CRISPR-Cas9 Gene Editing for Sickle Cell Disease: A Comprehensive Guide for Researchers and Developers
This article provides a detailed, technical overview of the CRISPR-Cas9 mechanism as applied to sickle cell disease (SCD) gene editing.
CRISPR-Cas9 Gene Editing for Sickle Cell Disease: A Comprehensive Guide for Researchers and Developers
Abstract
This article provides a detailed, technical overview of the CRISPR-Cas9 mechanism as applied to sickle cell disease (SCD) gene editing. We explore the foundational molecular basis of SCD and CRISPR-Cas9, examine the methodological strategies for targeting the HBB gene, discuss critical troubleshooting and optimization challenges in ex vivo and in vivo approaches, and validate outcomes through comparative analysis of recent clinical trials and emerging technologies. Designed for researchers, scientists, and drug development professionals, this review synthesizes current advancements and practical considerations for translating CRISPR-based therapies from bench to bedside.
Understanding the Target: Molecular Pathology of SCD and CRISPR-Cas9 Fundamentals
This whitepaper details the foundational molecular pathology of sickle cell disease (SCD), a monogenic disorder caused by a single nucleotide substitution in the HBB gene. This precise genetic understanding is the essential prerequisite for contemporary gene-editing therapeutics, most notably CRISPR-Cas9-based approaches. Current clinical strategies, such as ex vivo editing of the BCL11A enhancer to induce fetal hemoglobin or the direct correction of the HBB mutation, rely entirely on the mechanistic elucidation of hemoglobin S (HbS) polymerization presented herein. This document serves as a technical reference for researchers engineering next-generation gene therapies.
Molecular Genetics of the HBB Mutation
The primary genetic lesion is a homozygous A>T point mutation (E6V) at the seventh nucleotide of the HBB gene's sixth exon (HBB: c.20A>T; p.Glu6Val). This autosomal recessive mutation results in the production of the abnormal β-globin chain (βS).
Table 1: Quantitative Genetic and Molecular Data for SCD
| Parameter | Value / Specification | Notes / Clinical Correlation |
|---|---|---|
| Gene Locus | 11p15.4 | HBB gene encoding β-globin. |
| Mutation (Hg38) | chr11:5,248,233 (A>T) | rs334 (sickle cell variant). |
| mRNA Change | GAG (Glu) → GUG (Val) at codon 6 | Reference sequence NM_000518.5. |
| Inheritance Pattern | Autosomal Recessive | Heterozygotes (HbAS) have sickle cell trait. |
| Global Carrier Frequency | ~1 in 50 | Varies geographically; highest in sub-Saharan Africa. |
| HbS Polymer Critical Concentration | ~17 g/dL (deoxygenated) | Concentration-dependent polymerization kinetics. |
Hemoglobin S Polymerization: Mechanism & Pathophysiology
The substitution of valine for glutamic acid introduces a hydrophobic patch on the surface of the βS-globin chain. Upon deoxygenation, hemoglobin tetramers undergo a conformational change that exposes this valine. It can then interact with a complementary hydrophobic acceptor pocket (containing Phe85 and Leu88) on a β-chain of a neighboring hemoglobin tetramer.
This interaction initiates the formation of a double-stranded helical fiber. These fibers align and bundle into large, rigid polymers that distort the red blood cell (RBC) into the characteristic sickle shape. Polymerization is a delay-time event, governed by nucleation-controlled kinetics.
Diagram 1: HbS Polymerization Pathway & Cellular Consequences
Key Experimental Protocols for Studying HbS Polymerization
Protocol:In VitroHbS Polymerization Kinetics by Turbidimetry
Objective: To measure the delay time prior to polymer formation and the subsequent rate of polymerization under controlled deoxygenation.
- HbS Purification: Isolate HbS from patient RBCs via lysis, centrifugation, and ion-exchange chromatography (e.g., DEAE Sephadex). Confirm purity by HPLC and mass spectrometry.
- Sample Preparation: Dialyze HbS into a phosphate buffer (e.g., 0.15 M potassium phosphate, pH 7.35). Concentrate to a target Hb concentration (e.g., 25 g/dL) using centrifugal concentrators. Add sodium dithionite (Na₂S₂O₄) to a final concentration of 2-5 mg/mL as a reducing and deoxygenating agent.
- Turbidimetry: Immediately transfer the deoxygenated sample to a sealed cuvette in a spectrophotometer thermostatted to 37°C. Monitor absorbance at 700 nm (turbidity) over time.
- Data Analysis: The tracing will show a lag phase (delay time, t_d), followed by a rapid increase in turbidity. The delay time is inversely proportional to the ~30th power of the HbS concentration, demonstrating extreme concentration dependence.
Protocol: Electron Microscopy of HbS Fibers
Objective: To visualize the structure of deoxygenated HbS polymers.
- Polymerization: Induce polymerization of a concentrated HbS solution (as in 4.1) in a sealed chamber.
- Fixation & Embedding: Gently introduce glutaraldehyde (2.5% in buffer) to fix the polymerized fibers in situ. Dehydrate the sample through an ethanol series.
- Negative Staining: Apply a small aliquot of the fixed fiber suspension to a carbon-coated EM grid. Stain with 1-2% uranyl acetate for 30-60 seconds, then wick away excess.
- Imaging: Visualize using transmission electron microscopy (TEM) at 80-100 kV. Fibers appear as twisted, rope-like structures approximately 21 nm in diameter.
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for SCD Molecular Research
| Reagent / Material | Function & Application in SCD Research |
|---|---|
| Sodium Dithionite (Na₂S₂O₄) | Chemical deoxygenator used to induce HbS polymerization in vitro for kinetic and structural studies. |
| DEAE Sepharose / Cellulose | Ion-exchange chromatography medium for purification of hemoglobin variants (HbA, HbS, HbF) from hemolysates. |
| Hypoxanthine & Xanthine Oxidase | Enzymatic deoxygenation system for gentler, more controlled oxygen removal in polymerization assays. |
| CO-Saturated Buffer | Used to carbonmonoxy-ligate hemoglobin, preventing polymerization and stabilizing Hb during purification. |
| 2,3-Bisphosphoglycerate (2,3-BPG) Analogues | Allosteric effectors that reduce oxygen affinity, used to study polymer stability under physiological conditions. |
| CRISPR-Cas9 Ribonucleoprotein (RNP) | Pre-complexed Cas9 protein and sgRNA for precise genome editing in hematopoietic stem cells (HSCs). |
| Recombinant Adeno-Associated Virus (rAAV6) | Common delivery vector for donor DNA templates in HDR-based correction of the HBB E6V mutation in HSCs. |
| BCL11A-Targeting sgRNA | Guide RNA designed to disrupt the erythroid enhancer of BCL11A, a repressor of fetal hemoglobin (HbF). |
| Methylcellulose-based Colony Forming Unit (CFU) Assays | To assess the proliferative and differentiation potential of gene-edited CD34+ HSCs in vitro. |
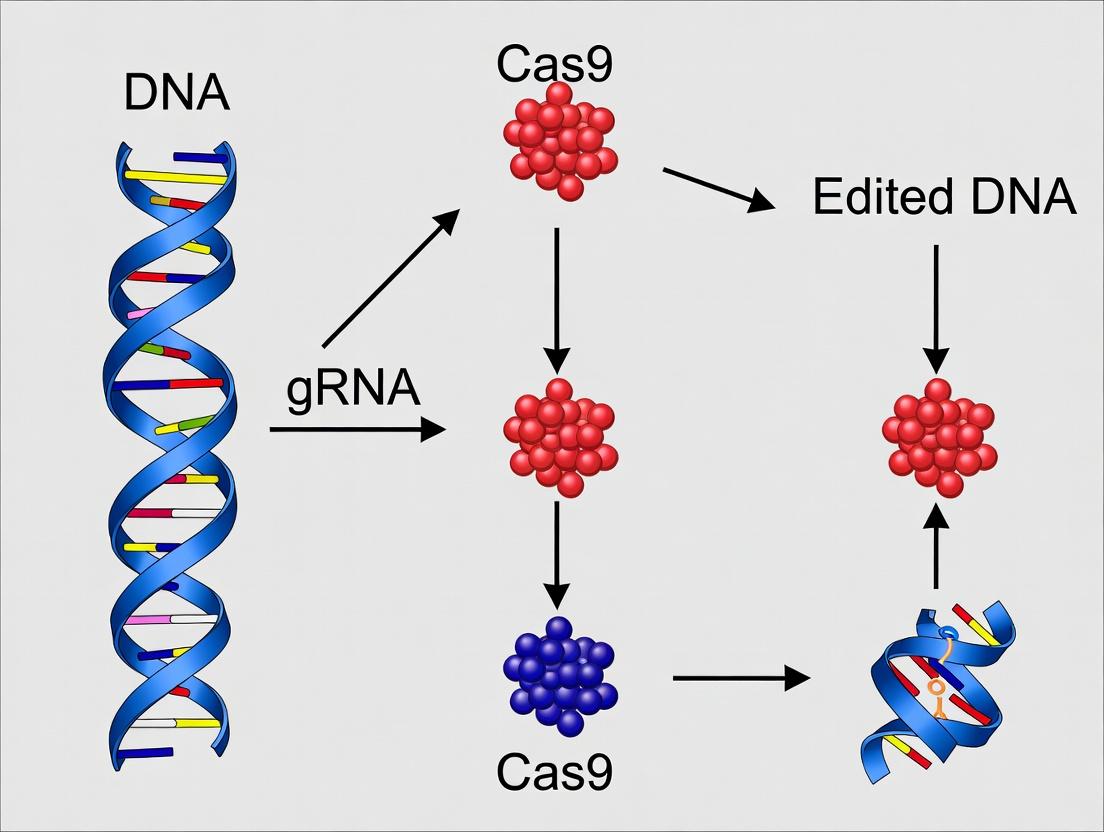
Diagram 2: CRISPR-Cas9 Gene Editing Strategies for SCD
This whitepaper provides an in-depth technical guide to the core CRISPR-Cas9 mechanism, framed within the critical context of its application for sickle cell disease (SCD) gene editing research. The pathological GAG-to-GTG point mutation in the β-globin gene (HBB) results in the production of sickle hemoglobin (HbS), and precise correction of this mutation represents a paradigm for therapeutic genome editing.
Core Mechanism: The Ribonucleoprotein (RNP) Complex
The functional unit is a ribonucleoprotein complex comprising the Cas9 endonuclease and a single-guide RNA (sgRNA). The sgRNA is a chimeric RNA with a 20-nucleotide spacer sequence at its 5’ end that dictates target specificity through Watson-Crick base pairing, and a scaffold sequence that binds Cas9.
Key Quantitative Parameters of CRISPR-Cas9 Targeting:
| Parameter | Typical Range/Value | Relevance to SCD Targeting |
|---|---|---|
| sgRNA Spacer Length | 20 nucleotides (nt) | Defines target site within the HBB gene. |
| Protospacer Adjacent Motif (PAM) | NGG (for S. pyogenes Cas9) | Must be present 3’ of the target DNA. Limits targetable sites. |
| DNA Cleavage Position | 3 bp upstream of PAM | Generates a blunt-ended double-strand break (DSB). |
| On-target Editing Efficiency (in vitro) | 40-80% (varies by cell type) | Must be high in hematopoietic stem/progenitor cells (HSPCs). |
| Off-target Potential | Site-dependent; can be >100 sites | Requires careful sgRNA design and validation (e.g., CIRCLE-seq). |
DNA Repair Pathways: NHEJ vs. HDR
The Cas9-induced DSB is resolved by endogenous cellular repair pathways, primarily Non-Homologous End Joining (NHEJ) or Homology-Directed Repair (HDR). The choice between these pathways is the fundamental determinant of the editing outcome.
Comparative Analysis of DNA Repair Pathways in SCD Gene Editing:
| Feature | Non-Homologous End Joining (NHEJ) | Homology-Directed Repair (HDR) |
|---|---|---|
| Primary Role | Error-prone repair; ligates broken ends. | High-fidelity repair using a homologous template. |
| Cell Cycle Phase | Active throughout, dominant in G0/G1. | Active primarily in S/G2 phases. |
| Template Required? | No. | Yes, exogenous donor DNA template. |
| Outcome for SCD | Knock-out: Can disrupt BCL11A enhancer to induce fetal hemoglobin (HbF) as a therapeutic strategy. | Knock-in/Correction: Precisely corrects the E6V point mutation in the HBB gene. |
| Efficiency in HSPCs | Typically high (>70% indels possible). | Typically low (often 10-40%, depends on delivery). |
| Byproducts | Small insertions/deletions (indels). | Precise nucleotide change with no indels (if designed correctly). |
Experimental Protocols for SCD-Relevant CRISPR-Cas9 Research
Protocol 1: Assessment of On- and Off-target Editing (Next-Generation Sequencing)
- Design & Cloning: Design sgRNA targeting the HBB gene or BCL11A erythroid enhancer. Clone into an expression plasmid (e.g., pSpCas9(BB)-2A-GFP).
- Delivery: Deliver plasmid or pre-formed RNP (Cas9 protein + synthetic sgRNA) into target cells (e.g., CD34+ HSPCs) via electroporation.
- Genomic DNA Extraction: Harvest cells 72-96 hours post-editing. Extract gDNA using a column-based kit.
- PCR Amplification: Amplify the on-target genomic locus and predicted top off-target sites (from algorithms like Cas-OFFinder) using high-fidelity polymerase.
- NGS Library Prep & Sequencing: Purify PCR products, tag with Illumina adapters, and sequence on a MiSeq or similar platform.
- Data Analysis: Align sequences to reference genome. Use CRISPResso2 or similar tool to quantify indel frequencies (for NHEJ) or HDR rates.
Protocol 2: HDR-Mediated Correction in HSPCs Using ssODN Donor
- RNP Formation: Complex recombinant S. pyogenes Cas9 protein with chemically modified, target-specific sgRNA at a 1:2 molar ratio in an electroporation buffer. Incubate 10 min at room temperature.
- Donor Design: Co-deliver a single-stranded oligodeoxynucleotide (ssODN) donor template (~100-200 nt). The template must contain the corrected nucleotide sequence (GAG) flanked by homologous arms (50-90 nt each) complementary to the target locus.
- Electroporation: Mix 1-2e5 CD34+ HSPCs with RNP and ssODN donor (e.g., 2 µM RNP, 1 µM donor). Electroporate using a optimized program (e.g., pulse code EH-115 on Lonza 4D-Nucleofector).
- Culture & Analysis: Culture cells in cytokine-enriched serum-free medium. After 48-72 hours, assay a subset for editing efficiency (by T7E1 assay or NGS). Proceed with differentiation assays (erythroid differentiation) and functional analysis (HPLC for HbA/HbS).
Visualizing the Core Mechanism and Pathways
Title: CRISPR-Cas9 RNP Complex Formation and DNA Targeting
Title: Cellular Decision Between NHEJ and HDR Repair Pathways
The Scientist's Toolkit: Key Reagent Solutions for SCD Gene Editing
| Research Reagent / Material | Function in SCD CRISPR Research |
|---|---|
| Recombinant S. pyogenes Cas9 Protein | High-purity, endotoxin-free nuclease for forming RNP complexes. Reduces plasmid DNA integration risk and enables rapid kinetics. |
| Chemically Modified sgRNA (synthetic) | Incorporates 2'-O-methyl and phosphorothioate modifications at terminal nucleotides to enhance stability and reduce immune activation in primary HSPCs. |
| Single-Stranded Oligodeoxynucleotide (ssODN) | ~150-nt donor template for HDR-mediated correction of the HBB point mutation. Homology arms flank the corrective change. |
| CD34+ Human Hematopoietic Stem/Progenitor Cells (HSPCs) | Primary target cell population for ex vivo SCD therapy. Source: mobilized peripheral blood or cord blood. |
| Electroporation System (e.g., Lonza 4D-Nucleofector) | Device for high-efficiency, low-toxicity delivery of RNP and donor template into hard-to-transfect HSPCs. |
| Cytokine Cocktail (SCF, TPO, FLT3L) | Essential for maintaining stemness and viability of edited CD34+ cells during post-electroporation culture. |
| Next-Generation Sequencing (NGS) Assay Kits | For comprehensive on-target efficiency and genome-wide off-target profiling (e.g., using GUIDE-seq or CIRCLE-seq methodologies). |
| Erythroid Differentiation Media | Specialized culture medium to differentiate edited HSPCs into erythroid lineages for functional validation of HbS correction via HPLC. |
Within the broader thesis on CRISPR-Cas9 mechanisms for sickle cell disease (SCD) research, a critical strategic decision is the choice between two fundamental editing approaches: directly correcting the causative point mutation in the HBB gene (β-globin, Glu6Val) or disrupting the mutant allele via targeted lesions to induce fetal hemoglobin (HbF). This whitepaper provides an in-depth technical comparison of these paradigms, detailing experimental protocols, quantitative outcomes, and essential research tools.
