cleavage fragments indicates the cut site(s) relative to the PAM. Comparison of single-strand vs. double-strand labeled substrates confirms staggered vs. blunt ends.
Visualizations
Diagram 1: CRISPR Type Mechanism Comparison
Diagram 2: PAM-SCAN Assay Workflow
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for CRISPR Mechanism Studies
| Reagent / Material | Supplier Examples (Illustrative) | Function in Key Experiments |
|---|---|---|
| Recombinant Cas Nuclease (His-/MBP-tagged) | Thermo Fisher Scientific, NEB, Sigma-Aldrich, in-house purification. | Purified protein for in vitro cleavage assays, PAM determination, and biochemical characterization. |
| Synthetic crRNA / sgRNA | IDT, Synthego, Horizon Discovery. | Guides the Cas effector to the specific target sequence. Chemical modifications can enhance stability. |
| PAM Library Plasmid Kits | Addgene (deposited plasmids), custom synthesis. | Provides a randomized substrate for high-throughput PAM identification (PAM-SCAN, SELEX). |
| Fluorescently-labeled DNA/RNA Substrates | IDT, Jena Bioscience. | For real-time kinetic measurement of cleavage activity (FRET-based assays) and gel-based analysis. |
| [γ-32P] ATP or [α-32P] dNTPs | PerkinElmer, Hartmann Analytic. | Radioactive labeling of oligonucleotide substrates for highly sensitive detection of cleavage products on gels. |
| High-Fidelity PCR Master Mix | NEB, QIAGEN, Thermo Fisher. | Amplification of PAM regions from plasmid libraries prior to sequencing. |
| High-Resolution Gel Electrophoresis System | Bio-Rad, Thermo Fisher. | Native or denaturing PAGE for precise separation and sizing of cleavage products. |
| Next-Generation Sequencing Service/Platform | Illumina, PacBio. | Deep sequencing of PAM library inserts for comprehensive sequence preference analysis. |
| Microfluidic Cartridges for Binding Kinetics | ForteBio (Blitz), Nicoya (LSPR). | Measures real-time binding affinity (KD) between Cas RNP and target sequences with different PAMs. |
Specificity and Fidelity Benchmarks Across Cas9, Cas12, and Cas13
The classification of CRISPR-Cas systems into six distinct types (I-VI) is based on signature effector proteins and the architecture of their interference complexes. Within this framework, Types II (Cas9), V (Cas12), and VI (Cas13) are the most widely adopted for genome and transcriptome engineering. Their utility in therapeutic development is fundamentally constrained by their "specificity" (the ability to discriminate between the intended target and off-target sequences) and "fidelity" (the precision of cleavage at the intended site without promiscuous activity). This guide provides a technical benchmark of these critical parameters across the three effector families, contextualized within the broader mechanistic diversity of Types I-VI.
Cas9 (Type II): A dual-RNA-guided (crRNA:tracrRNA) DNase that generates blunt double-strand breaks (DSBs). Off-targets arise primarily from toleration of mismatches and bulges in the DNA-RNA heteroduplex, particularly in the PAM-distal region, and are influenced by guide RNA structure, enzyme variants, and cellular context.
Cas12 (Type V): A single crRNA-guided DNase (e.g., Cas12a/Cpf1) that produces staggered DSBs. While also DNA-targeting, Cas12 systems exhibit a distinct PAM requirement (often T-rich) and demonstrate "collateral" single-stranded DNA (ssDNA) trans-cleavage activity post-target activation, a source of off-target nucleic acid degradation.
Cas13 (Type VI): A single crRNA-guided RNase that cleaves single-stranded RNA (ssRNA). Its primary off-target concern is collateral trans-cleavage of non-target ssRNA upon activation, a feature not found in DNA-targeting effectors and of significant consequence for cellular health.
Quantitative Benchmark Tables
Table 1: In Vitro Biochemical Specificity & Fidelity Metrics
| Effector (Example) | PAM/PFS Sequence | Cleavage Type | Reported in vitro Off-Target Rate (High-Fidelity Variants) | Key Determinant of Fidelity |
|---|---|---|---|---|
| SpCas9 (Type II) | 5'-NGG-3' | Blunt DSB | <0.1% (e.g., HypaCas9, eSpCas9) | DNA:RNA heteroduplex stability; RuvC/HNH domain conformation |
| AsCas12a (Type V) | 5'-TTTV-3' | Staggered DSB | ~0.1-1% (enAsCas12a) | PAM interrogation; guide DNA unwinding kinetics |
| LwaCas13a (Type VI) | Non-G PFS | ssRNA cleavage | High collateral RNase activity (uncatalyzed) | Protospacer Flanking Site (PFS); target activation threshold |
| Assay Name | Primary Application (Effector) | Detection Principle | Key Advantage | Key Limitation |
|---|---|---|---|---|
| CIRCLE-seq | Cas9, Cas12 | In vitro circularization & NGS | Sensitive, genome-agnostic | Does not reflect cellular context |
| GUIDE-seq | Cas9, Cas12 | Integration of dsODN tags at DSBs | In-cell, unbiased | Lower sensitivity; tag toxicity |
| DIG-seq | Cas12 | Detection of ssDNA ends | Specific to Cas12 collateral activity | Specialized for Cas12 systems |
| CLEAR-seq | Cas13 | Proximity ligation of cleaved RNA | Maps collateral RNA cleavage | Requires crosslinking and specific library prep |
Detailed Experimental Protocols
Protocol: GUIDE-seq for In-Cell Off-Target Profiling of Cas9 and Cas12
Objective: To identify genome-wide off-target sites of CRISPR-Cas nucleases in living cells.
Materials: See "Research Reagent Solutions" table.
Procedure:
- Cell Preparation & Transfection: Seed HEK293T cells (or relevant cell line) to reach 70-80% confluency at transfection. For a 24-well plate, prepare a transfection mix containing: 500 ng of Cas9/Cas12 expression plasmid, 100 ng of sgRNA/crRNA expression plasmid, and 100 pmol of phosphorylated, HPLC-purified GUIDE-seq dsODN tag. Use a suitable transfection reagent (e.g., Lipofectamine 3000) according to manufacturer protocol.
- Genomic DNA Extraction: 72 hours post-transfection, harvest cells and extract genomic DNA using a DNeasy Blood & Tissue Kit. Quantify DNA via fluorometry.
- Library Preparation: Digest 2 µg of gDNA with MseI (4-base cutter). Ligate annealed dsODN adapters compatible with MseI overhangs. Purify ligated DNA and perform PCR enrichment (14-18 cycles) using primers containing Illumina adapter sequences and barcodes. Use a proof-reading polymerase.
- Sequencing & Analysis: Pool libraries and sequence on an Illumina MiSeq or HiSeq platform (2x150 bp). Process reads using the published GUIDE-seq analysis pipeline (available on GitHub) to map tag integration sites, requiring a minimum of two unique reads per site.
Protocol:In VitroCollateral Cleavage Assay for Cas13
Objective: To quantify the collateral RNase activity of activated Cas13.
Procedure:
- Reaction Setup: In a black-walled 384-well plate, assemble a 20 µL reaction containing: 1x Reaction Buffer (20 mM HEPES pH 6.8, 50 mM KCl, 5 mM MgCl2), 50 nM purified LwaCas13a protein, 50 nM crRNA (targeting a specific ssRNA activator), 100 nM quenched fluorescent ssRNA reporter probe (e.g., 5'-/6-FAM/rUrUrUrUrUrU/3IABkFQ/-3').
- Activation: Initiate the reaction by adding the target ssRNA activator at varying concentrations (e.g., 0, 0.1, 1, 10 nM). Include a no-activator negative control and a reporter-only background control.
- Detection: Immediately transfer the plate to a fluorescent plate reader pre-warmed to 37°C. Measure fluorescence (Ex/Em: 485/535 nm) every 30 seconds for 60-90 minutes.
- Analysis: Calculate the maximum rate of fluorescence increase (RFU/sec) or endpoint fluorescence. Plot signal versus activator concentration to determine the activation kinetics and the threshold for collateral activity induction.
The Scientist's Toolkit: Research Reagent Solutions
| Reagent / Material | Function / Purpose | Example Product / Note |
|---|---|---|
| High-Fidelity Cas Variant Expression Plasmids | Delivery of engineered, high-specificity nucleases (e.g., SpCas9-HF1, HypaCas9, enAsCas12a). | Addgene vectors: #72247 (SpCas9-HF1), #120741 (enAsCas12a-HF). |
| Synthetic, Chemically Modified sgRNA/crRNA | Enhanced stability and potentially reduced off-target effects via 2'-O-methyl, phosphorothioate modifications. | Synthesized via IDT, Horizon Discovery. |
| GUIDE-seq dsODN Tag | Double-stranded oligodeoxynucleotide that integrates into DSBs for off-target site capture. | PAGE-purified, 5'-phosphorylated; Sequence as published (Tsai et al., 2015). |
| Quenched Fluorescent RNA Reporters | Detection of Cas13 collateral RNase activity; fluorescence increases upon cleavage. | Molecular Beacons or linear probes (e.g., from IDT, Biosearch Tech). |
| CIRCLE-seq Adapter Oligos | For circularization of in vitro cleaved genomic DNA prior to NGS library prep. | Required for sensitive in vitro off-target profiling. |
| DIG-seq Adapter | Specifically captures Cas12-generated ssDNA ends for sequencing. | Key reagent for the DIG-seq protocol (Li et al., 2020). |
| Next-Generation Sequencing Kit | For high-throughput sequencing of off-target amplicon libraries. | Illumina MiSeq Reagent Kit v3 (600-cycle). |
| Off-Target Analysis Software | Bioinformatics pipeline for mapping and quantifying off-target events from NGS data. | GUIDE-seq (GitHub), CIRCLE-seq (GitHub), Cas-OFFinder (web tool). |
Concluding Perspective: Towards Therapeutic-Grade Fidelity
Benchmarking specificity across Cas9, Cas12, and Cas13 reveals a fundamental trade-off between activity and fidelity, and highlights unique risks like collateral cleavage. The evolution of high-fidelity variants, coupled with increasingly sensitive in silico and experimental off-target detection methods, is critical for de-risking therapeutic applications. Future development within the CRISPR-Cas landscape (Types I-VI) may uncover new effectors with inherent high precision, further expanding the toolbox for accurate genomic and transcriptomic medicine. Continuous benchmarking using standardized protocols, as outlined herein, remains essential for the field.
Throughput and Multiplexing Capabilities of Different Systems
This technical guide provides an in-depth analysis of the throughput and multiplexing capabilities of CRISPR-Cas systems, contextualized within the broader research on the classification of Types I-VI. As CRISPR technologies evolve from basic gene editing to complex transcriptional regulation, base editing, and diagnostics, the demands on system throughput (the scale of perturbations possible in a single experiment) and multiplexing (the ability to target multiple genomic loci simultaneously) have become paramount. This whitepaper synthesizes current methodologies, quantitative capabilities, and experimental protocols, serving as a resource for researchers and drug development professionals.
CRISPR-Cas System Classification and Core Properties
CRISPR-Cas systems are broadly divided into two classes based on effector module complexity. Class 1 (Types I, III, and IV) utilizes multi-subunit effector complexes, while Class 2 (Types II, V, and VI) employs single effector proteins. This fundamental architectural difference has direct implications for multiplexing potential and experimental throughput.
Quantitative Comparison of Throughput and Multiplexing
The following table summarizes key performance metrics for representative systems from each type, based on current literature. Throughput is defined as the practical scale of targets addressable in a pooled screen. Multiplexing refers to the confirmed number of simultaneous, independent edits or manipulations.
Table 1: Throughput and Multiplexing Capabilities of Major CRISPR-Cas Systems
| System Type | Example Effector | Primary Function | Typical Multiplexing Capacity (Loci) | Screening Throughput (Library Size) | Key Enabling Technology |
|---|---|---|---|---|---|
| Type I | Cascade-Cas3 | DNA cleavage (long-range) | Moderate (5-10) | Up to 10⁵ | crRNA arrays, Cas3 processivity |
| Type II | Cas9 | DNA cleavage (blunt end) | High (10-100+) | >10⁶ (Common) | sgRNA, arrayed sgRNA libraries |
| Type III | Cas10 complex | RNA cleavage, DNA cleavage (collateral) | High (via transcription) | Up to 10⁵ | RNA targeting, transcription-dependent |
| Type IV | Csf1 complex | Putative DNA targeting | Low/Undefined | Limited Data | -- |
| Type V | Cas12a (Cpfl) | DNA cleavage (staggered) | High (10-100+) | >10⁶ | crRNA arrays, minimal constraints |
| Type VI | Cas13a | RNA cleavage (collateral) | High (via collateral effect) | Up to 10⁵ | SHERLOCK, REPORTER assays |
Detailed Experimental Protocols
Protocol 1: High-Throughput Knockout Screening with Type II (Cas9)
Objective: Perform a genome-scale loss-of-function screen to identify genes essential for cell viability. Workflow:
- Library Design: Design a pooled sgRNA library targeting ~20,000 human genes with 4-10 sgRNAs per gene (total library size ~100,000 constructs). Include non-targeting control sgRNAs.
- Library Delivery: Clone the sgRNA library into a lentiviral vector containing the Cas9 gene (all-in-one) or use a stable Cas9-expressing cell line.
- Viral Production & Transduction: Produce lentivirus at low MOI (<0.3) to ensure most cells receive a single sgRNA. Transduce the target cell population to achieve ~500x coverage of the library.
- Selection & Passaging: Apply puromycin selection (if vector contains a resistance marker) for 3-5 days. Passage cells for 14-21 population doublings, maintaining coverage.
- Genomic DNA Extraction & Sequencing: Harvest genomic DNA from the initial (T0) and final (Tend) cell populations. Amplify the integrated sgRNA sequences via PCR using primers containing Illumina adapters.
- Data Analysis: Perform next-generation sequencing. Quantify sgRNA abundance in T0 vs. Tend samples. Depleted sgRNAs (and their target genes) indicate essentiality. Use statistical algorithms (e.g., MAGeCK, DESeq2) for hit calling.
Protocol 2: Multiplexed Transcriptional Activation with Type V (dCas12a-VPR)
Objective: Simultaneously activate multiple endogenous genes to reprogram cell state. Workflow:
- crRNA Array Cloning: Synthesize a single DNA fragment containing a direct repeat (DR) followed by the ~20-24 nt spacer for each target gene, concatenated sequentially. Clone this array into a plasmid expressing Francisella novicida dCas12a fused to the VPR transcriptional activator.