Molecular Targets and Rationale
Correcting the Mutant Allele: Aims to restore wild-type β^A-globin sequence in hematopoietic stem and progenitor cells (HSPCs). This precise correction requires homology-directed repair (HDR) using an exogenous DNA template.
Disrupting the Mutant Allele: Primarily targets the HBG1/HBG2 gene promoters or the BCL11A erythroid enhancer to de-repress γ-globin (HbF) expression, compensating for defective β^S-globin. This relies on non-homologous end joining (NHEJ) to create disruptive insertions/deletions (indels).
Table 1: Comparison of Key Editing Outcomes from Recent Clinical & Preclinical Studies
| Parameter | Correction of HBB (HDR-based) | Disruption of BCL11A Enhancer (NHEJ-based) | Disruption of HBG Promoter (NHEJ-based) |
|---|---|---|---|
| Target Locus | HBB codon 6 | BCL11A +58 DHS erythroid enhancer | HBG1/HBG2 promoter (~200bp upstream of TSS) |
| Primary Edit | A>T reversion to wild-type | Indels disrupting GATA1 binding site | Indels disrupting BCL11A/ZBTB7A binding motifs |
| Therapeutic Effect | Produces β^A-globin | Reduces BCL11A, upregulates γ-globin | Directly de-represses γ-globin transcription |
| Typical Ex Vivo Editing Efficiency (HSPCs) | 15-30% HDR (with inhibitors) | 70-90% indels | 80-95% indels |
| HbF Increase (Post-Engraftment) | Not applicable (makes HbA) | 20-30% of total hemoglobin | 15-25% of total hemoglobin |
| Clinical Trial Phase (Example) | Phase 1/2 (NCT04819841) | Phase 1/2 (NCT03745287) | Phase 1/2 (NCT05444894) |
| Key Risk/Challenge | Lower HDR efficiency; requires precise template delivery. | Potential off-target effects at BCL11A coding region. | Potential disruption of HBG ORF; on-target specificity. |
Table 2: Key Biochemical Reagents for Enhancing Editing Outcomes
| Reagent | Function in Correction Strategy | Function in Disruption Strategy |
|---|---|---|
| Alt-R HDR Enhancer (IDT) | Inhibits NHEJ, improves HDR rates by 2-3 fold. | Not typically used. |
| Scaffold-Modified sgRNA (2'-O-methyl) | Increases stability and reduces immune response in primary cells. | Increases stability and on-target activity for high-efficiency disruption. |
| Recombinant Cas9 Protein (HiFi) | High-fidelity variant reduces off-target cleavage. | Critical for both strategies; HiFi variant preferred for therapeutic development. |
| AAV6 Serotype Vector | Delivers HDR template with high efficiency in HSPCs. | Not used for disruption-only strategies. |
| Small Molecule NHEJ Inhibitors (e.g., SCR7) | Synergizes with HDR enhancers. | Actively avoided to maximize disruptive indels. |
Experimental Protocols
Protocol 4.1: Ex Vivo Correction of HBB in CD34+ HSPCs via HDR
Objective: Precise correction of the HBB Glu6Val mutation using CRISPR-Cas9 RNP and an AAV6 HDR template.
- Mobilization & Isolation: Isolate human CD34+ HSPCs from mobilized peripheral blood or cord blood using clinical-grade CD34+ magnetic bead selection.
- Electroporation: Pre-complex Alt-R S.p. HiFi Cas9 protein (100 pmol) and chemically modified sgRNA (120 pmol) targeting near the HBB mutation to form RNP. Electroporate 1e5 cells per reaction using the Lonza 4D-Nucleofector (P3 kit, program DZ-100). Include Alt-R HDR Enhancer V2 at recommended concentration.
- AAV6 Transduction: Immediately post-electroporation, transduce cells with AAV6 vector containing the HDR template (homology arms ~800bp, containing corrected sequence and a silent restriction site for screening) at an MOI of 1e5 vg/cell.
- Culture & Analysis: Culture cells in StemSpan SFEM II with cytokines (SCF, TPO, FLT3-L). After 48-72h, harvest genomic DNA. Assess HDR efficiency via restriction fragment length polymorphism (RFLP) if a silent site was introduced, or by next-generation sequencing (NGS) of the target locus.
Protocol 4.2: Disruption of the BCL11A Erythroid Enhancer in HSPCs via NHEJ
Objective: Generate indels in the +58 enhancer region of BCL11A to reduce its expression and induce HbF.
- Cell Preparation: Isolate CD34+ HSPCs as in 4.1.
- RNP Electroporation: Pre-complex HiFi Cas9 protein (100 pmol) with sgRNA targeting the GATA motif in the BCL11A +58 DHS enhancer (sequence: 5'-GCCCACAGTGGCACCACGAG-3'). Electroporate 1e5 cells (identical conditions to 4.1). Do not add HDR enhancers.
- Differentiation & Validation: Culture a portion of cells for 3 days in expansion medium, then extract gDNA for T7 Endonuclease I or ICE analysis to determine indel %. Differentiate another portion in erythroid differentiation medium (IL-3, SCF, EPO) for 14-21 days.
- Functional Readout: Perform flow cytometry for HbF (F-cell staining) and HPLC for hemoglobin quantitation. Perform RT-qPCR for BCL11A and HBG mRNA levels.
The Scientist's Toolkit: Research Reagent Solutions
| Item | Vendor Examples (Catalog #) | Function & Application Notes |
|---|---|---|
| Clinical-Grade CD34+ Isolation Kit | Miltenyi Biotec (CliniMACS) | GMP-compatible immunomagnetic selection of target HSPCs. |
| S.p. HiFi Cas9 Nuclease V3 | Integrated DNA Technologies (Alt-R 1081060) | High-specificity Cas9 variant; reduces off-target effects for both strategies. |
| AAV6 Helper-Free System | Cell Biolabs (VPK-420) | Production of AAV6 vectors for high-efficiency HDR template delivery in HSPCs. |
| Genomic DNA Cleavage Detection Kit | NEB (T7 Endonuclease I, M0302L) | Fast, cost-effective validation of editing efficiency (indels). |
| Next-Gen Sequencing Kit for Editing | Illumina (Miseq CRISPR Amplicon) | Gold-standard for quantifying HDR and indel frequencies and profiling outcomes. |
| Erythroid Differentiation Media Kit | STEMCELL Technologies (HemaCare) | Standardized culture system for in vitro erythroid differentiation from HSPCs. |
| HbF Intracellular Flow Kit | BD Biosciences (FITC Anti-HbF, 552828) | Quantify percentage of F-cells (HbF-positive RBCs) post-editing/differentiation. |
Visualizations
Diagram 1: HDR Pathway for HBB Correction
Diagram 2: Disruption via BCL11A Enhancer
Diagram 3: SCD Gene Editing Strategy Decision Workflow
The application of CRISPR-Cas9 for sickle cell disease (SCD) gene editing research is founded on correcting the underlying genetic pathology. The primary mutation is an A-to-T transversion in the sixth codon of the β-globin gene (HBB), resulting in the production of pathological hemoglobin S (HbS). Current CRISPR-based therapeutic strategies bifurcate into two principal approaches: (1) Direct HBB Gene Correction, aimed at repairing the causative point mutation to restore adult hemoglobin (HbA) production, and (2) Disruption of the BCL11A Erythroid Enhancer, a trans-acting, developmentally regulated silencer of fetal hemoglobin (HbF). Reactivating HbF is therapeutic because it dilutes HbS and inhibits its polymerization. This whitepaper provides a technical comparison of these two target loci, detailing mechanisms, experimental protocols, and research tools.
Target Locus Comparison: Molecular Mechanisms and Quantitative Outcomes
BCL11A Erythroid Enhancer Disruption
The +58 BCL11A erythroid enhancer, located within intron 2 of the BCL11A gene, is a GATA1/TAL1/LDB1/LMO2 complex-binding site critical for high BCL11A expression in erythroid cells. BCL11A is a zinc-finger transcription factor that represses γ-globin (HBG1/HBG2) expression. Disruption of this enhancer via CRISPR-Cas9-induced double-strand breaks (DSBs) and non-homologous end joining (NHEJ) reduces BCL11A expression specifically in the erythroid lineage, leading to de-repression of HbF.
Direct HBB Gene Correction
This approach uses CRISPR-Cas9 to induce a DSB near the E6V mutation, co-delivered with a donor DNA template containing the correct sequence. Repair via homology-directed repair (HDR) results in the precise correction of the mutation, restoring normal β-globin (HBB) production.
Table 1: Comparative Quantitative Data of Key Genomic Targets
| Parameter | BCL11A Enhancer Targeting | Direct HBB Gene Correction |
|---|---|---|
| Primary Edit Type | NHEJ-mediated indel (knockout of enhancer function) | HDR-mediated precise point mutation correction |
| Therapeutic Effect | Reactivation of Fetal Hemoglobin (HbF) | Restoration of Adult Hemoglobin (HbA) |
| Typical Editing Efficiency (in CD34+ HSPCs) | 70-90% allele modification | 20-40% allele correction (HDR is less efficient) |
| Resulting HbF/HbA Levels | HbF can reach 20-40% of total hemoglobin | HbA restoration proportional to HDR efficiency; theoretical 100% in corrected cells |
| Key Risk/Challenge | Potential for BCL11A haploinsufficiency effects; off-targets in related enhancers. | Lower HDR efficiency in primitive HSPCs; requires donor template; risk of oncogenic indels at cut site. |
| Clinical Stage (as of 2024) | FDA-approved (exagamglogene autotemcel) | Multiple Phase 1/2 trials ongoing (e.g., EDIT-301, BIVV003) |
Experimental Protocols for Key Validation Experiments
Protocol: AssessingBCL11AEnhancer Editing and HbF Reactivation in Erythroid Differentiation Cultures
- Isolation and Editing: Isolate human CD34+ hematopoietic stem and progenitor cells (HSPCs) from mobilized peripheral blood or cord blood. Electroporate with RNP complexes comprising S. pyogenes Cas9 protein and a single guide RNA (sgRNA) targeting the +58 BCL11A erythroid enhancer (e.g., sequence: 5'-GAGTCTGTGCTCAGGAAGCA-3').
- Culture and Differentiation: Post-editing, culture cells in a three-phase erythroid differentiation medium.
- Phase 1 (Days 0-7): Expansion in SFEM II with cytokines (SCF, FLT3-L, TPO, IL-3, IL-6).
- Phase 2 (Days 7-11): Differentiation in media with erythropoietin (EPO), stem cell factor (SCF), and dexamethasone.
- Phase 3 (Days 11-18): Maturation in media with high EPO and insulin.
- Analysis:
- Flow Cytometry for HbF: On day 18, fix and permeabilize cells. Stain with FITC-conjugated anti-HbF antibody and APC-conjugated anti-glycophorin A (CD235a) antibody. Analyze HbF positivity in the erythroid (CD235a+) population.
- Molecular Analysis: Perform T7 Endonuclease I or next-generation sequencing (NGS) on genomic DNA from day 0 cells to quantify indel percentage. Use RT-qPCR on day 11-14 RNA to assess BCL11A and HBG mRNA levels.
Protocol: Assessing Direct HBB Correction via HDR
- Design and Delivery: Design a sgRNA to cut 5-10 bp upstream of the E6V mutation. Co-electroporate HSPCs with Cas9 RNP and a single-stranded oligodeoxynucleotide (ssODN) donor template (~100-200 nt). The donor should contain the wild-type codon (GAG) and silent blocking mutations in the PAM/protospacer to prevent re-cutting.
- Culture and Engraftment: Transplant edited HSPCs into immunodeficient NSG mice for in vivo assessment.
- Analysis:
- Primary HDR Assessment: 48-72 hours post-electroporation, harvest cells for genomic DNA. Use droplet digital PCR (ddPCR) with allele-specific probes (FAM for corrected, HEX for uncorrected) to quantify HDR efficiency.
- In Vivo Analysis: At 16 weeks post-transplant, analyze human cell engraftment in bone marrow. Isulate genomic DNA and perform NGS on the HBB locus to measure the percentage of perfectly corrected alleles and indels. Perform HPLC on peripheral blood to detect human HbA production.
Visualization of Pathways and Workflows
Title: BCL11A Enhancer Disruption Mechanism for HbF Reactivation
Title: Direct HBB Gene Correction Workflow via HDR
The Scientist's Toolkit: Essential Research Reagent Solutions
Table 2: Key Reagents for CRISPR-Based SCD Gene Editing Research
| Reagent / Material | Function / Application | Example/Note |
|---|---|---|
| Human CD34+ HSPCs | Primary cell model for ex vivo editing and differentiation. | Sourced from cord blood, mobilized peripheral blood, or commercially available frozen vials. |
| S. pyogenes Cas9 Nuclease | The effector enzyme that creates DSBs at the DNA target site. | Used as purified protein for RNP formation (enhanced kinetics, reduced off-target time). |
| Chemically Modified sgRNAs | Guides Cas9 to the specific genomic locus (BCL11A enhancer or HBB gene). | Chemical modifications (e.g., 2'-O-methyl, phosphorothioate) improve stability and reduce immunogenicity. |
| ssODN Donor Template | Provides the homologous template for precise HDR-mediated correction of the HBB gene. | Designed with silent blocking mutations; HPLC-purified. |
| Electroporation System | Method for delivering RNP and donor templates into hard-to-transfect HSPCs. | e.g., Lonza 4D-Nucleofector with P3 Primary Cell Kit. |
| Erythroid Differentiation Media Kits | Supports the in vitro maturation of HSPCs into enucleated erythrocytes. | e.g., STEMdiff Erythroid Differentiation Kit, or custom cytokine formulations. |
| Anti-HbF Antibody (FITC) | Flow cytometry-based detection and quantification of HbF-positive erythroid cells. | Critical for functional readout of BCL11A targeting experiments. |
| NGS-based Off-Target Assay Kits | Identifies potential off-target editing events across the genome. | e.g., CIRCLE-seq or GUIDE-seq workflows; essential for preclinical safety profiling. |
| Immunodeficient Mouse Model (NSG) | In vivo model to assess long-term engraftment, safety, and efficacy of edited HSPCs. | NOD.Cg-Prkdcscid Il2rgtm1Wjl/SzJ (NSG) mice are the standard. |
From Design to Delivery: Strategies for Editing Hematopoietic Stem Cells
Guide RNA (gRNA) Design and Optimization for HBB Locus or BCL11A Enhancer
Sickle cell disease (SCD) is caused by a single nucleotide substitution (A>T) in the β-globin gene (HBB), leading to the production of pathogenic hemoglobin S (HbS). CRISPR-Cas9 gene editing offers two primary therapeutic strategies targeting the HBB locus: direct correction of the sickle mutation or inhibition of BCL11A, a transcriptional repressor of fetal hemoglobin (HbF). The latter involves disrupting a erythroid-specific enhancer within the BCL11A gene to de-repress HbF production, which can compensate for defective adult β-globin. This guide details the technical design and optimization of guide RNAs (gRNAs) for these two critical targets, a foundational step in developing a curative CRISPR-based therapy for SCD.
Target Locus Genomics and Considerations
1HBBGene Target
The aim is to correct the E6V mutation (codon 6, GAG>GTG) or knock-in a corrective sequence via HDR. The target region is within exon 1 of HBB.
2BCL11AEnhancer Target
The target is a GATA1-binding site within the +58 DNase I hypersensitive site (DHS) of the erythroid-specific enhancer in intron 2 of BCL11A (chr2:60,466,389-60,467,368, hg38). Disruption here reduces BCL11A expression in erythroid cells, thereby increasing γ-globin expression.
Core gRNA Design Principles
General Design Parameters
- Length: 20 nucleotides preceding the 5'-NGG-3' Protospacer Adjacent Motif (PAM) for Streptococcus pyogenes Cas9 (SpCas9).
- Seed Region: Bases 1-12 proximal to the PAM are critical for specificity.
- GC Content: Optimal between 40-60%.