- Expression System Delivery: Deliver the crRNA array-dCas12a-VPR plasmid into target cells via nucleofection or lipid-based transfection.
- Array Processing Validation: Confirm in vivo processing of the crRNA array by the Cas12a protein. Extract total RNA 48-72 hours post-transfection.
- Downstream Analysis: Perform RT-qPCR for each target gene to quantify activation fold-change relative to non-targeting controls. For phenotypic assessment, conduct flow cytometry or functional assays at the appropriate time point.
Protocol 3: Multiplexed Nucleic Acid Detection with Type VI (Cas13)
Objective: Detect the presence of multiple specific RNA pathogens in a single reaction (SHERLOCK-based). Workflow:
- Sample Preparation: Extract RNA from clinical or environmental samples.
- Amplification & Detection Reaction: Set up a recombinase polymerase amplification (RPA) or RT-RPA step to amplify target sequences. In the same tube, include:
- Purified LwaCas13a or PspCas13b protein.
- Specific crRNAs for each target pathogen.
- A quenched fluorescent RNA reporter molecule (e.g., FAM-UU-BHQ1).
- Incubation & Reading: Incubate the reaction at 37°C for 30-60 minutes in a plate reader or real-time PCR machine capable of measuring fluorescence.
- Interpretation: A positive fluorescence signal above baseline indicates collateral cleavage of the reporter due to Cas13 activation by any of the targeted RNAs present. Using different fluorescent reporters (e.g., FAM, HEX) for different crRNAs in separate reactions allows multiplexed detection in a single instrument run.
Visualizing System Workflows and Relationships
Diagram 1: CRISPR System Selection and Experimental Workflow
Diagram 2: Guide Architecture and Multiplexing Pathways
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for CRISPR Throughput & Multiplexing Experiments
| Reagent / Material | Function / Description | Example Application |
|---|---|---|
| Lentiviral Packaging Plasmids (psPAX2, pMD2.G) | Third-generation system for producing replication-incompetent lentivirus to deliver CRISPR constructs with high efficiency into diverse cell types. | Creating stable Cas9-expressing lines; delivering pooled sgRNA libraries. |
| Pooled sgRNA/crRNA Library | A synthesized DNA pool containing thousands to hundreds of thousands of unique guide sequences, cloned into a delivery backbone. | Genome-wide knockout, activation, or inhibition screens. |
| Next-Generation Sequencing Kits (Illumina) | For preparing and sequencing amplicons of integrated guide sequences to quantify abundance changes in pooled screens. | Screen deconvolution and hit identification (e.g., Nextera XT). |
| Arrayed sgRNA/crRNA Collections | Individual guide RNAs or expression plasmids in multi-well plate format, allowing for parallel testing of specific targets. | Validation of screen hits; focused combinatorial multiplexing. |
| Cas9, Cas12a, Cas13 Protein (Purified) | Recombinantly expressed and purified effector proteins for in vitro applications. | DNA/RNA cutting assays; diagnostic tests (SHERLOCK, DETECTR). |
| CRISPR-Compatible Cell Lines (HEK293T, K562, etc.) | Cell lines with high transfection efficiency, stable Cas9/Cas12a expression, or robust phenotypic readouts. | Standardized screening and multiplex editing experiments. |
| Fluorescent Reporters (FAM-BHQ1, HEX-BHQ2 RNA) | Quenched oligonucleotide probes cleaved by activated Cas13 or Cas12a, producing a fluorescent signal. | Multiplexed, real-time detection of nucleic acids in diagnostic assays. |
| HDR Donor Template Pools | Libraries of single-stranded or double-stranded DNA oligonucleotides containing variant sequences for homology-directed repair. | Saturation mutagenesis screens; multiplex base editing studies. |
Within the broader thesis on CRISPR-Cas system classification (Types I-VI), a critical translational challenge for Type II (Cas9) and Type V (Cas12) systems is their potential for off-target activity. While novel Types (e.g., VI for RNA targeting) present different specificity profiles, the therapeutic application of DNA-targeting nucleases necessitates rigorous validation of target specificity. This guide details three cornerstone, NGS-based techniques for unbiased off-target profiling, which are essential for advancing any CRISPR-based therapeutic from research to clinic.
Core Techniques: Methodologies & Comparisons
NGS-Based Off-Target Analysis (Prediction-Dependent)
This method involves sequencing putative off-target sites identified by in silico prediction algorithms (e.g., based on sequence similarity and chromatin accessibility).
- Protocol: Genomic DNA is harvested, fragmented, and hybridized with biotinylated probes complementary to predicted off-target sites. Captured DNA is amplified and sequenced via NGS. Reads are aligned to the reference genome to identify indels.
- Limitation: Relies entirely on prediction algorithms, missing unpredicted sites.
GUIDE-seq (Genome-wide, Unbiased Identification of DSBs Enabled by Sequencing)
A genome-wide, in situ method that captures double-strand breaks (DSBs) in living cells by integrating a short, double-stranded oligodeoxynucleotide (dsODN) tag.
- Detailed Protocol:
- Transfection: Co-deliver Cas9-gRNA RNP and the dsODN tag into target cells.
- Tag Integration: Cellular repair machinery integrates the dsODN into DSBs generated by Cas9 (both on- and off-target).
- Genomic DNA Extraction & Shearing: Harvest genomic DNA and fragment it by sonication.
- Library Preparation & Enrichment: Perform adaptor ligation and PCR using one primer specific to the integrated dsODN tag and another to the adaptor, selectively amplifying tagged genomic fragments.
- NGS & Analysis: Sequence amplicons. Break sites are identified as genomic junctions adjacent to the dsODN sequence.
CIRCLE-seq (Circularization forIn VitroReporting of Cleavage Effects by Sequencing)
An in vitro, ultra-sensitive method that uses circularized genomic DNA as a substrate for Cas nuclease digestion.
- Detailed Protocol:
- Genomic DNA Circularization: Extract genomic DNA, fragment it, and enzymatically circularize the fragments to create a library of covalently closed circles.
- In Vitro Cleavage: Incubate the circularized DNA library with the Cas9-gRNA RNP. Cas9 cleaves circles, linearizing them at sites complementary to the gRNA.
- Selective Amplification of Cleaved Fragments: Treat with exonuclease to degrade all remaining linear DNA (uncleaved circles are resistant). The linearized, cleaved fragments are then purified.
- Library Prep & NGS: Add sequencing adaptors via PCR or ligation and perform deep sequencing. Breaks are mapped to the reference genome.
Table 1: Comparative Analysis of Key Off-Target Detection Techniques
| Feature | NGS-Based (Prediction-Dependent) | GUIDE-seq | CIRCLE-seq |
|---|---|---|---|
| Detection Basis | Hybrid capture & sequencing | In situ dsODN tag integration | In vitro cleavage of circularized DNA |
| Scope | Targeted (predicted sites only) | Genome-wide, unbiased in cells | Genome-wide, unbiased in vitro |
| Sensitivity | Moderate (Limited by probe design) | High (~0.1% modification frequency) | Extremely High (Can detect <0.01% of sites) |
| Cellular Context | In vivo (Cells or tissue) | In vivo (Living cells) | In vitro (Purified genomic DNA) |
| Primary Output | Indel frequency at predicted sites | Genome-wide map of DSB locations | Genome-wide map of cleavage susceptibility |
| Key Advantage | Straightforward for validating top candidates. | Captures cell-specific chromatin effects. | Highest sensitivity; no cellular toxicity concerns. |
| Key Limitation | Misses novel/ unpredicted off-targets. | dsODN can be cytotoxic; requires efficient delivery. | Lacks cellular context (repair, chromatin). |
Table 2: Typical Experimental Metrics from Recent Studies (Representative Data)
| Technique | Typical Sequencing Depth | Detection Threshold | Time to Result | Relative Cost |
|---|---|---|---|---|
| NGS-Based (Targeted) | 10,000x - 100,000x per site | ~0.1 - 0.5% indel frequency | 1-2 weeks | $$ |
| GUIDE-seq | 50 - 100 million total reads | ~0.1% integration frequency | 2-3 weeks | $$$ |
| CIRCLE-seq | 50 - 200 million total reads | <0.01% cleavage frequency | 1-2 weeks | $$ |
Experimental Workflow Diagrams
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for Off-Target Analysis Experiments
| Reagent / Material | Function / Purpose | Example Application |
|---|---|---|
| Recombinant Cas9 Nuclease | High-purity, active nuclease for RNP formation. | GUIDE-seq transfection; CIRCLE-seq in vitro cleavage. |
| Synthetic gRNA (chemically modified) | Provides targeting specificity; chemical modifications can enhance stability. | All techniques. Critical for defining the target. |
| dsODN Tag (e.g., GUIDE-seq Adapter) | Short, blunt, double-stranded DNA for integration into DSBs. | Essential for GUIDE-seq to label break sites in vivo. |
| Circularization Ligase | Enzymatically ligates DNA ends to form covalent circles. | Critical step in CIRCLE-seq library preparation. |
| Exonuclease (e.g., T5 or T7 Exo) | Degrades linear dsDNA, enriching for cleaved (linearized) circles. | Post-cleavage cleanup in CIRCLE-seq. |
| PCR Additives (e.g., Betaine) | Reduces amplification bias in GC-rich regions during enrichment PCR. | GUIDE-seq tag-specific amplification; CIRCLE-seq library prep. |
| High-Sensitivity DNA Assay Kit | Accurate quantification of low-concentration DNA libraries prior to NGS. | Quality control for all NGS library preparation steps. |
| Hybridization Capture Beads | Streptavidin-coated magnetic beads for biotinylated probe capture. | Enrichment of predicted off-target regions in targeted NGS. |
Within the expanding classification of CRISPR-Cas systems (Types I-VI), the primary objective remains precise genomic manipulation. Type II (Cas9) and Type V (Cas12) systems are the predominant workhorses for knockout (KO), knockdown (KD), and precise editing. However, the functional outcome of any editing event is paramount, regardless of the specific Cas protein or system type employed. Effective validation requires a multi-modal approach that confirms genetic change, assesses functional protein loss, and measures consequent phenotypic alterations. This guide details the core methodologies for robust validation.
Validation of Genetic Alterations
The first line of validation is confirming the intended DNA sequence change.
For Knockouts and Editing Events
- Sanger Sequencing & TIDE/ICE Analysis: Standard Sanger sequencing of PCR-amplified target regions is the baseline. For heterogeneous pools of edits (common in transfected cell populations), tools like Tracking of Indels by Decomposition (TIDE) or Inference of CRISPR Edits (ICE) quantitatively deconvolute sequencing chromatograms to provide indel percentages and spectra.
- Protocol: 1) Lysate edited cells. 2) PCR amplify target locus (~500-800 bp). 3) Purify PCR product. 4) Submit for Sanger sequencing. 5) Upload sequencing trace and reference sequence to TIDE/ICE web tool.
- Next-Generation Sequencing (NGS): The gold standard for quantitative, unbiased assessment of editing efficiency and specificity. Amplicon sequencing of target loci provides precise indel rates, haplotype analysis, and can detect off-target effects when combined with whole-genome or targeted-off-target sequencing.
- Protocol: 1) Design primers with overhangs for Illumina adapters. 2) Perform a two-step PCR: first to amplify the target, second to add full adapter indices. 3) Purify, quantify, pool libraries, and sequence on a MiSeq or similar platform. 4) Analyze with pipelines like CRISPResso2.
For Knockdowns (e.g., dCas9-based repression)
- Quantitative PCR (qPCR): Measures the reduction in target mRNA levels.
- Protocol: 1) Isolate total RNA from edited and control cells. 2) Synthesize cDNA. 3) Perform qPCR using primers specific to the target gene and housekeeping controls (e.g., GAPDH, ACTB). 4) Calculate fold-change using the ΔΔCt method.
Table 1: Quantitative Comparison of Genotypic Validation Methods
| Method | Primary Use | Key Metric | Throughput | Approximate Cost/Sample | Key Limitation |
|---|---|---|---|---|---|
| Sanger + TIDE/ICE | Indel quantification in pools | % Indel Efficiency | Low-Medium | $10-$30 | Low sensitivity for variants <5% frequency |
| Amplicon NGS | Precise editing & off-target analysis | Indel %, Haplotype distribution | High | $50-$200 | Requires bioinformatics expertise |
| qPCR | mRNA knockdown validation | Fold-change (ΔΔCt) | High | $5-$15 | Measures mRNA, not functional protein |
Validation of Functional Protein Loss
Genetic confirmation must be coupled with assessment of protein-level impact.
Western Blotting
The direct method to confirm protein knockout or severe knockdown.
- Protocol: 1) Lyse cells in RIPA buffer. 2) Separate proteins by SDS-PAGE. 3) Transfer to PVDF membrane. 4) Probe with target-specific and loading control antibodies. 5) Quantify band intensity via densitometry.
Flow Cytometry / Immunofluorescence (IF)
Essential for assessing efficiency in mixed cell populations and for cell-surface or intracellular proteins.
- Protocol (Flow): 1) Harvest cells (for intracellular targets, fix and permeabilize). 2) Stain with antibody conjugated to a fluorophore. 3) Analyze on a flow cytometer. The shift in fluorescence intensity directly quantifies the percentage of protein-negative cells.
- Protocol (IF): 1) Fix and permeabilize cells on a coverslip. 2) Stain with primary then fluorescent secondary antibodies. 3) Image with a microscope. Provides spatial protein localization data.
Table 2: Quantitative Protein-Level Validation Metrics
| Method | Key Metric | Typical Validation Threshold | Time to Result | Advantage |
|---|---|---|---|---|
| Western Blot | Band intensity vs. control | >70-80% reduction | 1-2 days | Direct, uses standard lab equipment |
| Flow Cytometry | % of protein-negative cells | >70% negative population | <1 day | Quantitative, single-cell resolution, high-throughput capable |
| Immunofluorescence | Visual protein absence & localization | Complete loss of signal | 1-2 days | Adds morphological context |
Validation of Phenotypic Consequences
Ultimate validation requires linking genetic and protein-level changes to a relevant biological output.
Cell Viability/Proliferation Assays
For genes essential in a given context (e.g., oncogenes in cancer cell lines).
- Protocol (MTT/CCK-8): Seed edited and control cells in a 96-well plate. At multiple time points, add MTT or CCK-8 reagent. Measure absorbance. Plot growth curves.