- Off-Target Prediction: Essential to minimize potential cleavage at homologous genomic sites.
Target-Specific Optimization
- For HBB Correction: The gRNA must bind immediately adjacent to the mutant nucleotide to facilitate precise homology-directed repair (HDR).
- For BCL11A Enhancer Disruption: gRNAs are designed to create double-strand breaks (DSBs) within the core enhancer sequence, ideally disrupting transcription factor binding motifs, leading to error-prone non-homologous end joining (NHEJ).
Table 1: Representative gRNA Sequences and Predicted Efficiency Scores
| Target | gRNA Sequence (5' to 3', PAM excluded) | PAM | Strand | Predicted On-Target Efficiency* | Primary Purpose |
|---|---|---|---|---|---|
| HBB (Corrective) | GGTGAAGCTGGTGGCGTAG | CGG | + | 68 | HDR template for E6V correction |
| BCL11A Enhancer | GACAAGGGTAGGAGAAATGC | TGG | - | 85 | Disruption via NHEJ |
| BCL11A Enhancer | GTCACTGCCACACCTGGCA | AGG | + | 72 | Disruption via NHEJ |
*Efficiency scores are illustrative, based on tools like Chop-Chop or CRISPick (scale 0-100). Actual scores require current tool consultation.
Table 2: Key In Vitro and Preclinical Editing Outcomes (Illustrative)
| Target | Cell Model | Delivery Method | Editing Efficiency (%) | Key Functional Outcome | Citation Context |
|---|---|---|---|---|---|
| HBB (Corrective) | CD34+ HSPCs | Electroporation (RNP) | 20-40% HDR | HbA production restored | Frangoul et al. (2021) NEJM |
| BCL11A Enhancer | CD34+ HSPCs | Electroporation (RNP) | ~80% INDELs | HbF induction to >20% | Wu et al. (2019) Nature Medicine |
Detailed Experimental Protocol: gRNA Validation
Protocol:In VitroCutting Assay (Surveyor/T7E1)
Objective: To validate gRNA nuclease activity prior to cellular experiments. Materials: Synthetic gRNA, purified SpCas9 protein, target PCR amplicon, T7 Endonuclease I. Method:
- Assembly: Combine 200 ng of target DNA amplicon, 100 nM SpCas9 protein, and 120 nM synthetic gRNA in NEBuffer 3.1. Incubate at 37°C for 1 hour.
- Purification: Clean up reaction using a PCR purification kit.
- Heteroduplex Formation: Denature/reanneal purified DNA: 95°C for 10 min, ramp down to 85°C at -2°C/sec, then to 25°C at -0.1°C/sec.
- Digestion: Add 1μL T7E1 enzyme to 10μL of reannealed DNA. Incubate at 37°C for 1 hour.
- Analysis: Run products on a 2% agarose gel. Cleavage efficiency (%) = (1 - sqrt(1 - (b+c)/(a+b+c))) * 100, where a=uncut band, b and c=cut bands.
Protocol: Delivery and Analysis in Hematopoietic Stem/Progenitor Cells (HSPCs)
Objective: Assess editing and functional outcomes in therapeutically relevant cells. Materials: Mobilized peripheral blood CD34+ cells, SpCas9 protein, synthetic gRNA, electroporator, erythroid differentiation media. Method:
- RNP Complex Formation: Pre-complex 60μM synthetic gRNA with 40μM SpCas9 protein in Cas9 buffer at room temp for 10 min.
- Electroporation: Mix 1e5 CD34+ cells with RNP complex in electroporation cuvette. Electroporate using optimized program (e.g., Lonza 4D-Nucleofector, pulse code EO-115).
- Culture & Differentiation: Recover cells in cytokine-supplemented media for 2 days, then transfer to erythroid differentiation medium for 14-21 days.
- Analysis:
- INDEL Efficiency: At day 3-4, extract genomic DNA. PCR amplify target site and perform next-generation sequencing (NGS) or TIDE analysis.
- Functional Readout: At terminal differentiation (day 18+), perform HPLC or mass spectrometry to quantify HbF (%) and HbS/HbA ratios.
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for gRNA Testing in SCD Editing
| Item | Function in Experiment | Example/Supplier |
|---|---|---|
| Synthetic sgRNA (chemically modified) | Directs Cas9 to specific genomic locus; chemical modifications enhance stability. | Synthego, IDT (Alt-R) |
| Recombinant SpCas9 Nuclease | Engineered protein that creates DSB at gRNA-specified site. | Aldevron, IDT (Alt-R S.p.) |
| CD34+ Hematopoietic Stem Cells | Primary human cell model for evaluating therapeutic editing. | AllCells, StemCell Technologies |
| Electroporation System | For efficient delivery of RNP complexes into sensitive HSPCs. | Lonza 4D-Nucleofector, Neon (Thermo Fisher) |
| NGS-based INDEL Analysis Kit | Quantitative, high-throughput measurement of editing efficiency and profile. | Illumina MiSeq, amplicon-EZ service (Genewiz) |
| Hemoglobin Analysis Kit | Functional validation via quantification of HbF, HbA, and HbS proteins. | HPLC (Bio-Rad Variant II), MSD assay |
Visualizations
Title: HBB Correction via CRISPR-Cas9 HDR Pathway
Title: BCL11A Enhancer Editing Workflow for HbF Induction
This whitepaper details a critical delivery methodology within the broader thesis of applying CRISPR-Cas9 gene editing to cure Sickle Cell Disease (SCD). The therapeutic goal is to correct the causative point mutation in the HBB gene or induce fetal hemoglobin (HBG) expression. Precise ex vivo editing of patient-derived Hematopoietic Stem and Progenitor Cells (HSPCs) is paramount, as these cells must be reinfused to reconstitute the entire hematopoietic system with genetically corrected cells. Among delivery methods, electroporation of pre-assembled Ribonucleoprotein (RNP) complexes—comprising Cas9 protein and guide RNA (sgRNA)—offers significant advantages for HSPC editing: transient exposure minimizes off-target edits, high efficiency is achievable, and it avoids genomic integration associated with viral vectors.
Table 1: Comparative Performance of Electroporation Parameters for CD34+ HSPC RNP Delivery
| Parameter / Condition | Editing Efficiency (% INDELs) | Cell Viability (Day 3 Post-Electroporation) | Erythroid Differentiation & Fetal Hemoglobin (HbF) Induction | Key Citation (Example) |
|---|---|---|---|---|
| Electroporation System: Lonza 4D-Nucleofector | ||||
| └ Program EO-100, P3 Buffer | 70-85% | 40-55% | High enucleation, >30% HbF+ cells | DeWitt et al., Sci Transl Med, 2016 |
| └ Program DZ-100, P3 Buffer | 60-75% | 50-65% | Comparable erythroid output | |
| Electroporation System: Thermo Fisher Neon | ||||
| └ 1400V, 10ms, 3 pulses, Buffer T | 65-80% | 35-50% | Robust engraftment potential in NSG mice | |
| RNP Concentration | ||||
| └ 60 µM Cas9, 180 µM sgRNA | ~80% | ~45% | Optimal for BCL11A targeting | Wu et al., Nature, 2019 |
| └ 30 µM Cas9, 90 µM sgRNA | ~65% | ~60% | Good balance of efficiency/viability | |
| HSPC Source & Pre-stimulation | ||||
| └ Mobilized Peripheral Blood (mPB), 48h | 75-90% | 50-60% | Highest editing in primitive subsets | |
| └ Cord Blood (CB), 24h | 70-85% | 55-65% | High viability, slightly lower efficiency | |
| └ Bone Marrow (BM), 48h | 65-80% | 45-55% | Variable donor-to-donor |
Table 2: Key Functional Outcomes Post-Editing in SCD Models
| Experimental Outcome | Measurement Method | Typical Result Post-BCL11A Erythroid Enhancer Editing | Implication for SCD Therapy |
|---|---|---|---|
| In Vitro Erythroid Differentiation | HbF% via HPLC/FACS | 25-50% HbF of total hemoglobin | Demonstrates phenotypic correction |
| Clonogenic Potential | CFU (Colony-Forming Unit) Assay | 60-80% of mock-electroporated control | Indicates maintained progenitor function |
| Long-Term Engraftment | Human CD45+ chimerism in NSG mice (16 wks) | Comparable to unedited transplanted cells | Confirms stem cell integrity post-editing |
| Genomic Safety | GUIDE-seq / rhAmpSeq for off-targets | 0-2 predicted off-target sites with INDELs <0.5% | Supports favorable safety profile of RNP delivery |
Detailed Experimental Protocols
Protocol: CD34+ HSPC Culture and Pre-stimulation
- Isolation: Isolate CD34+ cells from mobilized peripheral blood, cord blood, or bone marrow using clinical-grade magnetic-activated cell sorting (MACS) columns. Maintain cells in cryopreservation media in liquid nitrogen until use.
- Thawing: Rapidly thaw cryovials in a 37°C water bath. Transfer cells dropwise to 10mL of pre-warmed thawing medium (e.g., IMDM + 20% FBS + 100U/mL DNase I). Centrifuge at 300g for 10 minutes.
- Pre-stimulation: Resuspend cells in pre-stimulation medium: StemSpan SFEM II supplemented with 100ng/mL human SCF, 100ng/mL human TPO, 100ng/mL human FLT3-Ligand, and 50ng/mL human IL-6. Culture at a density of 0.5-1x10^6 cells/mL in a 37°C, 5% CO2 incubator for 24-48 hours.
Protocol: RNP Complex Assembly and Electroporation (4D-Nucleofector)
- RNP Formation:
- Dilute chemically modified, synthetic sgRNA (targeting, e.g., BCL11A erythroid enhancer or HBB sickle allele) in nuclease-free duplex buffer.
- Combine 60µM of high-purity, recombinant Cas9 protein (e.g., Alt-R S.p. HiFi Cas9) with a 3:1 molar ratio of sgRNA (180µM) in a sterile microcentrifuge tube.
- Incubate at room temperature for 10-20 minutes to allow RNP complex formation.
- Cell Preparation: After pre-stimulation, collect HSPCs, count, and assess viability. Centrifuge and resuspend in P3 Primary Cell Solution (Lonza) at a concentration of 1-2 x 10^6 cells per 20µL (for a 20µL cuvette).
- Electroporation:
- Mix 20µL of cell suspension with 2-5µL of prepared RNP complex. Do not vortex.
- Transfer the entire mixture to a 20µL Nucleocuvette strip.
- Insert the strip into the 4D-Nucleofector X unit and run the designated program (e.g., EO-100 for high efficiency, DZ-100 for higher viability).
- Immediately after electroporation, add 80µL of pre-warmed, antibiotic-free culture medium to the cuvette.
- Recovery: Gently transfer the cells (~100µL) to a well containing 1-2mL of pre-warmed, cytokine-supplemented recovery medium. Incubate at 37°C, 5% CO2 for 15-30 minutes before transferring to long-term culture or functional assays.
Protocol: Assessment of Editing Efficiency (T7 Endonuclease I Assay)
- Genomic DNA Extraction: Harvest cells 48-72 hours post-electroporation. Extract gDNA using a commercial kit (e.g., QuickExtract DNA Solution).
- PCR Amplification: Design primers flanking the target site (~500bp amplicon). Perform PCR with a high-fidelity polymerase.
- Heteroduplex Formation: Purify PCR product. Denature and reanneal: 95°C for 10 min, ramp down to 85°C at -2°C/sec, then to 25°C at -0.1°C/sec.
- Digestion: Incubate 200ng of reannealed DNA with 0.5µL T7 Endonuclease I (NEB) in 1X NEBuffer 2 at 37°C for 30 minutes.
- Analysis: Run digest on a 2% agarose gel. Calculate INDEL frequency using band intensity: % INDEL = 100 x [1 - sqrt(1 - (b+c)/(a+b+c))], where a is the undigested band, and b & c are cleavage products.
Visualizations
Workflow for HSPC Editing via RNP Electroporation
Logical Framework: RNP Delivery within SCD Thesis
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for RNP Electroporation of CD34+ HSPCs
| Category | Item/Reagent | Function & Rationale |
|---|---|---|
| Source Cells | G-CSF Mobilized Peripheral Blood CD34+ Cells | Primary human target cells with high engraftment potential; the clinically relevant source. |
| Cell Culture | StemSpan SFEM II (StemCell Tech) | Serum-free, cytokine-free basal medium optimized for HSPC expansion without differentiation. |
| Recombinant Human Cytokines (SCF, TPO, FLT3-L, IL-6) | Essential pre-stimulation cocktail to prime HSPCs for editing and improve survival post-electroporation. | |
| Editing Components | Alt-R S.p. HiFi Cas9 Nuclease V3 (IDT) | High-purity, recombinant Cas9 protein with reduced off-target activity compared to wild-type. |
| Alt-R CRISPR-Cas9 sgRNA (IDT) / Synthego sgRNA | Chemically modified synthetic sgRNA with enhanced stability and RNP formation efficiency. | |
| Delivery Hardware | 4D-Nucleofector X Unit with 20µL Cuvettes (Lonza) | Gold-standard electroporation system for primary cells; provides optimized, pre-set programs. |
| P3 Primary Cell 4D-Nucleofector Solution (Lonza) | Low-conductivity, xenofree buffer formulated for HSPCs, maximizing viability and delivery efficiency. | |
| Analysis Reagents | QuickExtract DNA Solution (Lucigen) | Rapid, single-tube gDNA extraction for genotyping assays from small cell numbers. |
| T7 Endonuclease I (NEB) | Enzyme for mismatch cleavage assay to quickly quantify INDEL efficiency at target locus. | |
| Next-Generation Sequencing Kit (Illumina) | For comprehensive, quantitative assessment of on-target editing and off-target analysis. | |
| Functional Assays | MethoCult H4434 Enriched (StemCell Tech) | Methylcellulose-based medium for CFU assays to quantify progenitor function post-editing. |
| Erythroid Differentiation Media (SCF, EPO, IL-3, etc.) | Cytokine cocktail to drive edited HSPCs toward erythroid lineage for HbF analysis via FACS/HPLC. |
Within the paradigm of CRISPR-Cas9-based therapeutic development for sickle cell disease (SCD), strategies can be broadly categorized into two approaches: corrective editing of the HBB gene and genetic reactivation of fetal hemoglobin (HbF). The latter leverages the natural developmental silencing of γ-globin (HBG) genes, which constitute HbF. This whitepaper focuses on a pivotal strategy within this category: the functional knockout of a key regulatory element—the +58 BCL11A enhancer—to disrupt the expression of the transcriptional repressor BCL11A, thereby de-repressing HBG genes and inducing HbF. This approach exemplifies the application of CRISPR-Cas9 for non-coding, cis-regulatory element editing to achieve a potent therapeutic phenotype.
Mechanistic Rationale and Pathway
BCL11A is a master transcription factor essential for the developmental switch from fetal to adult hemoglobin. It functions by repressing HBG expression. The gene encoding BCL11A is regulated by a set of enhancer elements, with the erythroid-specific +58 kb enhancer (also referred to as the GATA1 motif) being critical for its expression in erythroid cells. Disruption of this enhancer via CRISPR-Cas9-mediated knockout (e.g., through small deletions or insertions) selectively reduces BCL11A expression in the erythroid lineage, lifting the repression on the HBG genes.
Diagram: BCL11A Enhancer Knockout Pathway to HbF Induction
Table 1: Quantitative Outcomes from Preclinical & Clinical BCL11A Enhancer Editing
| Study Model | Editing Efficiency (Indels) | BCL11A Reduction | HbF Induction (F-cells or %HbF) | Key Readout | Reference (Example) |
|---|---|---|---|---|---|
| Human CD34+ HSPCs (in vitro) | 80-90% | ~70-80% protein knockdown | >40% F-cells; HbF ~25-30% of total Hb | Terminal erythroid differentiation | Wu et al., 2019 |
| SCD Mouse Model (xenograft) | ~80% in engrafted human cells | Significant knockdown in erythroid cells | >30% HbF; Reduced sickling | In vivo pathology correction | Frangoul et al., 2020 (preclinical) |
| Clinical Trial (CLIMB SCD-121) | High allele editing in bone marrow | Not directly reported | ~40% HbF at 18 months; >95% F-cells | Resolution of vaso-occlusive events | Frangoul et al., NEJM 2020 |
Table 2: Comparison of Key SCD Gene Editing Strategies Targeting HbF
| Parameter | BCL11A Enhancer Knockout | BCL11A Erythroid Exon 2 KO | HBG Promoter Editing |
|---|---|---|---|
| Target | Non-coding enhancer (+58 kb) | Coding exon (Erythroid-specific) | Non-coding HBG promoter |
| Primary Effect | Reduces BCL11A transcription | Disrupts BCL11A protein in erythroid cells | Disrupts repressor binding sites |
| HbF Induction Level | Very High (>>20%) | Very High (>>20%) | High (Variable) |
| Specificity | Erythroid-specific (enhancer-dependent) | Erythroid-specific (exon choice) | Universal |
| Clinical Stage | Approved (exa-cel) | Phase 3 Trials | Phase 1/2 Trials |
Detailed Experimental Protocol: BCL11A Enhancer Editing in Human CD34+ HSPCs
This protocol outlines the key steps for ex vivo gene editing of hematopoietic stem and progenitor cells (HSPCs) for preclinical analysis.