Reporter Assays
Engineered fluorescent or luminescent readouts for pathway activity (e.g., Wnt, NF-κB) disrupted by target gene editing.
- Protocol: Co-transfect cells with the CRISPR component and a reporter plasmid containing the pathway-responsive element driving luciferase. After editing, lyse cells and measure luminescence.
Functional Rescue (Critical Control)
Re-introducing a CRISPR-resistant, wild-type version of the target gene should restore the wild-type phenotype, confirming specificity.
- Protocol: After creating a KO clone, transfect with a plasmid expressing the target cDNA with silent mutations in the gRNA target site. Perform the relevant phenotypic assay (e.g., proliferation, reporter) to confirm rescue.
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in Validation |
|---|---|
| High-Fidelity DNA Polymerase (e.g., Q5, KAPA HiFi) | Accurate amplification of target loci for sequencing and NGS library prep. |
| CRISPResso2 Software | Standardized, open-source NGS analysis pipeline for quantifying genome editing outcomes. |
| Validated Antibodies (for WB/Flow/IF) | Specific detection of target protein loss; validation for the application is critical. |
| Fluorescent-Conjugated Secondary Antibodies | Enable detection in flow cytometry and immunofluorescence. |
| Cell Viability Dye (e.g., MTT, CCK-8) | Quantitative measurement of proliferation changes post-editing. |
| Dual-Luciferase Reporter Assay System | Quantifies changes in transcriptional activity of pathways affected by gene editing. |
| CRISPR-Resistant cDNA Clone | Essential reagent for performing a functional rescue experiment to confirm specificity. |
Experimental Workflows & Pathway Logic
Validation Workflow for CRISPR Edits
Linking CRISPR Action to Functional Readouts
Comparative Suitability for Diagnostic Sensitivity, Speed, and Cost
1. Introduction Within the broader research thesis on CRISPR-Cas system classification (Types I-VI), a critical applied goal is the development of next-generation molecular diagnostics. The suitability of each system for diagnostic applications is governed by a triad of parameters: analytical sensitivity (limit of detection), assay speed (time-to-result), and cost (per-test, inclusive of reagents and instrumentation). This technical guide provides a comparative analysis of CRISPR-Cas Types II (Cas9, Cas12, Cas13), V (Cas12f), and VI (Cas13) for diagnostic use, focusing on their inherent biochemical mechanisms that directly influence these key metrics.
2. Core Mechanisms & Diagnostic Performance
- Cas9 (Type II-A): Functions primarily as a double-stranded DNA (dsDNA) cleaver guided by a crRNA. Its diagnostic application (e.g., SHERLOCKv1) relies on pre-amplification of target nucleic acid, as its collateral activity is minimal. It offers high specificity but generally lower inherent sensitivity without amplification.
- Cas12 (Type V-A): Upon binding and cleaving its target dsDNA, exhibits promiscuous, non-specific single-stranded DNA (ssDNA) collateral cleavage. This trans-activity enables signal amplification from a single target recognition event, directly boosting sensitivity.
- Cas12f (Type V-F): A hypercompact variant (e.g., Cas12f1, ~400-700 amino acids). While also a dsDNA-targeting, ssDNA collateral-cleaving enzyme, its smaller size can reduce reagent costs and improve cellular delivery, but often with initially lower catalytic efficiency than Cas12a, requiring protein engineering for robust diagnostic performance.
- Cas13 (Type VI): Upon binding and cleaving its target single-stranded RNA (ssRNA), exhibits promiscuous cleavage of surrounding non-target ssRNA (collateral activity). This makes it uniquely suited for RNA pathogen detection and direct viral surveillance.
3. Quantitative Comparison Table
Table 1: Comparative Diagnostic Performance of Key CRISPR-Cas Systems
| CRISPR-Cas Type | Example Enzyme | Target | Collateral Activity Substrate | Reported Sensitivity (LoD) | Typical Assay Time (Post-Amplification) | Key Cost Drivers |
|---|---|---|---|---|---|---|
| II-A | SpCas9 | dsDNA | None (Requires reporter cleavage via separate enzyme) | ~aM - fM* | 60 - 90 min | High-cost protein production; qPCR instrument |
| V-A | LbCas12a | dsDNA | ssDNA | ~aM - single copy | 10 - 30 min | Recombinant protein; fluorescent reader |
| V-F | AsCas12f1 | dsDNA | ssDNA | ~fM - pM (improved variants reach aM) | 30 - 60 min | Lower protein production cost; engineered variants |
| VI-B | LwCas13a | ssRNA | ssRNA | ~aM - single copy | 5 - 20 min | Recombinant protein; RNA handling protocols |
aM = attomolar (10^-18 M); fM = femtomolar (10^-15 M). *Note: Sensitivity for Cas9-based diagnostics is contingent on the pre-amplification method used (e.g., RPA, PCR).
Table 2: Suitability Scoring (1-Low, 5-High) for Diagnostic Parameters
| Parameter | Cas9 (II) | Cas12a (V) | Cas12f (V) | Cas13 (VI) |
|---|---|---|---|---|
| Inherent Sensitivity | 2 | 5 | 3 (4 for engineered) | 5 |
| Assay Speed | 2 | 4 | 3 | 5 |
| Reagent Cost Potential | 2 | 3 | 5 | 3 |
| Instrument Simplicity | 2 | 4 | 4 | 5 |
| Multiplexing Potential | 3 | 3 | 2 | 5 |
4. Detailed Experimental Protocols
4.1 Protocol: DETECTR Assay using Cas12a for DNA Virus Detection
- Principle: Recombinase Polymerase Amplification (RPA) followed by Cas12a target recognition and collateral cleavage of a quenched ssDNA reporter.
- Reagents: Target DNA sample, RPA primers, TwistAmp Basic RPA kit, purified LbCas12a protein, crRNA, ssDNA-FQ reporter (e.g., 6-FAM/TTATT/3BHQ_1), Nuclease-Free Water, Reaction Buffer (20 mM HEPES, 100 mM NaCl, 5 mM MgCl2, pH 6.8).
- Procedure:
- Isothermal Amplification: Prepare a 50 µL RPA reaction per manufacturer's protocol. Incubate at 37-42°C for 15-20 minutes.
- CRISPR Detection Mix: In a separate tube, combine 100 nM LbCas12a, 120 nM crRNA, 500 nM ssDNA-FQ reporter in 1x Reaction Buffer. Incubate at 25°C for 10 minutes for RNP complex formation.
- Detection Reaction: Transfer 2 µL of the completed RPA reaction into 23 µL of the CRISPR Detection Mix. Mix gently.
- Signal Measurement: Incubate the final 25 µL reaction at 37°C. Monitor fluorescence (Ex/Em: 485 nm/520 nm) in real-time using a plate reader or endpoint measurement after 10-30 minutes.
- Data Analysis: A positive sample shows a time-dependent increase in fluorescence signal exceeding a threshold defined by negative controls (no template).
4.2 Protocol: SHERLOCK Assay using Cas13 for RNA Virus Detection
- Principle: Reverse Transcription-RPA (RT-RPA) followed by Cas13 target recognition and collateral cleavage of a quenched ssRNA reporter.
- Reagents: Target RNA sample, RT-RPA primers, Purified LwCas13a protein, crRNA, ssRNA-FQ reporter (e.g., Uracil-containing, 5'-/6-FAM/rUrUrUrUrU/3IAbRQSp/-3'), Nuclease-Free Water, Reaction Buffer (40 mM Tris-HCl, 60 mM NaCl, 6 mM MgCl2, pH 7.3).
- Procedure:
- Reverse Transcription & Amplification: Prepare a 50 µL RT-RPA reaction. Incubate at 42°C for 20-30 minutes.
- CRISPR Detection Mix: Combine 50 nM LwCas13a, 50 nM crRNA, 500 nM ssRNA-FQ reporter in 1x Reaction Buffer. Incubate at 25°C for 10 minutes.
- Detection Reaction: Transfer 2 µL of the RT-RPA product into 23 µL of the CRISPR Detection Mix.
- Signal Measurement: Incubate at 37°C. Monitor fluorescence (Ex/Em: 485 nm/520 nm) in real-time. Signal accrual is often faster than Cas12-based assays.
- Data Analysis: Similar to DETECTR, with positive calls based on fluorescence kinetic curves or endpoint values.
5. Visualizations
Diagram 1: Cas12 & Cas13 Diagnostic Signal Transduction
Diagram 2: One-Pot CRISPR Diagnostic Workflow
6. The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for CRISPR-Cas Diagnostic Development
| Reagent / Material | Function | Example / Notes |
|---|---|---|
| Recombinant CRISPR-Cas Protein | Core detection enzyme. Purity and activity are critical. | Purified LbCas12a, AapCas12b, LwCas13a. Commercial sources (IDT, NEB) or in-house expression/purification. |
| Synthetic crRNA | Guides Cas protein to specific target sequence. Requires careful design to avoid off-target effects. | Chemically synthesized, HPLC-purified. Contains spacer sequence complementary to target. |
| Isothermal Amplification Kit | Amplifies target nucleic acid to detectable levels without complex thermocycling. | TwistAmp RPA kits, LAMP kits. Enables field-deployable, rapid pre-amplification. |
| Fluorescent-Quenched (FQ) Reporter | Signal-generating molecule cleaved during collateral activity. | ssDNA or ssRNA oligo with 5' fluorophore (FAM, HEX) and 3' quencher (BHQ, Iowa Black). |
| Lateral Flow Strip | For visual, instrument-free endpoint readout. Uses labelled reporter particles. | Often uses biotin and FAM labels captured on test/control lines. Requires adaptation of the detection reaction. |
| Nuclease-Free Buffers & Water | To prevent degradation of RNA/DNA reagents and ensure reaction specificity. | Essential for maintaining integrity of crRNAs, reporters, and templates. |
| Positive Control Template | Synthetic gene fragment or RNA transcript containing the target sequence. | Critical for assay validation, optimization, and establishing limit of detection (LoD). |
| Fluorescence Plate Reader or Handheld Fluorometer | For quantitative, real-time, or endpoint signal measurement. | Devices like BioTek Synergy or DeNovix DS-11 for lab use; low-cost portable readers for field use. |
Regulatory and Safety Considerations for Clinical Translation of Each System Type
Within the broader thesis of CRISPR-Cas system classification (Types I-VI) research, the translation of these powerful gene-editing tools into clinical applications presents a complex landscape of regulatory and safety challenges. Each system type—distinguished by its molecular architecture, effector complex, and mechanism of action—carries unique risk profiles and technical considerations that directly inform the pathway to clinical use. This guide provides a technical analysis of these considerations, focusing on the requirements for preclinical characterization, safety pharmacology, and regulatory submission for each major CRISPR-Cas type.
Core Safety Profiles by CRISPR-Cas Type
Table 1: Key Safety & Technical Characteristics by System Type
| CRISPR-Cas Type | Primary Risk Domains | Key Off-Target Potential Determinants | Immunogenicity Concerns | Primary Regulatory Focus (e.g., FDA, EMA) |
|---|---|---|---|---|
| Type II (Cas9) | Double-strand breaks (DSBs), large deletions, chromosomal rearrangements. | sgRNA design, PAM specificity, Cas9 variant fidelity, delivery method. | High (bacterial-derived protein; pre-existing antibodies). | Genotoxicity, tumorigenicity, off-target characterization. |
| Type V (Cas12a) | DSBs with staggered ends, potential for larger indel spectrum. | RuvC nuclease domain activity, PAM (TTTV) specificity. | Moderate to High. | Similar to Cas9, with added focus on unique indel profile consequences. |
| Type VI (Cas13) | Collateral RNA cleavage, transcriptome-wide perturbations. | RNA target specificity, collateral activity levels in human cells. | Moderate. | On-target RNA knockdown efficiency, collateral activity quantification, immunostimulation. |
| Type I (Cascade) | Large multi-protein complex, "shredding" of displaced strand. | Complex assembly requirements, processivity. | High (multiple bacterial proteins). | Manufacturing complexity, immunogenicity, precise cleavage control. |
| Type III (Csm/Cmr) | Dual RNA/DNA cleavage, cyclic oligoadenylate signaling. | Non-specific RNA cleavage upon DNA binding (Csm). | High (multiple proteins). | Complex activity control, potential for aberrant immune activation. |
| Base Editors (BE) | Single-base changes, but potential for bystander editing, guide-independent off-target DNA/RNA edits. | Editing window width, deaminase activity kinetics, gRNA-independent off-targets. | Moderate (fusion proteins). | Comprehensive off-target analysis (DNA & RNA), bystander edit quantification. |
| Prime Editors (PE) | Small insertions/deletions, reverse transcriptase template-driven edits. | PegRNA design, reverse transcriptase fidelity, potential for large deletions. | Moderate (fusion proteins). | Precision of intended edit, analysis of byproduct indels and large deletions. |
Preclinical Experimental Protocols for Safety Assessment
Protocol: CIRCLE-seq for Off-Target Profiling (Cas9, Cas12a)
Objective: Genome-wide, unbiased identification of off-target sites.
- Genomic DNA Isolation: Extract genomic DNA from target cell type (e.g., HEK293T, primary T-cells).
- In Vitro Cleavage: Incubate 1-5 µg of genomic DNA with purified RNP complex (Cas protein:sgRNA at 3:1 molar ratio) in appropriate reaction buffer for 16h at 37°C.
- Circularization: Blunt-end repair (if necessary), then ligate digested DNA using T4 DNA ligase to form circles. This step enriches for cleaved fragments.
- Phi29 Amplification: Perform rolling circle amplification (RCA) with Phi29 polymerase to linearly amplify circularized fragments.
- Library Prep & Sequencing: Fragment amplified DNA, prepare sequencing library (e.g., Nextera XT), and sequence on an Illumina platform.
- Bioinformatic Analysis: Map reads to reference genome, identify junctions indicative of in vitro cleavage sites. Validate top candidate sites in cells using targeted deep sequencing.
Protocol: RATESeq for Cas13 Collateral RNA Editing Assessment
Objective: Quantify transcriptome-wide collateral RNA degradation.
- Cell Treatment: Transfect target cells with Cas13 RNP or mRNA/gRNA.
- Total RNA Extraction: At 24h and 72h post-treatment, extract RNA (TRIzol method). Include untreated and catalytically dead (dCas13) controls.
- RNA-Seq Library Preparation: Deplete ribosomal RNA, prepare stranded RNA-seq libraries.