Materials:
- Mobilized peripheral blood or cord blood-derived human CD34+ HSPCs.
- Cas9 Protein: High-purity, recombinant S. pyogenes Cas9 nuclease.
- sgRNA: Synthetic, chemically modified sgRNA targeting the GATA1 motif within the +58 BCL11A enhancer (e.g., sequence: 5'-GCCCACAGTGGCACCATGAG-3').
- Electroporation Buffer: Optimized, such as P3 Primary Cell Solution.
- Electroporator: 4D-Nucleofector (Lonza) or Neon (Thermo Fisher).
- Erythroid Differentiation Media: Multi-stage media containing SCF, EPO, IL-3, dexamethasone, etc.
- QC Reagents: T7 Endonuclease I or next-generation sequencing (NGS) kits for indel analysis; flow cytometry antibodies for BCL11A and HbF (F-cells).
Procedure:
Design and Preparation of RNP:
- Resuspend the sgRNA in nuclease-free buffer.
- Pre-complex the Cas9 protein and sgRNA at a molar ratio of 1:2.5 (e.g., 100 pmol Cas9: 250 pmol sgRNA) in a total volume of ~10 µL. Incubate at room temperature for 10-20 minutes to form the ribonucleoprotein (RNP) complex.
CD34+ HSPC Preparation and Electroporation:
- Thaw and pre-stimulate CD34+ cells in serum-free expansion medium containing cytokines (SCF, TPO, FLT3-L) for 24-48 hours.
- Harvest 1x10^5 to 1x10^6 cells, wash, and resuspend in the appropriate volume of electroporation buffer.
- Mix the cell suspension with the pre-formed RNP complex. Transfer the mixture to a certified cuvette or strip.
- Electroporate using a validated program (e.g., [DS-138] on a 4D-Nucleofector).
- Immediately transfer cells to pre-warmed recovery medium.
Post-Electroporation Culture and Analysis:
- Day 0-2: Culture cells in expansion medium. Assess viability 24h post-electroporation.
- Day 3: Harvest a sample for genomic DNA extraction. Assess editing efficiency at the target locus via T7E1 assay or, preferably, NGS.
- Day 4-16: Initiate erythroid differentiation. Transfer cells to stage-specific differentiation media, maintaining high cell density.
- Day 14-18: Harvest terminally differentiated erythroid cells for functional analysis.
Functional Readouts:
- Flow Cytometry: Stain cells for CD235a (glycophorin A), intracellular BCL11A, and HbF (using anti-γ-globin antibody) to quantify the percentage of F-cells and BCL11A knockdown.
- HPLC: Perform hemoglobin electrophoresis or HPLC on cell lysates to quantify the percentage of HbF relative to total hemoglobin.
- In Vivo Assessment (Optional): Transplant edited CD34+ cells into immunodeficient mice (e.g., NSG) and analyze human erythroid chimerism, HbF induction, and sickling in circulating cells after 16-20 weeks.
Diagram: Experimental Workflow for HSPC Editing and Validation
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagent Solutions for BCL11A Enhancer Editing Research
| Reagent / Material | Function / Role | Example Vendor / Catalog |
|---|---|---|
| Recombinant S. pyogenes Cas9 Nuclease | The endonuclease that creates a double-strand break at the target DNA site directed by the sgRNA. | Integrated DNA Technologies (IDT), Thermo Fisher Scientific |
| Chemically Modified sgRNA (targeting +58 enhancer) | Guides the Cas9 protein to the specific GATA1 motif within the BCL11A erythroid enhancer. Chemical modifications enhance stability. | Synthego, IDT, Trilink BioTechnologies |
| Human CD34+ Cell Isolation Kit | For the positive selection of hematopoietic stem and progenitor cells from source material (mPB, CB). | Miltenyi Biotec (CD34 MicroBead Kit) |
| 4D-Nucleofector X Kit and Unit | System for high-efficiency, low-toxicity delivery of RNP into hard-to-transfect primary CD34+ cells. | Lonza |
| StemSpan SFEM II with Cytokines | Serum-free, optimized medium for expansion and maintenance of undifferentiated CD34+ HSPCs pre- and post-editing. | StemCell Technologies |
| Erythroid Differentiation Media Kit | Multi-stage, cytokine-driven system to drive edited HSPCs through terminal erythroid maturation. | STEMCELL Technologies (HemaTox) |
| Anti-Human Fetal Hemoglobin Antibody (FITC) | Flow cytometry antibody for detecting γ-globin protein and quantifying the percentage of F-cells. | BD Biosciences, Invitrogen |
| BCL11A-XL Antibody (for flow/IF) | For detecting and quantifying the reduction in BCL11A protein levels in differentiated erythroid cells. | Cell Signaling Technology |
| T7 Endonuclease I | Enzyme for initial, rapid assessment of indel formation via mismatch cleavage assay. | NEB |
| NGS Library Prep Kit for Amplicon Sequencing | For precise, quantitative measurement of editing efficiency and indel spectrum at the target locus. | Illumina (MiSeq), IDT (xGen) |
Within the broader thesis on utilizing the CRISPR-Cas9 mechanism for sickle cell disease (SCD) gene editing research, correcting the causative point mutation in the HBB gene (A>T, Glu6Val) is paramount. Traditional CRISPR-Cas9 homology-directed repair (HDR) is inefficient in non-dividing cells like hematopoietic stem cells and can induce uncontrolled indels. This whitepaper details two precise, next-generation strategies—direct base editing and prime editing—that directly rectify point mutations without requiring double-strand DNA breaks (DSBs) or donor DNA templates.
Direct Base Editing: Principles and Application to SCD
Base editors (BEs) are fusion proteins comprising a catalytically impaired Cas9 (nickase or dead) tethered to a nucleobase deaminase enzyme. They facilitate the direct, irreversible conversion of one target DNA base pair into another without creating a DSB.
- Cytosine Base Editors (CBEs): Convert C•G to T•A. The SCD mutation (A>T) is not directly reversible with a CBE. However, a compensatory strategy targets a different T in the codon (GAG to GTG) to recreate the sickling mutation in healthy donor cells for research, or potentially install a benign SNP that upregulates fetal hemoglobin.
- Adenine Base Editors (ABEs): Convert A•T to G•C. This is therapeutically relevant for SCD, as the reverse mutation (T>A) correction requires an A•T to G•C conversion on the opposite strand. An ABE can directly correct the pathogenic Glu6Val mutation back to the wild-type Glu codon.
Table 1: Comparison of Base Editor Systems for HBB Point Mutation Correction
| Editor Type | Core Components | Target SCD Mutation (HBB, codon 6) | Conversion | Editing Window (Protospacer Position) | Primary Outcome | Key Limitation |
|---|---|---|---|---|---|---|
| ABE8e | dCas9 or nCas9 + evolved TadA-8e deaminase | A>T (GAG->GTG) | A•T → G•C | ~ positions 4-8 (counting PAM as 21-23) | Correction to wild-type GAG (Glu) | Off-target RNA editing; bystander editing possible. |
| BE4max | nCas9 + rAPOBEC1 deaminase | Not directly applicable. Can install research model mutation. | C•G → T•A | ~ positions 4-8 | Can create sickling GTG (Val) codon in wild-type sequence. | Cannot correct the primary SCD mutation. |
| Dual BE | Sequential ABE + CBE delivery | Comprehensive correction or modulation. | A•T → G•C & C•G → T•A | Dependent on individual BE | Potential for multiplexed correction of mutation and associated SNPs. | Complex delivery; increased risk of indels. |
Experimental Protocol: In Vitro Correction of SCD Mutation in CD34+ HSPCs using ABE8e
- Design: Create a sgRNA with a protospacer sequence positioning the target adenine within the editing window (typically positions 4-8, counting from the distal end of the non-target strand) of an NGG PAM.
- Delivery: Electroporate CD34+ hematopoietic stem and progenitor cells (HSPCs) with ABE8e mRNA and synthetic sgRNA.
- Culture: Maintain cells in cytokine-supplemented serum-free media supporting stemness.
- Analysis:
- Efficiency: 3-5 days post-editing, harvest genomic DNA. Perform PCR amplification of the HBB target locus and sequence via next-generation sequencing (NGS) to quantify A•T to G•C conversion rates and indel frequencies.
- Functional Assessment: Differentiate edited HSPCs into erythroid lineage and perform HPLC to measure hemoglobin S (HbS) reduction and western blot for HBB protein correction.
- Safety: Perform whole-genome sequencing (WGS) or targeted off-target analysis (e.g., GUIDE-seq) to assess unintended edits.
Title: ABE Workflow for SCD Correction
Prime Editing: A Search-and-Replace Tool for SCD
Prime editors (PEs) are versatile fusion proteins consisting of a Cas9 nickase (H840A) reverse transcriptase (RT) enzyme. A prime editing guide RNA (pegRNA) directs the system to the target site and also encodes the desired edit. PEs can install all 12 possible base-to-base conversions, small insertions, and deletions, without DSBs.
Application to SCD: A PE can be designed to precisely correct the exact A>T transversion in the HBB gene. The pegRNA would specify the change from "GTG" (Val) back to "GAG" (Glu).
Table 2: Prime Editing System Specifications for HBB Correction
| Component | Specification for SCD (HBB codon 6) | Function |
|---|---|---|
| PE Protein | Cas9(H840A)-RT fusion (e.g., PE2, PEmax) | Nicks target strand and reverse transcribes new DNA from pegRNA. |
| pegRNA | Contains: 1) sgRNA spacer, 2) RT template with GAG correction, 3) Primer Binding Site (PBS). | Guides PE to locus and provides template for correction. |
| nicking sgRNA | Optional (for PE3/PE3b systems). Guides a second nick on the non-edited strand to increase efficiency. | Promotes cellular repair to incorporate the edited strand. |
| Typical Efficiency in HSPCs | 10-40% correction with PEmax, depending on design and delivery. | |
| Primary Advantage | High precision and versatility; minimal indel byproducts. | |
| Key Challenge | Lower efficiency than base editors; complex pegRNA design optimization required. |
Experimental Protocol: Prime Editing in HEK293T Cells (Model for Optimization)
- pegRNA Design: Design multiple pegRNAs varying in PBS length (8-15 nt) and RT template length. Include the corrected base(s) and necessary synonymous mutations to prevent re-editing.
- Delivery: Co-transfect HEK293T cells with plasmids encoding PEmax and the pegRNA (and optional nicking sgRNA for PE3).
- Screening: Extract genomic DNA 72 hours post-transfection. Amplify the target region via PCR and analyze by Sanger sequencing or NGS to identify the most efficient pegRNA designs.
- Translation to HSPCs: Deliver the optimized PE system as RNP (PEmax protein + in vitro transcribed pegRNA) into CD34+ HSPCs via electroporation. Assess correction efficiency and purity via NGS.
Title: Prime Editing Search-and-Replace Mechanism
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Base and Prime Editing Research in SCD
| Reagent / Material | Supplier Examples | Function in Experiment |
|---|---|---|
| ABE8e or PEmax Expression Plasmid | Addgene | Source of editor DNA for RNP production or viral packaging. |
| Chemically Modified sgRNA/pegRNA | Synthego, IDT | Enhances stability and editing efficiency in primary cells; reduces immune response. |
| Recombinant ABE8e or PEmax Protein | ToolGen, Thermo Fisher | For RNP assembly and delivery, offering rapid kinetics and reduced off-target risk. |
| Human CD34+ HSPCs | Lonza, StemCell Technologies | Primary, therapeutically relevant cell model for SCD gene editing. |
| Electroporation System (Neon, 4D-Nucleofector) | Thermo Fisher, Lonza | High-efficiency delivery platform for RNP or mRNA into sensitive HSPCs. |
| HSPC Expansion Media (SFEM II) | StemCell Technologies | Serum-free media supporting HSPC maintenance during editing. |
| Next-Generation Sequencing Kit (Illumina MiSeq) | Illumina | For deep sequencing of target loci to quantify editing efficiency, purity, and byproducts. |
| Guide-seq / CHANGE-seq Kit | IDT, Custom | For unbiased, genome-wide identification of potential off-target sites. |
| Erythroid Differentiation Kit | StemCell Technologies | To differentiate edited HSPCs into erythroid cells for functional hemoglobin analysis. |
Direct base editing and prime editing represent transformative advances beyond standard CRISPR-Cas9 for point mutation correction in sickle cell disease. ABEs offer a relatively efficient, one-step correction of the pathogenic HBB variant. Prime editors provide unparalleled versatility and precision, capable of installing the exact correction with minimal genotoxic risk. The choice of strategy involves a critical trade-off between efficiency (favoring ABEs) and versatility/cleanliness (favoring PEs). Ongoing optimization of editor proteins, delivery methods, and pegRNA design is rapidly enhancing both platforms, moving them closer to clinical translation for SCD and other genetic disorders.
Manufacturing and Scalability Considerations for Clinical-Grade Cell Products
Introduction The advent of CRISPR-Cas9 gene editing has ushered in a new era of advanced therapeutic products. A seminal application is the ex vivo editing of hematopoietic stem and progenitor cells (HSPCs) for sickle cell disease (SCD), as exemplified by the approved therapies Casgevy (exagamglogene autotemcel). This success underscores a critical challenge: translating a precise laboratory edit into a robust, scalable, and reproducible process for manufacturing clinical-grade cell products. This guide details the technical considerations, from vector systems to process analytics, essential for this translation, framed within the context of CRISPR-Cas9 SCD research.
1. Core CRISPR-Cas9 Editing Workflow for SCD The SCD therapeutic strategy involves ex vivo editing of patient-derived HSPCs to induce fetal hemoglobin (HbF) via BCL11A erythroid enhancer disruption. The manufacturing workflow is a linear sequence of interdependent unit operations.
Diagram Title: Ex Vivo CRISPR-Cas9 HSPC Manufacturing Workflow
2. Key Manufacturing & Scalability Components
2.1. Reagent and Vector Systems The choice of editing components is foundational. For clinical use, sourcing Good Manufacturing Practice (GMP)-grade materials is non-negotiable.
Table 1: Comparison of CRISPR-Cas9 Delivery Modalities for HSPCs
| Delivery Method | Format | Key Advantage | Key Scalability/Manufacturing Challenge | Typical Editing Efficiency (HSPCs) |
|---|---|---|---|---|
| Electroporation of RNP | Cas9 Protein + sgRNA Complex | Rapid kinetics, reduced off-target risk, no DNA integration | GMP-grade protein production & characterization; cost at scale | 80-95% |
| Viral Vector (AAV6) | Recombinant AAV carrying sgRNA template | High delivery efficiency to HSPCs | Complex, high-cost GMP viral production; pre-existing immunity concerns | 50-80% |
| mRNA Electroporation | Cas9 mRNA + sgRNA | Transient expression, no DNA integration | Stability of mRNA; potential higher immune response | 60-85% |
2.2. Cell Processing and Culture Scalability requires moving from flask-based to closed, automated bioreactor systems.
- Pre-stimulation: Typically uses serum-free media with recombinant cytokines (SCF, TPO, FLT3L) for 1-2 days to prime HSPCs for editing.
- Post-edition Culture: Short-term (1-3 days) culture to allow for edit completion and protein turnover before cryopreservation. Extended culture risks differentiation and loss of stemness.
- Scale-up Platforms: Transition from static culture bags to closed, automated bioreactors (e.g., rocking-motion or hollow-fiber systems) is critical for lot sizes > 1e9 cells. These systems ensure consistent gas exchange, nutrient delivery, and pH control.