- Sequencing & Analysis: Sequence to high depth (>50M reads/sample). Align reads to transcriptome. Analyze for global reduction in transcript counts unrelated to the on-target, and specifically identify downregulated transcripts lacking complementarity to the gRNA.
Protocol: GOTI (Genome-wide Off-target analysis by Two-cell embryo Injection) for In Vivo Assessment
Objective: Detect off-target mutations in a whole-organism context.
- Mouse Embryo Generation: Harvest zygotes from superovulated female mice.
- Microinjection: Inject CRISPR components into one blastomere of a two-cell embryo. The other blastomere serves as an internal control.
- Embryo Transfer & Development: Transfer embryos to foster mothers, harvest at E14.5.
- Cell Sorting & WGS: Dissociate embryonic cells, sort edited and control cells from the same embryo (using a fluorescent co-injection marker). Perform whole-genome sequencing (WGS) on both populations to >30x coverage.
- Variant Calling: Identify somatic mutations present exclusively in the edited cell population.
Regulatory Pathway Visualization
Diagram Title: CRISPR Therapy Regulatory Pathway from Preclinical to Approval
Immunogenicity Assessment Workflow
Diagram Title: Immunogenicity Risk Assessment Workflow for CRISPR Proteins
The Scientist's Toolkit: Essential Reagents for Safety Assessment
Table 2: Key Research Reagent Solutions for Safety Studies
| Reagent/Material | Function in Safety Assessment | Example Vendor/Catalog | Critical Parameters |
|---|---|---|---|
| Recombinant Cas Nuclease (Active) | Positive control for in vitro off-target assays (CIRCLE-seq). | Integrated DNA Technologies, Thermo Fisher Scientific | High purity (>95%), endotoxin-free, verified specific activity. |
| Guide RNA (synthetic, chem mod) | Ensures consistent gRNA for comparing system variants. | Synthego, TriLink BioTechnologies | Chemical modifications (e.g., 2'-O-methyl, phosphorothioates) for stability. |
| Control gRNA (Non-targeting) | Essential control for distinguishing on/off-target effects in cellular assays. | Horizon Discovery, Dharmacon | Validated lack of targeting in human genome, matched length/modification. |
| dCas Protein (Catalytically Dead) | Critical negative control for separating editing from binding effects. | Applied Biological Materials, GenScript | Sequence-verified dead mutation(s) (e.g., D10A, H840A for SpCas9). |
| Genomic DNA from Target Cell Line | Substrate for unbiased off-target discovery assays. | ATCC, Coriell Institute | High molecular weight (>50 kb), from relevant cell type (e.g., iPSCs). |
| Targeted Deep Sequencing Panel | Validates predicted off-target sites from discovery assays. | Illumina TruSeq, Twist Bioscience | Must cover all candidate loci with high coverage (>1000x). |
| Relevant Animal Model | In vivo safety, biodistribution, immunogenicity studies. | Jackson Laboratory, Charles River | Immunocompetent model for immunogenicity; disease model for efficacy/tox. |
| Anti-Cas Antibody (ELISA) | Quantifies anti-drug antibodies in preclinical/clinical sera. | Antibody-specific vendors (custom) | Must be validated for sensitivity and specificity against the Cas variant used. |
| Primary Human Cells (PBMCs, T-cells) | Assess immunogenicity and functional editing in relevant human cells. | STEMCELL Technologies, AllCells | Donor variability requires multiple donors (n≥3). |
| NGS Library Prep Kit | For CIRCLE-seq, RNA-seq, WGS off-target analyses. | New England Biolabs, KAPA Biosystems | High fidelity, low bias, compatible with input from relevant assays. |
Table 3: Comparative Off-Target Rates (Representative Published Data)
| System | Target Locus | Detection Method | Reported Off-Target Sites | Validation Method | Reference (Example) |
|---|---|---|---|---|---|
| SpCas9 (WT) | VEGFA Site 3 | BLESS | >150 | Targeted Sequencing | Tsai et al., Nat Biotechnol, 2015 |
| SpCas9-HF1 | EMX1, VEGFA | GUIDE-seq | 0-1 (vs 4-10 for WT) | Targeted Sequencing | Kleinstiver et al., Nature, 2016 |
| AsCas12a | DNMT1 | Digenome-seq | ~10 | Targeted Sequencing | Kim et al., Nat Biotechnol, 2016 |
| LwaCas13a | Reporter RNA | RATESeq | Widespread collateral RNA effects | RNA-seq | Metsky et al., Nat Biotechnol, 2019 |
| ABE8e (Adenine BE) | HEK Site 4 | OT-seq | Very low DNA; detectable RNA deamination | Targeted & RNA-seq | Grünewald et al., Nature, 2019 |
Table 4: Regulatory Submission Elements with System-Specific Requirements
| IND Section | Cas9/Cas12a-Specific Requirements | Cas13-Specific Requirements | Base/Prime Editor-Specific Requirements |
|---|---|---|---|
| Pharmacology/Toxicology | Comprehensive genotoxicity (e.g., γH2AX, comet, karyotype). | Transcriptome-wide analysis (RNA-seq) for collateral damage. | Bystander editing analysis at target site; gRNA-independent DNA/RNA off-target screens. |
| Biodistribution/PK | Persistence of nuclease activity (time-course of indels). | Duration of RNA knockdown vs. protein presence. | Quantification of editing efficiency over time (stable vs. transient). |
| Immunogenicity | Assessment of pre-existing and treatment-induced anti-Cas humoral/cellular response. | Assessment of innate immune activation via RNA sensing pathways (RIG-I, MDA5). | Assessment of response to deaminase/RT fusion components. |
| CMC | Characterization of RNP complex or vector titer/identity; sgRNA purity. | Characterization of crRNA and protein components; stability of RNA guide. | Characterization of fusion protein integrity and linker stability. |
CRISPR-Cas systems are broadly classified into two classes, six types, and numerous subtypes based on their effector module architecture. Class 1 (Types I, III, IV) utilize multi-protein effector complexes, while Class 2 (Types II, V, VI) employ a single, large effector protein (e.g., Cas9, Cas12, Cas13). This classification provides the evolutionary and mechanistic backdrop against which engineered variants are developed. The drive to miniaturize these systems for viral vector delivery (especially AAV with its ~4.7 kb cargo limit) and to discover novel functionalities has led to the creation of hyper-compact tools like CasΦ and protein-engineered systems like CasMINI, pushing beyond the natural Class 2 archetypes.
Core Systems: CasMINI vs. CasΦ
2.1 CasMINI: A Protein-Engineered Hyper-Compact Variant CasMINI is not a naturally occurring protein but a computationally designed and extensively engineered derivative of the Type V-F Cas12f (formerly Cas14) system. The natural Cas12f (Un1Cas12f1) is compact (~400-700 amino acids) but exhibits low activity in mammalian cells. Through iterative rounds of protein engineering (directed evolution and rational design), the Stanford team created CasMINI, which shows robust gene activation and editing in human cells despite its small size (529 aa).
2.2 CasΦ (Cas12j): A Naturally Occurring Ultra-Compact System CasΦ (Cas12j) is a naturally occurring, ultra-compact (700-800 aa) CRISPR system discovered in huge bacteriophages. It belongs to the Type V CRISPR-Cas family (Class 2) but forms a distinct phylogenetic clade. Unlike other Cas12 proteins, CasΦ processes its own CRISPR RNA (crRNA) from a precursor, eliminating the need for a host-derived RNase III and tracrRNA. This self-sufficient, minimal system is highly active in vitro and in both prokaryotic and eukaryotic cells.