2.3. Critical Quality Attributes (CQAs) & Analytics A multi-parameter release specification is required to ensure product safety and potency.
Table 2: Essential Release Tests for an SCD-edited HSPC Product
| CQA Category | Specific Test | Target Specification | Rationale |
|---|---|---|---|
| Identity/Potency | INDEL Frequency at BCL11A target (NGS) | > 60% | Primary mechanism of action (HbF induction) |
| HbF Expression (HPLC/FACS) | > 20% F-cells | Functional potency correlate | |
| CD34+ Viability (Flow Cytometry) | > 70% | Ensures engraftment potential | |
| Safety | Vector Copy Number (ddPCR) | < 0.5 copies/cell (if using viral donor) | Assesses risk of insertional mutagenesis |
| Off-target Analysis (GUIDE-seq/CHANGE-seq) | No significant hits in pre-defined risk loci | Assesses genomic specificity | |
| Sterility (BacT/ALERT) | No growth | Prevents infection | |
| Purity/Vector Safety | Replication Competent AAV (RCAAV) Assay | Negative (if using AAV6) | Ensures viral safety |
| Dose | Total Viable CD34+ Cell Count | Defined per protocol | Determines therapeutic dose |
| Colony-Forming Unit (CFU) Assay | > specific threshold per kg | Functional measure of progenitor content |
3. Detailed Experimental Protocol: Electroporation of CRISPR-Cas9 RNP into HSPCs This protocol is adapted from published clinical-scale methods for SCD.
Objective: To achieve high-efficiency editing of the BCL11A erythroid enhancer in mobilized human CD34+ HSPCs using Cas9 RNP electroporation.
Materials (The Scientist's Toolkit): Table 3: Key Research Reagent Solutions for Clinical-Scale HSPC Electroporation
| Item | Function | Example (GMP-grade if available) |
|---|---|---|
| Human CD34+ HSPCs | Starting cellular raw material | Mobilized peripheral blood apheresis product, >90% purity. |
| StemSpan SFEM II | Serum-free expansion medium | Provides defined, xeno-free culture conditions. |
| Recombinant Cytokines (SCF, TPO, FLT3L) | Pre-stimulation | Primes HSPCs for efficient editing and survival. |
| Alt-R S.p. Cas9 Nuclease V3 | GMP-grade Cas9 protein | Catalyzes the DNA double-strand break at the target locus. |
| Alt-R CRISPR-Cas9 sgRNA | Target-specific guide RNA | Complexes with Cas9 to direct it to the BCL11A enhancer. |
| Electroporation Buffer | Low-conductivity solution | Maximizes cell viability and delivery efficiency during electroporation. |
| Lonza 4D-Nucleofector & P3 Kit | Electroporation device & cuvettes | Enables high-efficiency, scalable non-viral delivery. |
| DNase I | Degrades residual plasmid DNA | Critical safety step if RNP was produced using in vitro transcription. |
Methodology:
- Cell Thaw and Pre-stimulation: Thaw cryopreserved CD34+ HSPCs in a 37°C water bath. Wash in pre-warmed medium, count, and assess viability. Seed cells at 1-2e6 cells/mL in StemSpan SFEM II supplemented with 100 ng/mL each of SCF, TPO, and FLT3L. Culture for 24-48 hours at 37°C, 5% CO2.
- RNP Complex Formation: For a 100 µL reaction (sufficient for ~1e6 cells), combine 60 pmol of GMP-grade Cas9 protein with 180 pmol of target-specific sgRNA in nuclease-free duplex buffer. Incubate at room temperature for 10-20 minutes to form the RNP complex.
- Cell Preparation and Electroporation: Harvest pre-stimulated cells, wash once with PBS, and resuspend in P3 primary cell electroporation solution at a concentration of 1e7 cells/mL. Combine 10 µL of cell suspension (1e5 cells) with 10 µL of pre-formed RNP complex. Transfer to a 20 µL Nucleocuvette strip. Electroporate using the Lonza 4D-Nucleofector with the recommended program for HSPCs (e.g., EO-100 or DS-138). Include an RNP-negative control (cells + buffer only).
- Post-Electroporation Recovery: Immediately after electroporation, add 80 µL of pre-warmed, cytokine-supplemented medium to the cuvette. Gently transfer cells to a culture plate. Place in incubator for 15-30 minutes, then perform a full medium change to remove debris and electroporation buffer.
- Post-edition Culture and Harvest: Culture cells for 48-72 hours. Analyze an aliquot for editing efficiency (via T7 Endonuclease I assay or NGS) and viability. For clinical production, cells are then formulated in cryomedium (containing DMSO and human serum albumin), filled into cryobags, and controlled-rate frozen for cryopreservation.
4. Scaling and Process Control The logical relationship between scaling stages and control systems is critical.
Diagram Title: Scaling Pathway with Process Controls
Conclusion Manufacturing clinical-grade CRISPR-edited cell products for SCD is a multidisciplinary feat integrating molecular biology, cell process engineering, and rigorous quality control. The transition from research to clinic hinges on the adoption of GMP-grade reagents, closed automated systems, and a comprehensive analytical framework centered on defined CQAs. As the field progresses, innovations in all-in-one editing systems, inline process monitoring (PAT), and automated fill-finish will further enhance scalability, consistency, and access to these transformative therapies.
Navigating Challenges: Off-Target Effects, Efficiency, and Translational Hurdles
The therapeutic promise of CRISPR-Cas9 for sickle cell disease (SCD) hinges on precise editing of the HBB gene to induce fetal hemoglobin (HbF) expression, typically via BCL11A enhancer disruption or direct HBB correction. While clinical trials show efficacy, the potential for off-target editing remains a primary safety concern. Unwanted double-strand breaks (DSBs) at loci with sequence homology to the single guide RNA (sgRNA) could disrupt tumor suppressors, oncogenes, or other vital genomic regions. This guide details integrated computational and empirical frameworks essential for profiling and mitigating these risks in SCD therapeutic development.
Computational Prediction of Off-Target Sites
In silico tools predict potential off-target sites by scanning the genome for sequences with imperfect matches to the sgRNA spacer, especially in the seed region proximal to the PAM.
Key Algorithms and Tools:
- Basic Alignment Tools (BLAST, Bowtie): Identify sites with a limited number of mismatches.
- CFD (Cutting Frequency Determination) Score: Predicts off-target activity based on mismatch type and position.
- MIT CRISPR Design Tool: Incorporates CFD and MIT specificity scores.
- Cas-OFFinder: Searches for potential off-targets across genomes, allowing bulges (insertions/deletions).
Quantitative Comparison of Predictive Tools:
Table 1: Comparison of Computational Off-Target Prediction Tools
| Tool Name | Core Algorithm | Inputs | Key Outputs | Advantages | Limitations |
|---|---|---|---|---|---|
| Cas-OFFinder | Genome-wide exhaustive search | sgRNA seq, PAM, mismatch/bulge tolerance | List of genomic loci | Allows bulge searches; fast | Does not predict cleavage likelihood |
| MIT CRISPR Design | CFD & MIT Specificity Scores | sgRNA seq, reference genome | Ranked off-target list, specificity score | Validated scoring model; user-friendly | Limited to pre-defined genomes; no bulge consideration |
| CHOPCHOP | Multiple (including MIT, CFD) | Target gene or sequence | On/Off-target scores, primer design | Integrated design and validation suite | Predictive accuracy varies by algorithm chosen |
| CCTop | Empirical rules from GUIDE-seq data | sgRNA seq | Off-targets ranked by likelihood | Incorporates experimental data trends | May miss sites not represented in training data |
Empirical Detection of Off-Target Effects
Computational prediction requires empirical validation. Unbiased, genome-wide methods are now standard.
3.1. GUIDE-seq (Genome-wide, Unbiased Identification of DSBs Enabled by Sequencing)
- Principle: Captures in situ DSBs via integration of a double-stranded oligodeoxynucleotide (dsODN) tag.
- Detailed Protocol:
- Transfection: Co-deliver CRISPR-Cas9 RNP (ribonucleoprotein) and the GUIDE-seq dsODN tag into target cells (e.g., HUDEP-2 or CD34+ HSPCs for SCD models).
- Tag Integration: Cellular NHEJ machinery integrates the dsODN into DSB sites.
- Genomic DNA Extraction & Shearing: Harvest cells 48-72h post-transfection. Extract and fragment DNA.
- Enrichment & Library Prep: Use PCR to enrich tagged fragments and prepare sequencing libraries.
- Sequencing & Analysis: Perform high-throughput sequencing. Map reads to identify genomic junctions between the dsODN tag and the genome, denoting DSB loci.
3.2. CIRCLE-seq (Circularization for In Vitro Reporting of Cleavage Effects by Sequencing)
- Principle: An in vitro, highly sensitive method using circularized genomic DNA as a substrate for Cas9 cleavage.
- Detailed Protocol:
- Genomic DNA Isolation & Circularization: Extract genomic DNA from target cell type. Fragment, end-repair, and ligate into circular molecules.
- In Vitro Cleavage: Incubate circularized DNA with pre-complexed Cas9-sgRNA RNP. Linearized circles indicate cleavage events.
- Adapter Ligation & Linear Molecule Capture: Ligate adapters to broken ends. Use exonuclease to degrade remaining circular DNA, enriching cleaved fragments.
- Library Preparation & Sequencing: Amplify and sequence adapter-ligated fragments.
- Bioinformatics Analysis: Map sequence reads to the reference genome. Peak calling identifies cleavage sites with single-nucleotide resolution.
Quantitative Comparison of Empirical Methods:
Table 2: Comparison of Empirical Off-Target Detection Methods
| Method | Sensitivity | Throughput | Context | Key Advantage | Key Limitation |
|---|---|---|---|---|---|
| GUIDE-seq | High (detects sites at ~0.1% frequency) | Genome-wide | In cellulo (native chromatin) | Captures cellular context & repair | Requires dsODN transfection; background noise possible |
| CIRCLE-seq | Very High (detects sites at <0.01% frequency) | Genome-wide | In vitro (protein-free DNA) | Extremely sensitive; low background | May miss chromatin-influenced sites |
| Digenome-seq | High | Genome-wide | In vitro (cell-free genomic DNA) | Uses native genomic DNA sequence | High input DNA requirement; computationally intensive |
| SITE-seq | High | Genome-wide | In vitro (biochemical) | Controlled cleavage conditions; precise mapping | Does not account for cellular repair |
Strategies for Minimizing Off-Target Editing
- sgRNA Design Optimization: Select sgRNAs with high on-target and minimal predicted off-target scores using tools in Table 1. Prioritize unique genomic sequences.
- High-Fidelity Cas9 Variants: Use engineered Cas9 nucleases (e.g., SpCas9-HF1, eSpCas9(1.1), HiFi Cas9) with reduced non-specific DNA binding.
- RNP Delivery: Deliver pre-complexed Cas9 protein and sgRNA as RNP. This shortens exposure time, reducing off-target effects compared to plasmid DNA delivery.
- Dosage Control: Use the minimum effective concentration of CRISPR components.
- Modified sgRNA Scaffolds: Incorporate chemical modifications (e.g., 2'-O-methyl-3'-phosphorothioate) to enhance stability and specificity.
- Prime Editing: For SCD point correction strategies, use prime editing which nicks DNA and does not create full DSBs, drastically reducing off-target profiles.
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Off-Target Assessment in SCD Gene Editing Research
| Reagent / Material | Function in Off-Target Analysis | Example Product/Catalog |
|---|---|---|
| High-Fidelity Cas9 Nuclease | Engineered protein for reduced off-target cleavage while maintaining on-target activity. | HiFi Cas9 Protein, Alt-R S.p. HiFi Cas9 Nuclease V3 |
| Chemically Modified sgRNA | Enhanced stability and specificity guides; often synthetic, tracrRNA modifications. | Alt-R CRISPR-Cas9 sgRNA (2'-O-methyl, 3' phosphorothioate) |
| GUIDE-seq dsODN Tag | Double-stranded tag for integration into DSBs during NHEJ for genome-wide break mapping. | Truseq GUIDE-seq Oligo Kit, Custom dsODN |
| CIRCLE-seq Adapters & Enzymes | Specialized adapters and exonucleases for circularization and enrichment of cleaved fragments. | CIRCLE-seq Kit, NEBNext Ultra II DNA Library Prep Kit |
| Next-Generation Sequencing Kit | For preparing libraries from enriched fragments for high-throughput sequencing. | Illumina DNA Prep, Swift Accel-NGS 2S Plus |
| Positive Control sgRNA/Oligos | Validated sgRNA and oligos for known off-target sites to serve as assay controls. | Synthetic oligonucleotides for predicted off-target loci |
| Genomic DNA Isolation Kit (High MW) | To obtain high-quality, high-molecular-weight DNA for CIRCLE-seq or Digenome-seq. | Qiagen Genomic-tip, MagAttract HMW DNA Kit |
| Human CD34+ HSPCs or HUDEP-2 Cells | Clinically relevant cell models for SCD gene editing and off-target profiling. | Mobilized Peripheral Blood CD34+ Cells, HUDEP-2 cell line |
Visualized Workflows and Relationships
Title: Integrated Off-Target Assessment and Mitigation Workflow
Title: CIRCLE-seq Experimental Workflow
Title: GUIDE-seq Experimental Workflow
Optimizing Editing Efficiency and HDR Rates in Quiescent Hematopoietic Stem Cells
The application of CRISPR-Cas9 for curing sickle cell disease (SCD) represents a paradigm shift in genetic medicine. The primary therapeutic goal is to correct the causative Glu6Val point mutation in the β-globin (HBB) gene or to induce fetal hemoglobin (HbF) via BCL11A enhancer editing. While clinical successes have been achieved, a key biological bottleneck remains: the quiescent nature of the most primitive human hematopoietic stem cells (HSCs). These long-term repopulating HSCs predominantly reside in the G0 phase of the cell cycle, a state that favors non-homologous end joining (NHEJ) over the precise homology-directed repair (HDR) pathway. This technical guide details strategies to overcome this barrier, optimizing both total editing efficiency and HDR rates in quiescent HSCs, thereby enhancing the therapeutic potential and safety profile of SCD gene therapies.
Table 1: Comparison of Strategies to Enhance HDR in Quiescent HSCs
| Strategy | Target/Mechanism | Reported HDR Increase | Key Trade-off/Consideration |
|---|---|---|---|
| Small Molecule Inhibition (e.g., SCR7) | DNA Ligase IV (NHEJ) | 1.8- to 3.2-fold | Potential for increased genomic instability from prolonged NHEJ inhibition. |
| RS-1 (RAD51 stimulant) | Enhances RAD51 nucleofilament stability | 2- to 4-fold | Can be cytotoxic at higher concentrations; variable efficacy across cell types. |
| Cell Cycle Synchronization (e.g., Nocodazole) | Arrest in G2/M phase (higher HDR) | ~4-fold increase in HDR/NHEJ ratio | May compromise stemness and long-term engraftment potential. |
| Modified gRNA Designs (Alt-R HDR) | Chemically modified gRNAs | ~1.5-fold HDR efficiency | Modest improvement, best used in combination with other methods. |
| AAV6 HDR Donor Delivery | High-efficiency transduction & template delivery | Can achieve >40% HDR in mobilized CD34+ cells | Immunogenicity concerns; size limitations for donor template. |
| Inhibitor of 53BP1 (e.g., i53) | Blocks 53BP1 recruitment to DSBs | Up to 5.9-fold HDR increase in primary T cells | Shifts repair balance profoundly; long-term safety data needed. |
Table 2: Core Experimental Outcomes in Edited Quiescent HSCs
| Metric | Baseline (Untreated Quiescent HSCs) | Optimized Protocol (e.g., RS-1 + AAV6) | Measurement Method |
|---|---|---|---|
| Total Editing (% Indels) | 20-40% | 60-80% | NGS of target locus; T7E1 assay. |
| HDR Efficiency (%) | 5-15% | 25-45% | NGS detecting precise template incorporation. |
| Cell Viability Post-Electroporation | 40-60% | 65-80% | Flow cytometry (Annexin V/PI). |
| Long-term Engraftment in NSG Mice | ~10-20% human CD45+ | Maintained or slightly reduced (~15-25%) | Multilineage analysis 16+ weeks post-transplant. |
Detailed Experimental Protocols
Protocol 1: Combined RS-1 and AAV6 Donor Delivery for HDR Enhancement in Quiescent CD34+ HSCs
- Materials: Mobilized peripheral blood CD34+ cells, Cas9 RNP (Alt-R S.p. Cas9 + synthetic gRNA), Alt-R HDR donor template (ssODN) or AAV6-HDR donor vector, RS-1 (Sigma), Serum-Free Expansion Medium II (SFEM II), Cytokines (SCF, TPO, FLT3L).
- Procedure:
- Pre-stimulation: Culture CD34+ cells in SFEM II with cytokines (100 ng/mL each) for 24-48 hours.
- RNP Complex Formation: Complex 60 µg of S.p. Cas9 protein with 200 pmol of synthetic gRNA (targeting HBB or BCL11A enhancer) at room temperature for 10 minutes.
- Electroporation: Use the Lonza 4D-Nucleofector (program DZ-100). Resuspend 1e5 cells in 20 µL P3 buffer with RNP complex. Transfer to cuvette and electroporate.
- HDR Enhancement: Immediately post-electroporation, add cells to pre-warmed medium containing 40 µM RS-1. For AAV6 delivery, transduce cells at an MOI of 1e5 vg/cell concurrently.
- Recovery & Analysis: Culture cells for 3-7 days. Analyze editing efficiency by flow cytometry (for fluorescent reporters) or harvest genomic DNA for NGS at day 7. For transplantation assays, inject cells into sub-lethally irradiated NSG mice within 24 hours of editing.
Protocol 2: Cell Cycle Profiling and Sorting of Quiescent HSCs Pre-/Post-Editing
- Materials: Pyronin Y, Hoechst 33342, DAPI, FACS sorter.
- Procedure:
- Staining: At 24h post-editing, harvest cells. Stain with 1 µg/mL Hoechst 33342 (DNA content) and 0.5 µg/mL Pyronin Y (RNA content) for 45 min at 37°C.
- Gating Strategy: Use FACS to identify G0 (Hoechst[low]/Pyronin[low]), G1 (Hoechst[low]/Pyronin[high]), and S/G2/M (Hoechst[high]/Pyronin[high]) populations.
- Analysis: Sort populations and extract genomic DNA. Use NGS to compare HDR and NHEJ outcomes specifically in the purified G0 population versus cycling cells.
Signaling Pathways and Workflow Visualizations
Title: HDR Bottleneck and Intervention Points in Quiescent HSCs
Title: Optimized HSC Editing and Validation Workflow
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Editing Quiescent HSCs
| Reagent/Category | Example Product (Supplier) | Function in Experiment |
|---|---|---|
| CRISPR-Cas9 Delivery | Alt-R S.p. Cas9 Nuclease V3 (IDT) | High-purity, pre-complexed Cas9 protein for RNP formation, reduces off-target effects vs. plasmid. |
| Synthetic gRNA | Alt-R CRISPR-Cas9 sgRNA (IDT) | Chemically modified for enhanced stability and reduced immunogenicity. |
| HDR Donor Template | AAV6 HBB Donor Vector (VectorBuilder) | High-efficiency delivery of long homology arm templates for precise HDR. |
| HDR Enhancer (Small Molecule) | RS-1 (Sigma-Aldrich, 554189) | RAD51 stimulator; promotes strand invasion during homologous recombination. |
| NHEJ Inhibitor | SCR7 (Sigma-Aldrich, SML1546) | DNA Ligase IV inhibitor; temporarily biases repair towards HDR. |
| Cell Cycle Modulator | Nocodazole (Sigma-Aldrich, M1404) | Microtubule polymerizer; synchronizes cells at G2/M phase where HDR is more active. |
| Stem Cell Maintenance Media | StemSpan SFEM II (StemCell Tech) | Serum-free, cytokine-supportive medium for maintaining HSC potency ex vivo. |
| Electroporation System | 4D-Nucleofector X Unit (Lonza) | High-efficiency delivery of RNP complexes into sensitive primary HSCs. |
| Cell Viability Dye | Zombie NIR Fixable Viability Kit (BioLegend) | Accurate flow cytometry-based discrimination of live/dead cells post-editing. |
| NGS for Editing Analysis | Illumina MiSeq Amplicon-EZ (Genewiz) | Deep sequencing to quantify precise HDR rates and indel patterns at target locus. |
The therapeutic application of CRISPR-Cas9 for sickle cell disease (SCD), exemplified by the recently approved ex vivo therapy exagamglogene autotemcel, involves the targeted disruption of the BCL11A erythroid enhancer to induce fetal hemoglobin. While clinically transformative, the underlying gene-editing process can induce cellular stress responses that pose potential risks. This guide details the technical underpinnings of three major toxicity concerns—p53-mediated DNA damage response, on-target and off-target chromosomal aberrations, and pre-existing or induced adaptive immune responses to the Cas9 nuclease—within the specific context of SCD hematopoietic stem/progenitor cell (HSPC) editing. Mitigating these toxicities is critical for enhancing the safety and efficacy profile of next-generation in vivo SCD therapies.
p53 Activation and the DNA Damage Response in Edited HSPCs
The introduction of a double-strand break (DSB) by Cas9 is recognized by the cell as DNA damage, triggering the classical p53 pathway. In HSPCs, a robust p53 response can lead to transient cell cycle arrest, senescence, or apoptosis, potentially reducing engraftment efficiency and long-term repopulating potential.
Key Quantitative Findings:
| Parameter | Findings in HSPCs/Cell Lines | Impact on SCD Editing |
|---|---|---|
| p53 Upregulation | 2- to 5-fold increase in protein levels and transcriptional activity post-editing. | May deplete the primitive HSPC pool, reducing engraftable cells. |
| Cell Cycle Arrest | Increase in G1/S phase population (from ~45% to ~65%) within 24h. | Delays ex vivo culture, may alter cell state. |
| Apoptosis Rate | Baseline apoptosis increases by 5-15% in edited vs. control HSPCs. | Direct loss of therapeutic cell product. |
| p53-Dependent Senescence | Observed in 10-20% of edited primary HSPCs in extended culture. | Impacts long-term regenerative capacity. |
Experimental Protocol: Assessing p53 Response in Edited CD34+ HSPCs
- HSPC Isolation & Culture: Isolate human CD34+ HSPCs from mobilized peripheral blood or cord blood. Pre-stimulate for 48h in serum-free medium supplemented with SCF, TPO, FLT3-L, and IL-3.
- Electroporation & Editing: Complex Cas9 ribonucleoprotein (RNP) with a sgRNA targeting the BCL11A enhancer. Electroporate 200,000-500,000 cells using a optimized protocol (e.g., Lonza 4D-Nucleofector, P3 Primary Cell kit).
- Control Samples: Include a non-electroporated control and an RNP with a non-targeting sgRNA.
- Time-Course Sampling: Harvest cells at 6, 24, 48, and 72 hours post-electroporation.
- Western Blot Analysis: Lyse cells, run protein extracts on SDS-PAGE, and probe for p53, phospho-p53 (Ser15), p21, and a loading control (e.g., β-actin).
- Flow Cytometry: Fix and permeabilize cells at 24h. Stain for intracellular p53 and p21, alongside viability dye (e.g., DAPI) and a proliferation marker (e.g., Ki67). Analyze the correlation between p53 levels and cell cycle status.
- qPCR for p53 Target Genes: Iscribe RNA and perform RT-qPCR for canonical targets (CDKN1A/p21, MDM2, PUMA).
Title: p53 Pathway Activation After Cas9 Cleavage in HSPCs
Chromosomal Aberrations: From On-Target Complexities to Off-Target Risks
Beyond intended edits, CRISPR-Cas9 can generate large, deleterious genomic rearrangements. These include on-target large deletions and chromosomal translocations resulting from concurrent cutting at off-target sites.
Quantitative Data on Aberrations:
| Aberration Type | Detection Method | Reported Frequency in HSPCs/Lymphocytes | Therapeutic Concern |
|---|---|---|---|
| On-Target Large Deletions (>1kb) | Long-range PCR, WGS | 1-10% of edited alleles (sgRNA-dependent). | Potential disruption of non-targeted regulatory regions or genes. |
| Chromosomal Deletions/Inversions | Karyotyping, FISH, WGS | <1-5% of edited cell populations. | Could inactivate tumor suppressor genes or create oncogenic fusions. |
| Chromosomal Translocations | WGS, CAST-Seq | Rare (<0.1%) but detectable between concurrently cut loci. | Driver event for genomic instability. |
| Complex On-Target Rearrangements (e.g., chromothripsis) | WGS | Rare, but reported in stem cell models. | Catastrophic, potentially oncogenic outcome. |
Experimental Protocol: Comprehensive Aberration Detection via WGS Analysis
- Cell Culture & Editing: Edit CD34+ HSPCs targeting BCL11A as described. Include a non-edited control. Culture cells for 7-14 days to allow potential aberrations to manifest.
- Single-Cell Derived Colony Picking: Plate edited cells in methylcellulose. After 14 days, pick 100-200 individual hematopoietic colonies (CFUs).
- Genomic DNA Extraction: Pool colonies from the same experimental condition (bulk analysis) or extract DNA from individual colonies (clonal analysis).
- Whole Genome Sequencing (WGS): Prepare sequencing libraries (e.g., Illumina PCR-free). Aim for high coverage (>30x for bulk, >50x for clonal).
- Bioinformatic Analysis Pipeline:
- Alignment: Map reads to human reference genome (hg38) using BWA-MEM.
- Variant Calling: Use DRAGEN or GATK for small indels.
- Structural Variant (SV) Calling: Employ multiple callers (e.g., Manta, DELLY, LUMPY) specialized for detecting deletions, duplications, inversions, and translocations.
- On-Target Analysis: Examine the ±50 kb region around the BCL11A cut site for SVs.
- Off-Target & Translocation Analysis: Cross-reference predicted off-target sites from in silico tools (Cas-OFFinder) with SV calls to identify translocations between on- and off-target loci.
- Clonal Analysis: For single-colony data, determine if an SV is present in all reads (clonal) or a subset (subclonal).
Title: WGS Workflow to Detect CRISPR-Induced Chromosomal Aberrations
Immune Responses to Cas9: Pre-existing Immunity and Therapeutic Implications
Cas9 proteins, derived from bacterial Streptococcus pyogenes (SpCas9) or Staphylococcus aureus (SaCas9), are foreign antigens. Seroprevalence studies indicate pre-existing humoral and cellular immunity in humans, posing risks for in vivo delivery, including potential inflammation, reduced editing efficiency, and immunotoxicity.
Quantitative Immune Prevalence Data:
| Immune Parameter | Assay | Reported Prevalence in Human Population | Implication for In Vivo SCD Therapy |
|---|---|---|---|
| Anti-SpCas9 Antibodies (IgG) | ELISA | 58-78% of adults seropositive. | May neutralize systemically delivered Cas9 RNP or vector. |
| Anti-SaCas9 Antibodies (IgG) | ELISA | Up to 95% seropositive. | High prevalence complicates in vivo use of SaCas9. |
| Cas9-Specific T Cells (SpCas9) | IFN-γ ELISpot | ~46% of donors reactive. | Risk of cytotoxic T-cell-mediated clearance of edited cells, especially HSPCs. |
| Cas9-Specific T Cells (SaCas9) | IFN-γ ELISpot | ~67% of donors reactive. | Even higher cellular immunity risk. |
Experimental Protocol: Assessing Pre-existing T-cell Responses to Cas9
- Donor PBMC Isolation: Collect peripheral blood from healthy donors (or SCD patients, if possible). Isolate PBMCs via density gradient centrifugation (Ficoll-Paque).
- Peptide Library Design: Synthesize overlapping 15-mer peptides spanning the entire SpCas9 or SaCas9 protein (e.g., 11-amino acid overlap). Pool peptides into sub-pools (e.g., by protein domain) for screening.
- IFN-γ ELISpot Assay: Coat ELISpot plates with anti-IFN-γ capture antibody overnight. Plate 200,000-400,000 PBMCs per well. Stimulate with:
- Individual peptide sub-pools (2 µg/mL per peptide).
- Positive control: Phytohemagglutinin (PHA) or CEF peptide pool.
- Negative control: DMSO (peptide solvent) or media alone.
- Critical Control: Include peptides from a non-Cas9 bacterial protein to assess baseline reactivity.
- Incubation & Development: Incubate plates for 40-48h at 37°C. Develop according to manufacturer's protocol (biotinylated detection antibody, streptavidin-ALP, BCIP/NBT substrate).
- Analysis: Count spots using an automated ELISpot reader. A response is considered positive if the mean spot-forming units (SFU) in the test well exceeds the mean of negative controls by a predetermined threshold (e.g., >2-fold and >10 SFU/10^6 PBMCs). Confirmatory intracellular cytokine staining (ICS) for IFN-γ and CD8/CD4 can be performed on positive hits.
The Scientist's Toolkit: Key Research Reagent Solutions
| Reagent/Material | Supplier Examples | Function in Toxicity Studies |
|---|---|---|
| Human CD34+ MicroBead Kit | Miltenyi Biotec | Isolation of high-purity HSPCs from source material for editing experiments. |
| Cas9 Nuclease (SpCas9, HiFi) | IDT, Thermo Fisher, Sigma-Aldrich | High-fidelity enzyme to reduce off-target cutting while maintaining on-target activity. |
| CRISPR-Cas9 sgRNA (Synthetgic) | IDT, Synthego | Chemically modified, high-purity sgRNA for complexing with Cas9 as RNP. |
| 4D-Nucleofector System & P3 Kit | Lonza | High-efficiency, low-toxicity delivery of RNP into primary HSPCs. |
| Anti-p53 (DO-1) Antibody | Santa Cruz Biotechnology | Detection of total p53 protein by western blot or flow cytometry. |
| Phospho-p53 (Ser15) Antibody | Cell Signaling Technology | Detection of activated p53 by western blot. |
| Annexin V Apoptosis Detection Kit | BioLegend, BD Biosciences | Quantification of early and late apoptotic cells post-editing. |
| PCR-free WGS Library Prep Kit | Illumina (TruSeq DNA PCR-Free) | Preparation of sequencing libraries without amplification bias for accurate SV detection. |
| SpCas9/SaCas9 Peptide Libraries | JPT Peptide Technologies | Overlapping peptide sets for comprehensive T-cell epitope mapping via ELISpot. |
| Human IFN-γ ELISpot Kit | Mabtech, BD Biosciences | Sensitive detection of antigen-specific T-cell responses from PBMCs. |
| Structural Variant Caller (Manta) | Illumina (via DRAGEN), Open Source | Bioinformatics tool specifically designed for sensitive detection of SVs from paired-end WGS data. |
Engraftment and Long-Term Repopulation Potential of Edited HSCs
Within the broader thesis on CRISPR-Cas9 mechanisms for sickle cell disease (SCD) gene editing research, a critical translational challenge is ensuring that edited hematopoietic stem cells (HSCs) successfully engraft in patients and sustain long-term, multilineage hematopoiesis. This whitepaper details the technical parameters, experimental protocols, and key findings central to evaluating this potential, focusing on preclinical and clinical data from SCD-focused therapies.
Table 1: Summary of Key Clinical/Preclinical Engraftment Data for SCD Gene Editing Therapies
| Therapy / Study (Reference) | Editing Platform | Target Gene | Patient/Model | Time to Neutrophil Engraftment (Days) | Time to Platelet Engraftment (Days) | Edited Cell Chimerism (% in PBMCs) at >12 Months | Vector Copy Number / Indel Efficiency |
|---|---|---|---|---|---|---|---|
| CLIMB SCD-121 (CTX001 exa-cel) | CRISPR-Cas9 RNP | BCL11A Erythroid Enhancer | SCD Patients (Phase 1/2) | ~30 (median) | ~40 (median) | ~80% (stable) | N/A (non-viral) |
| BMT CTN 1507 | CRISPR-Cas9 (mRNA) | BCL11A Enhancer | SCD Patients (Early Phase) | 28-35 (range) | 35-45 (range) | 70-90% (reported) | >80% allele editing in CD34+ |
| Preclinical NHP Study | CRISPR-Cas9 RNP | BCL11A Enhancer | Non-Human Primate | 10-15 | 15-20 | ~20-30% (long-term) | 50-70% in infused product |
| SCD CD34+ Xenograft (NSG) | Cas9-sgRNA RNP | HBB (Correction) | NSG Mouse Model | N/A | N/A | 15-25% human cell engraftment at 16-24 wks | 20-40% HDR correction rate |
Table 2: Factors Impacting Long-Term Repopulation Potential
| Factor | Optimal Range/Characteristic | Negative Impact |
|---|---|---|
| CD34+ Cell Viability Post-Editing | >80% | Reduced engraftment pool, delayed recovery |
| HSC Preservation (Phenotype CD34+CD90+CD45RA-) | High frequency maintained | Shift to progenitor dominance, short-term engraftment only |
| Myeloablative Conditioning | Busulfan AUC ~80-100 mg·h/L | Insufficient niche opening; excessive toxicity |
| Off-Target Editing Profile | No indels in predicted sensitive loci | Risk of clonal dominance or malignancy |
| Inflammatory Cytokine Storm | Minimal (post-infusion) | Potential for niche damage and reduced engraftment |
Detailed Experimental Protocols
Protocol:Ex VivoCRISPR Editing of Human SCD CD34+ HSCs for Engraftment Studies
Objective: To generate BCL11A enhancer-edited HSCs for functional assessment of engraftment in immunodeficient mice.