Quantitative Comparison Table
Table 1: Core Characteristics of Next-Generation Compact CRISPR Tools
| Feature | CasMINI (Engineered Cas12f) | CasΦ (Cas12j; e.g., Cas12j2) | Standard Cas9 (SpCas9) |
|---|---|---|---|
| Class/Type | Class 2, Type V (engineered from V-F) | Class 2, Type V (distinct clade) | Class 2, Type II |
| Size (aa) | ~529 | ~700-800 | ~1368 |
| Size (kb, coding DNA) | ~1.6 kb | ~2.1-2.4 kb | ~4.1 kb |
| Guide RNA Structure | Single crRNA (no tracrRNA required) | Single, self-processed pre-crRNA (no tracrRNA) | crRNA + tracrRNA duplex |
| PAM Requirement | 5'-T-rich (e.g., TTTN) | 5'-TBN (B=C/G/T) for Cas12j2; T-rich for others | 5'-NGG (SpCas9) |
| Cleavage Mechanism | Creates staggered DSBs with 5-7 nt overhangs | Creates staggered DSBs with 6-8 nt overhangs | Creates blunt-ended DSBs |
| In vivo Activity | Robust gene editing & activation in human cells | Demonstrated editing in human, plant, and bacterial cells | High activity in diverse cells |
| Key Advantage | Smallest engineered effector; ideal for AAV delivery | Naturally compact, self-contained RNA processing | High efficiency, well-characterized |
Detailed Experimental Protocols
4.1 Protocol for Assessing Gene Knockout Efficiency with CasΦ in Mammalian Cells
A. Materials & Reagents
- HEK293T cells
- CasΦ expression plasmid (e.g., pCMV-Cas12j2)
- Guide RNA expression plasmid (U6-driven pre-crRNA scaffold)
- Transfection reagent (e.g., Lipofectamine 3000)
- Lysis buffer (QuickExtract DNA Solution)
- PCR primers flanking target site
- T7 Endonuclease I (T7EI) or ICE analysis reagents
- NGS library prep kit for deep sequencing (optional)
B. Procedure
- Cell Seeding: Seed HEK293T cells in a 24-well plate to reach 70-80% confluency at transfection.
- Plasmid Transfection: For each target, prepare a transfection mix containing 500 ng of CasΦ plasmid and 250 ng of gRNA plasmid. Use a suitable transfection reagent per manufacturer's protocol.
- Incubation: Incubate cells for 48-72 hours at 37°C, 5% CO₂.
- Genomic DNA Harvest: Aspirate medium, wash with PBS, and add 100 µL of QuickExtract solution. Incubate at 65°C for 15 min, 98°C for 10 min, then hold at 4°C.
- Target Site Amplification: Use 2 µL of lysate as template for PCR with high-fidelity polymerase to amplify a 300-500 bp region surrounding the target site.
- Editing Analysis:
- T7EI Assay: Denature and reanneal PCR products. Digest with T7EI for 1 hour. Analyze fragments on a 2% agarose gel. Editing efficiency = 100% × [1 - √(1 - (b+c)/(a+b+c))], where a is intact band, b & c are cleavage products.
- NGS Analysis: Purify PCR products, prepare sequencing libraries, and perform high-depth sequencing (e.g., Illumina MiSeq). Analyze indels at the target site using CRISPResso2 software.
4.2 Protocol for Directed Evolution of CasMINI for Enhanced Activity
A. Materials & Reagents
- Yeast surface display library of Cas12f mutants
- Magnetic beads conjugated with anti-c-Myc antibody
- Biotinylated target DNA duplex containing PAM and protospacer
- Fluorescent streptavidin (e.g., SA-PE)
- FACS sorter
- Recovery media (SDCAA and SGCAA for yeast)
- Mammalian cell transfection system for secondary validation
B. Procedure
- Library Sorting: Induce expression of the Cas12f mutant library on yeast surface. Label yeast with anti-c-Myc antibody (to detect expression) and biotinylated target DNA + fluorescent streptavidin (to detect functional DNA binding).
- FACS Gating: Use FACS to sequentially gate for: 1) Healthy yeast, 2) High c-Myc signal (good expressors), 3) High biotin/streptavidin signal (strong DNA binders). Collect the top 1-2% of the double-positive population.
- Recovery & Amplification: Grow sorted yeast in recovery media, then extract plasmid DNA.
- Iterative Rounds: Use the plasmid DNA to create a new, mutated library for the next round of selection, increasing selection stringency (e.g., shorter incubation time).
- Mammalian Cell Screening: Clone individual enriched mutants into mammalian expression vectors, co-transfect with a gRNA and a fluorescent reporter (e.g., GFP activation assay), and quantify activity via flow cytometry to identify top performers like CasMINI.
Visualization of System Architecture & Workflows
Diagram Title: CasMINI Protein Engineering Workflow
Diagram Title: CasΦ Discovery and Application Pathway
Diagram Title: CasΦ Self-Processing and Cleavage Mechanism
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for Compact CRISPR Tool Research
| Reagent/Material | Function & Application |
|---|---|
| AAV Packaging System (serotype 9) | For in vivo delivery of compact Cas genes; small size allows room for regulatory elements and multiple gRNAs. |
| Yeast Surface Display Library Kit | Critical for directed evolution projects (e.g., creating CasMINI) to select for improved binding/function mutants. |
| T7 Endonuclease I (T7EI) | Rapid, cost-effective tool for initial assessment of nuclease-induced indel formation in mammalian cells. |
| Next-Generation Sequencing (NGS) Service | Gold-standard for quantitative, unbiased measurement of editing efficiency, specificity, and off-target analysis. |
| U6-sgRNA Cloning Vector | Standardized backbone for expressing gRNAs (crRNAs) in mammalian cells; easily adaptable for CasΦ or CasMINI scaffolds. |
| QuickExtract DNA Solution | Enables fast, column-free genomic DNA extraction from cultured cells for PCR-based genotyping. |
| Lipofectamine 3000/CRISPRMAX | High-efficiency transfection reagents for delivering plasmid or RNP complexes into hard-to-transfect cell lines. |
| Anti-c-Myc Magnetic Beads | For purification or detection of epitope-tagged (c-Myc) Cas proteins during surface display or pull-down assays. |
Conclusion
The intricate classification of CRISPR-Cas systems from Types I to VI represents a vast, programmable toolkit for biomedical science, each with unique architectural and functional profiles. While Type II (Cas9) remains foundational for DNA editing, Types V (Cas12) and VI (Cas13) have revolutionized diagnostics and RNA targeting, and lesser-used types offer novel mechanisms for future exploitation. Successful application requires matching the system's inherent properties—its PAM specificity, cleavage mechanism, and molecular size—to the experimental goal, whether it's high-fidelity genome editing, sensitive detection, or multiplex regulation. Future directions will focus on engineering novel effectors with enhanced capabilities, improving delivery vectors for therapeutic use, and deepening our understanding of type-specific immunogenicity and safety. For researchers and drug developers, a strategic, comparative approach to selecting and validating the appropriate CRISPR-Cas system is paramount for advancing both basic research and next-generation clinical interventions.