Materials: See "Scientist's Toolkit" below.
Procedure:
Mobilized Peripheral Blood CD34+ Cell Isolation: Isolate CD34+ cells from leukapheresis product of SCD patient (or healthy donor) using clinical-grade immunomagnetic beads. Assess viability (trypan blue) and purity (flow cytometry for CD34). Culture overnight in serum-free expansion medium (SFEM) supplemented with SCF (100 ng/mL), TPO (100 ng/mL), FLT3-L (100 ng/mL), and UM171 (35 nM).
Electroporation and RNP Delivery: Harvest cells, count, and resuspend in electroporation buffer at 1-2 x 10^6 cells/100 µL. For a single reaction, complex 60 µg of purified Cas9 protein with 200 pmol of synthesized sgRNA targeting the BCL11A erythroid enhancer (e.g., sequence: 5'-GATTATGGAGACCATTGTGG-3') to form ribonucleoprotein (RNP). Incubate 10 min at room temperature. Add RNP to cell suspension. Electroporate using a 4D-Nucleofector (program EO-100). Immediately transfer cells to pre-warmed recovery medium.
Post-Editing Culture and Analysis: Culture edited cells for 48-72 hours in cytokine-supplemented SFEM. Harvest an aliquot for analysis:
- Indel Efficiency: Extract genomic DNA. Amplify target locus by PCR. Perform T7 Endonuclease I (T7E1) assay or next-generation sequencing (NGS) to quantify indel percentage.
- Viability/Phenotype: Assess by flow cytometry using Annexin V/7-AAD and antibodies against CD34, CD90, CD45RA.
Mouse Transplantation (NSG Model): Irradiate 8-12 week-old NSG mice with a sublethal dose (1.5 Gy) 24 hours prior to transplant. Resuspend 2-5 x 10^5 edited or mock-treated CD34+ cells in PBS. Inject via tail vein. Provide supportive antibiotics in drinking water.
Peripheral Blood and Bone Marrow Analysis:
- Engraftment Tracking: From week 8 onwards, collect peripheral blood monthly. Lyse red blood cells and stain for human CD45, CD33 (myeloid), CD19 (B-cell), and CD3 (T-cell). Analyze by flow cytometry to determine % human chimerism and lineage distribution.
- Long-Term Assessment: Sacrifice cohorts at 16, 24, and 36 weeks. Flush bone marrow from femurs/tibiae. Perform detailed flow cytometry for human lineage markers and progenitor phenotypes (CD34+CD38-, etc.). Isulate genomic DNA from sorted human cells for NGS to determine editing persistence and clonal composition.
Protocol: Analysis of Long-Term Repopulating HSCs via Limiting Dilution Transplantation
Objective: Quantitatively measure the frequency of functional long-term HSCs (LT-HSCs) in edited vs. control grafts.
Procedure:
- Prepare a series of cell doses from the edited and control CD34+ cell products (e.g., 5x10^4, 1x10^5, 2x10^5, 5x10^5 cells/mouse).
- Transplant each dose into 10-12 irradiated NSG mice per group.
- At 16 and 24 weeks post-transplant, define a mouse as "positively engrafted" if human CD45+ chimerism in bone marrow is >0.1%.
- Input cell doses and positive engraftment data into ELDA software (http://bioinf.wehi.edu.au/software/elda/) to calculate the frequency of SCID-repopulating cells (SRC) and their 95% confidence intervals for each condition.
Diagrams
Title: Clinical Workflow for SCD HSC Gene Therapy
Title: CRISPR Repair Pathways in HBB Gene Correction
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for HSC Editing and Engraftment Assays
| Reagent/Material | Supplier Examples | Function in Protocol |
|---|---|---|
| StemSpan SFEM II | StemCell Technologies | Serum-free basal medium for human HSC culture, maintains stemness. |
| Recombinant Human SCF, TPO, FLT3-L | PeproTech | Essential cytokines for pre-stimulation and survival of primitive HSCs. |
| UM171 | STEMCELL Technologies | Small molecule agonist that expands LT-HSCs in culture. |
| Clinical-Grade Cas9 Protein | Aldevron, Thermo Fisher | High-purity, endotoxin-free Cas9 for RNP complex formation. |
| Synthetic sgRNA (target-specific) | Synthego, IDT | Chemically modified for stability; guides Cas9 to genomic target. |
| P3 Primary Cell 4D-Nucleofector Kit | Lonza | Optimized buffer and cuvettes for efficient RNP delivery into CD34+ cells. |
| Anti-human CD34 MicroBead Kit | Miltenyi Biotec | Immunomagnetic separation for high-purity CD34+ cell isolation. |
| Mouse Anti-Human CD45 APC | BioLegend | Antibody for flow cytometric detection of human cell engraftment in mouse blood/bone marrow. |
| T7 Endonuclease I | NEB | Enzyme for initial detection of CRISPR-induced indel mutations. |
| NGS Library Prep Kit for Amplicons | Illumina, Twist Bioscience | Prepares targeted amplicons for deep sequencing to quantify editing efficiency and spectrum. |
| Busulfan | Various | Myeloablative conditioning agent used in mouse models to open marrow niches. |
Overcoming Delivery Barriers for In Vivo CRISPR-Cas9 Approaches
The clinical success of ex vivo CRISPR-Cas9 therapy for sickle cell disease (SCD), as exemplified by Casgevy (exagamglogene autotemcel), represents a landmark achievement. However, this approach is complex, costly, and involves myeloablative conditioning. In vivo delivery of CRISPR-Cas9 components directly to a patient's hematopoietic stem and progenitor cells (HSPCs) or other relevant tissues offers a transformative alternative. This technical guide delves into the core barriers—cellular targeting, payload packaging, intracellular trafficking, and immune evasion—and outlines current experimental strategies to overcome them, specifically within the context of SCD gene editing targeting the BCL11A erythroid enhancer or the HBB gene.
Core Delivery Barriers & Quantitative Analysis
| Barrier Category | Specific Challenge | Quantitative Metrics (Current State) | Primary Investigational Solutions |
|---|---|---|---|
| Physical/Cellular | Targeting HSPCs in bone marrow niche | <5% of IV-injected LNPs localize to bone marrow; HSPC engagement <0.1% of dose. | Antibody-conjugated lipid nanoparticles (Ab-LNPs); Bone marrow-homing peptides (e.g., E-selectin ligands). |
| Payload Packaging | Co-delivery of large Cas9 mRNA (~4.5kb) and sgRNA(s) | LNP encapsulation efficiency: mRNA ~80-95%, sgRNA ~70-85%. Viral vector (AAV) cargo limit: <4.7kb. | Split-Cas9 systems; Smaller Cas9 orthologs (SaCas9 ~3.2kb); Dual-AAV or mRNA/sgRNA + AAV donor systems. |
| Intracellular Trafficking | Endosomal escape of cargo | <2% of internalized LNPs successfully release cargo into cytosol. | Ionizable lipids with pKa ~6.4; Endosomolytic peptides (e.g., GALA, HA2). |
| Immunogenicity | Pre-existing & adaptive immunity to Cas9 & vectors | Anti-Cas9 antibodies present in ~50-80% of population; Anti-AAV neutralizing antibodies in ~30-60%. | Engineered Cas9 variants with reduced immunogenicity; PEGylation of LNPs; Transient immunosuppression regimens. |
| Off-Target Editing | Unwanted genomic modifications in non-target cells | In vivo, off-target rate varies from <0.1% to >10% depending on delivery method and sgRNA design. | High-fidelity Cas9 variants (e.g., HiFi Cas9); Purified RNP delivery; Improved sgRNA design algorithms. |
Detailed Experimental Protocols
Protocol 1: Formulation & In Vivo Evaluation of Targeted LNPs for Murine HSPCs
Objective: To deliver Cas9 mRNA and sgRNA targeting the murine Bcl11a enhancer to HSPCs via intravenously injected CD117 (c-Kit) antibody-conjugated LNPs.
Materials: Ionizable lipid (e.g., DLin-MC3-DMA), phospholipid, cholesterol, PEG-lipid, maleimide-functionalized PEG-lipid. Cas9 mRNA (FLuc mRNA for biodistribution), Cy5-labeled sgRNA. Anti-mouse CD117 antibody (ACK2). Thiolation reagent (2-iminothiolane). β-mercaptoethanol. C57BL/6 mice.
Methodology:
- LNP Formation: Formulate LNPs via microfluidic mixing using an aqueous phase containing Cas9 mRNA and sgRNA at a 1:2 mass ratio and an ethanol phase containing lipids. Include 0.5 mol% maleimide-PEG-lipid.
- Antibody Conjugation: Reduce the anti-CD117 antibody with β-mercaptoethanol to generate free thiols. Purify via desalting column. Incubate thiolated antibody with LNPs at a 50:1 molar ratio (antibody:maleimide group) for 1h at room temperature. Quench with cysteine.
- Biodistribution: Inject 0.5 mg mRNA dose via tail vein. Image mice at 1, 4, 24, and 48h post-injection using an IVIS system (for FLuc) or isolate organs for Cy5 fluorescence quantification.
- Editing Analysis: Sacrifice mice at day 7. Harvest bone marrow, lineage-deplete, and enrich for HSPCs (c-Kit+). Extract genomic DNA. Assess Bcl11a enhancer editing efficiency via T7E1 assay or next-generation sequencing (NGS).
- Functional Readout: At 16 weeks post-treatment, analyze peripheral blood for HbF-positive red cells via FACS and perform HPLC for hemoglobin profiling.
Protocol 2: Dual AAV6 Delivery for HBB Gene Correction in a Humanized SCD Mouse Model
Objective: To co-deliver SaCas9 and an HBB donor template via two AAV6 vectors for homology-directed repair (HDR) in vivo.
Materials: AAV6 vectors: AAV6-SaCas9-U6-sgRNA (targeting the SCD mutation site) and AAV6-HDR-donor (containing ~800bp homology arms with corrective sequence). NSG mice engrafted with human CD34+ cells from an SCD patient.
Methodology:
- Vector Administration: At 12 weeks post-engraftment, inject mice intravenously with a 1:1 mixture of the two AAV6 vectors at a total dose of 5e11 vg/mouse.
- Kinetic Analysis: Bleed mice weekly for 8 weeks. Isolate human CD45+ cells from peripheral blood using FACS.
- Editing Assessment: Extract genomic DNA from sorted cells. Quantify HDR and indel frequencies at the HBB locus using droplet digital PCR (ddPCR) with allele-specific probes and NGS.
- Phenotypic Correction: At terminal timepoint, analyze erythroid differentiation in bone marrow. Perform single-cell RNA-seq on BFU-E colonies. Measure sickling propensity of derived red blood cells under hypoxia.
Visualization of Key Concepts
The Scientist's Toolkit: Research Reagent Solutions
| Reagent/Material | Supplier Examples | Function in In Vivo CRISPR Delivery |
|---|---|---|
| Ionizable Lipids (e.g., SM-102, DLin-MC3-DMA) | Avanti Polar Lipids, MedChemExpress | Critical for mRNA encapsulation and endosomal escape via pH-dependent membrane disruption. |
| AAV Serotype 6 (AAV6) | Vector Biolabs, SignaGen, Vigene Biosciences | High-efficiency transduction of human and murine HSPCs; preferred for hematopoietic gene therapy. |
| High-Fidelity Cas9 (HiFi Cas9) mRNA | Trilink Biotechnologies, Aldevron | Reduces off-target editing while maintaining on-target activity; crucial for safety. |
| Chemically Modified sgRNA (ms, ps) | Synthego, Integrated DNA Technologies | Enhances stability, reduces immunogenicity, and improves editing efficiency. |
| Anti-human CD117 (c-Kit) Antibody | BioLegend, Miltenyi Biotec | Targeting moiety for conjugation to LNPs or other carriers to direct them to HSPCs. |
| T7 Endonuclease I (T7E1) | New England Biolabs | Rapid, semi-quantitative assay for detecting indel mutations at target genomic loci. |
| ddPCR Assay for HDR Quantification | Bio-Rad | Absolute, sensitive quantification of low-frequency precise gene correction events. |
| Humanized SCD Mouse Model (e.g., Townes model) | The Jackson Laboratory | In vivo model expressing human α- and β-globin genes, enabling study of sickling and correction. |
Benchmarks and Efficacy: Analyzing Clinical Outcomes and Competing Technologies
This analysis situates pivotal clinical trials for sickle cell disease (SCD) within the broader mechanistic thesis of CRISPR-Cas9 gene editing. The fundamental thesis posits that precise, targeted disruption of the BCL11A gene enhancer or correction of the HBB gene point mutation via CRISPR-Cas9 can induce fetal hemoglobin (HbF) expression or restore adult hemoglobin (HbA), thereby ameliorating the pathophysiology of SCD. Trials such as CLIMB SCD-121 (exa-cel) and NCT04819841 (lovotibeglogene autotemcel) are the direct clinical translation of this hypothesis, testing the safety, efficacy, and durability of ex vivo hematopoietic stem/progenitor cell (HSPC) editing.
Detailed Trial Analysis
CLIMB SCD-121 (NCT03745287) – exagamglogene autotemcel (exa-cel)
Thesis Connection: This trial tests the hypothesis that CRISPR-Cas9-mediated disruption of the erythroid-specific enhancer region of the BCL11A gene in autologous CD34+ HSPCs will sufficiently reduce BCL11A expression to de-repress γ-globin synthesis, leading to clinically relevant increases in HbF and resolution of vaso-occlusive events (VOEs).
Experimental Protocol (Key Steps):
- Mobilization & Apheresis: Patients undergo mobilization with plerixafor and granulocyte colony-stimulating factor (G-CSF), followed by leukapheresis to collect peripheral blood CD34+ HSPCs.
- Manufacturing & Editing: CD34+ cells are electroporated with CRISPR-Cas9 components (SpCas9 protein and a single guide RNA, sgRNA, targeting the BCL11A enhancer).
- Myeloablative Conditioning: Patients receive busulfan conditioning to clear the bone marrow niche.
- Reinfusion: The CRISPR-edited autologous CD34+ cells are infused back into the patient.
- Engraftment & Follow-up: Patients are monitored for neutrophil/platelet engraftment, HbF levels, VOEs, and long-term safety.
NCT04819841 – lovotibeglogene autotemcel (lovo-cel)
Thesis Connection: This trial tests an alternative but complementary hypothesis: that using CRISPR-Cas9 to directly correct the causative E6V point mutation in the HBB gene in HSPCs can produce functional adult hemoglobin (HbA), thereby addressing the root genetic cause.
Experimental Protocol (Key Steps):
- Bone Marrow Harvest: Autologous CD34+ HSPCs are obtained via bone marrow aspiration under general anesthesia.
- Manufacturing & Editing: Cells are electroporated with Cas9 protein, a sgRNA targeting the HBB gene locus, and a single-stranded DNA template (ssODN) encoding the corrective sequence.
- Myeloablative Conditioning: Busulfan conditioning is administered.
- Reinfusion: Edited cells are reinfused.
- Engraftment & Follow-up: Monitoring includes engraftment, HbA production, total hemoglobin, and VOE frequency.
Table 1: Key Efficacy and Safety Outcomes from Pivotal Trials
| Parameter | CLIMB SCD-121 (exa-cel) | NCT04819841 (lovo-cel) |
|---|---|---|
| Primary Endpoint (VOC/ACS) | Freedom from severe VOEs for ≥12 consecutive months: 96.7% (29/30) | Resolution of severe VOEs (≥2/yr baseline): 88% (30/34) at 24 months |
| Mean/Median HbF (HbAT87Q) Level | Mean HbF (% of total Hb): ~40% at Month 24 | Median HbAT87Q (% of alleles): ~47% at Month 24 |
| Mean Total Hemoglobin | Increased to >11 g/dL from baseline (~9 g/dL) | Increased to >11 g/dL from baseline (~8.5 g/dL) |
| Neutrophil Engraftment | Median time: 29 days (range, 16-48) | Median time: 30 days (range, 17-54) |
| Platelet Engraftment | Median time: 35 days (range, 18-85) | Median time: 39 days (range, 26-108) |
| Key Safety Events | All patients had ≥1 AE. Most common: nausea, vomiting, fever. No malignancies related to treatment. | All patients had ≥1 AE. Most common: stomatitis, nausea, vomiting. No insertional mutagenesis events. |
| Follow-up Duration | Median: ~46.5 months (as of 2024) | Median: ~42 months (as of 2024) |
Visualizing CRISPR-Cas9 Mechanisms & Clinical Workflow
Diagram Title: CRISPR Thesis and Clinical Workflow for SCD
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for CRISPR-Cas9 SCD Research & Translation
| Reagent / Material | Function in SCD Gene Editing Research |
|---|---|
| CD34+ HSPCs (Primary Cells) | Target cell population for editing; sourced from mobilized peripheral blood or bone marrow. The "raw material" for therapy. |
| CRISPR-Cas9 RNP Complex | The core editing machinery. Recombinant SpCas9 protein pre-complexed with target-specific sgRNA (e.g., targeting BCL11A enhancer or HBB locus) for efficient delivery and rapid degradation to limit off-target exposure. |
| Single-Stranded DNA Donor Template (ssODN) | For homology-directed repair (HDR)-based correction (as in lovo-cel). Provides the correct DNA sequence to replace the pathogenic HBB mutation. |
| Electroporation System | (e.g., Lonza 4D-Nucleofector). Enables efficient, non-viral delivery of RNP complexes and donor templates into sensitive primary CD34+ cells. |
| Myeloablative Agent (Busulfan) | Critical pre-conditioning drug. Creates bone marrow niche space for the engraftment and dominance of the infused, edited HSPCs. |
| Cell Culture Media (Serum-Free) | Specialized formulations (e.g., StemSpan) that support the survival, maintenance, and minimal expansion of CD34+ HSPCs during the ex vivo editing process. |
| QC Assays: NGS for On/Off-target | Deep sequencing (e.g., GUIDE-seq, CIRCLE-seq, targeted NGS) to quantify editing efficiency at the intended locus and screen for potential off-target modifications. |
| HPLC / Mass Spectrometry | To quantify hemoglobin variant levels (HbS, HbF, HbAT87Q) in vitro and in patient samples, providing a direct readout of functional editing efficacy. |
| Clonogenic Assays (CFU, LTC-IC) | Functional assays to ensure the ex vivo editing process does not impair the long-term multi-lineage differentiation potential and stemness of HSPCs. |
This technical guide frames the clinical and hematological efficacy metrics for sickle cell disease (SCD) therapies within the context of CRISPR-Cas9 gene-editing research. The primary therapeutic goal of CRISPR-based interventions, such as ex vivo editing of hematopoietic stem/progenitor cells (HSPCs) to reactivate fetal hemoglobin (HbF), is to alter the underlying pathophysiology of SCD. The quantifiable clinical outputs of this genetic manipulation are measured through three core efficacy endpoints: elevation of HbF levels, reduction in the frequency of vaso-occlusive crises (VOCs), and achievement of stable, non-anemic hemoglobin levels. This document details the methodologies for measuring these endpoints and their interrelationships in clinical trial design.
Core Efficacy Metrics: Definitions and Measurement Protocols
HbF (Fetal Hemoglobin) Levels
HbF (α2γ2) is a potent inhibitor of HbS polymerization. Its reactivation via CRISPR-Cas9-mediated disruption of the BCL11A erythroid enhancer or similar targets is a primary mechanism of action in current SCD gene-editing trials.
- Quantitative Measurement: Expressed as the percentage of total hemoglobin or as HbF concentration (g/dL). The most critical submetric is HbF-containing erythrocytes (F-cells), specifically the F-cell proportion and the amount of HbF per F-cell (pF-cell).
- Standard Analytical Protocol:
- Sample: Peripheral blood collected in EDTA tubes.
- Primary Method: High-Performance Liquid Chromatography (HPLC).
- Protocol: Hemolysate is prepared. A fixed volume is injected into an HPLC system equipped with a cation-exchange column (e.g., Bio-Rad VARIANT II). Hemoglobins are eluted using a gradient of increasing ionic strength buffers (e.g., phosphate buffers, pH ~6.8). HbF elutes at a characteristic retention time and is quantified by integrating the area under the peak, reported as a percentage of total hemoglobin.
- Confirmatory/Supplementary Methods:
- Capillary Electrophoresis (CE): Separates hemoglobins based on charge in an alkaline buffer.
- Flow Cytometry for F-cells: Cells are fixed, permeabilized, and stained with a fluorescently labeled monoclonal anti-HbF antibody (e.g., FITC-conjugated). Analysis identifies the percentage of RBCs containing detectable HbF.
Vaso-Occlusive Crisis (VOC) Reduction
VOCs are the acute, painful hallmark of SCD. Reduction in VOC rate is the leading primary endpoint for pivotal Phase 3 trials.
- Definition: A VOC is typically defined as an acute episode of pain with no medically determined cause other than a vaso-occlusion, requiring parenteral narcotics or an increase in home analgesic management, and lasting >2 hours.
- Measurement Protocol in Clinical Trials:
- Prospective, Event-Driven Documentation: Patients (or guardians) use an electronic diary to record pain scores and analgesic use daily.
- Clinical Validation: Each reported event is adjudicated by a blinded clinical endpoint committee against the trial's protocol-defined criteria.
- Key Metric Calculation: The annualized rate of VOCs is calculated for each patient as: (Total number of adjudicated VOCs over the efficacy period / Total number of days in the period) * 365.25.
- Statistical Analysis: The primary comparison is often the proportion of patients achieving a pre-defined threshold reduction (e.g., >50% reduction) or the difference in mean annualized VOC rates between treatment and control arms using a negative binomial regression model.
Hemoglobin Stabilization
This metric assesses the improvement in chronic anemia and hemolytic rate.
- Components:
- Hemoglobin Level: Steady-state total hemoglobin concentration (g/dL).
- Hemolysis Markers: Indirect bilirubin, lactate dehydrogenase (LDH), aspartate aminotransferase (AST), and reticulocyte count.
- Red Blood Cell (RBC) Survival: Measured via half-life of biotin-labeled RBCs in research settings.
- Measurement Protocol:
- Steady-State Sampling: Blood samples are taken during a "well" period, typically defined as >30 days from any acute event, transfusion, or hospitalization.
- Standard Clinical Assays: Hemoglobin is measured via automated hematology analyzer. Reticulocyte count is performed via flow cytometry. LDH and bilirubin are measured via standard clinical chemistry platforms.
- Stability Definition: An increase in mean hemoglobin of >1 g/dL from baseline sustained over multiple visits (e.g., at 6, 12, and 18 months post-treatment) is considered clinically significant stabilization.
Table 1: Efficacy Outcomes from Selected CRISPR-Cas9 SCD Clinical Trials (as of 2023-2024)
| Trial / Intervention (Target) | Patients (n) | Mean HbF Increase (ppt. or g/dL) | Mean Hb Increase (g/dL) | VOC Rate Reduction | Follow-up Duration |
|---|---|---|---|---|---|
| CLIMB SCD-121 (exa-cel, BCL11A enhancer) | >30 | HbF%: ~40% (from <10%) F-cells: ~95% | ~3.0 g/dL | 98.3% (0 VOC vs 3.9/yr baseline) | 24-36 months |
| BMT-CTN 1904 (lovo-cel, BCL11A enhancer) | >30 | HbF%: ~30-40% | ~2.0-3.0 g/dL | ~90% reduction | 12-24 months |
| Standard Care (Historical) | - | <10% (typically) | 6-9 g/dL (baseline) | N/A | N/A |
Table 2: Correlation of Hematological and Clinical Efficacy Metrics
| Hematological/Biomarker Change | Direct Physiological Consequence | Resulting Clinical Efficacy Metric |
|---|---|---|
| ↑ HbF % (>20-30%) & ↑ F-cells | Inhibition of intracellular HbS polymerization | Reduced RBC sickling & hemolysis |
| ↓ Hemolysis (↓ LDH, ↓ Bilirubin) | Improved RBC survival, ↓ endothelial damage | Hemoglobin Stabilization (↑ Hb) |
| ↑ Total Hb & ↓ Sickling | Improved microvascular flow, ↓ inflammation | VOC Reduction |
Experimental Workflow: From CRISPR Editing to Efficacy Readouts
(Title: CRISPR to Clinical Efficacy Workflow in SCD)
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents and Materials for SCD Gene-Editing Efficacy Research
| Item | Function/Application in SCD Research | Example/Note |
|---|---|---|
| CRISPR-Cas9 RNP Complex | Direct delivery of Cas9 protein and target-specific gRNA for precise BCL11A enhancer editing. Ensures rapid activity and degradation to minimize off-targets. | Synthesized gRNA targeting HS2 of BCL11A + recombinant S. pyogenes Cas9 protein. |
| CD34+ HSPC Isolation Kit | Positive selection of human hematopoietic stem/progenitor cells from mobilized peripheral blood or bone marrow for ex vivo editing. | Miltenyi Biotec CD34 MicroBead Kit (human). |
| StemSpan SFEM II Medium | Serum-free, cytokine-supplemented medium optimized for maintenance and expansion of human HSPCs during the editing and culture process. | From STEMCELL Technologies. |
| Anti-HbF Antibody (FITC) | Flow cytometric identification and quantification of F-cells (HbF-containing RBCs) from peripheral blood or in vitro erythroid differentiation cultures. | Clone HbF-1 (e.g., from Santa Cruz Biotechnology). |
| HPLC System for Hemoglobin | Gold-standard quantitative separation and measurement of HbA, HbS, HbF, and other hemoglobin variants. | Bio-Rad VARIANT II HPLC system with the β-Thalassemia Short Cellulose cartridge. |
| Human Erythroid Differentiation Media | To differentiate edited CD34+ cells into mature erythroid lineages in vitro to assess HbF induction potency pre-clinically. | A three-phase system using SCF, EPO, IL-3, etc. |
| ddPCR/NGS Off-Target Assay Kits | Detection of potential off-target editing events. Critical for safety assessment. | Assays designed against in silico predicted off-target sites. |
Context within Thesis on CRISPR-Cas9 for SCD: This analysis directly compares the safety architectures of the two primary gene therapy modalities—CRISPR-Cas9 gene editing and lentiviral vector (LV)-mediated gene addition—currently approved or in advanced development for sickle cell disease (SCD). It critically evaluates their risk profiles to inform therapeutic development strategies.
CRISPR-Cas9 (e.g., exagamglogene autotemcel/exa-cel): Utilizes a Cas9 nuclease and a single guide RNA (sgRNA) to create a targeted double-strand break (DSB) in the BCL11A gene enhancer. This disrupts BCL11A expression, thereby inducing fetal hemoglobin (HbF) production to compensate for defective adult hemoglobin. Safety revolves around the precision of on-target editing and the potential for off-target genomic alterations.
Lentiviral Gene Therapy (e.g., lovotibeglogene autotemcel/lovo-cel): Employs a replication-incompetent, self-inactivating (SIN) lentiviral vector to deliver a functional β-globin gene (β^A-T87Q) into patient hematopoietic stem and progenitor cells (HSPCs). The therapeutic gene integrates semi-randomly into the host genome to produce anti-sickling hemoglobin. Safety centers on vector integration site profiles and genotoxicity risks.
Table 1: Key Clinical Safety Metrics from Approved/Late-Stage SCD Therapies
| Safety Parameter | CRISPR-Cas9 (exa-cel) | Lentiviral (lovo-cel) |
|---|---|---|
| Therapy Name | Exagamglogene autotemcel | Lovotibeglogene autotemcel |
| Target/Transgene | BCL11A Erythroid Enhancer | β^A-T87Q Globin Gene |
| Primary Safety Events | Myelosuppression from conditioning; Theoretical off-target editing. | Myelosuppression from conditioning; Clonal expansion monitoring. |
| Incidence of Vector/Editor-Related SAEs | Low; No clear pattern linking DSBs to SAEs. | Low; No reported vector-derived replication-competent lentivirus (RCL). |
| Risk of Insertional Oncogenesis | Very Low (Theoretical, due to on-target DSB repair). | Low (Managed by SIN design and integration site monitoring). |
| Immunogenicity to Component | Anti-Cas9 antibodies common; clinical significance unclear. | Anti-transgene product antibodies rarely reported. |
| Key Monitoring Requirements | Off-target analysis (computational & cellular), long-term clonal tracking. | Integration site analysis (LAM-PCR, NGS) for >15 years. |
Table 2: Laboratory Safety Assessment Parameters
| Assessment | CRISPR-Cas9 Platform | Lentiviral Vector Platform |
|---|---|---|
| Genotoxicity Assays | GUIDE-seq, CIRCLE-seq, in silico prediction, long-term clonal outgrowth assays. | In vitro genotoxicity (e.g., murine myeloid proliferation assay), vector copy number (VCN) distribution. |
| Primary Readout | Off-target site identification & frequency; chromosomal rearrangements. | Integration site preference (relative distance from TSS, oncogenes); clonal dominance. |
| Typical Finding in SCD Therapies | Off-target sites identified at very low frequencies (<0.1%), not in coding regions. | Polyclonal integration pattern; no evidence of preferential integration near oncogenes in clinical samples. |
Detailed Experimental Protocols for Safety Assessment
Protocol 3.1: In Vitro Off-Target Analysis for CRISPR-Cas9 (GUIDE-seq)
Purpose: Genome-wide identification of CRISPR-Cas9 off-target double-strand breaks. Reagents: Cas9:sgRNA RNP complex, GUIDE-seq oligonucleotides, transfection reagent, HeLa or HEK293T cells, PCR reagents, NGS library prep kit. Procedure:
- Transfect cells with Cas9 RNP and double-stranded GUIDE-seq oligos.
- Incubate for 48-72 hours. Harvest genomic DNA.
- Perform tag-specific PCR amplification to enrich for genomic junctions containing the integrated oligo.
- Prepare NGS libraries and sequence. Analyze sequences using the GUIDE-seq computational pipeline to map double-strand break sites.
- Validate top candidate off-target sites via targeted amplicon sequencing.
Protocol 3.2: Integration Site Analysis for Lentiviral Therapies (Linear Amplification-Mediated PCR - LAM-PCR)
Purpose: To map genomic locations of lentiviral vector integrations in transduced cell populations. Reagents: Genomic DNA from transduced cells, biotinylated LV LTR-specific primer, restriction enzymes, linkers, streptavidin magnetic beads, PCR reagents. Procedure:
- Digest genomic DNA with a frequent-cutter restriction enzyme (e.g., MseI).
- Ligate a known double-stranded linker to the digested fragments.
- Perform linear PCR using a biotinylated primer specific to the LV LTR.
- Capture biotinylated products using streptavidin beads.
- Perform nested PCR using primers for the linker and LV LTR.
- Purity and sequence PCR products. Align sequences to the reference genome to identify integration sites. Analyze for clustering near oncogenes.
Visualization of Mechanisms and Safety Assessments
Diagram Title: CRISPR-Cas9 SCD Therapy Safety Workflow
Diagram Title: Lentiviral SCD Therapy Safety Workflow
Diagram Title: Comparative Primary Risk Profiles
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for Safety Assessment Experiments
| Reagent/Material | Function | Example Vendor/Cat. No. (Illustrative) |
|---|---|---|
| Recombinant S.p. Cas9 Nuclease | Creates targeted double-strand breaks in CRISPR editing. | IDT, Thermo Fisher Scientific. |
| Chemically Modified sgRNA | Guides Cas9 to specific genomic locus; modifications enhance stability. | Synthego, Horizon Discovery. |
| GUIDE-seq Oligonucleotides | Double-stranded oligos that tag and mark sites of DSBs for NGS identification. | Integrated DNA Technologies. |
| CIRCLE-seq Kit | In vitro genome-wide circularization assay for unbiased off-target site profiling. | Published protocol; requires Tn5 transposase, NGS reagents. |
| SIN Lentiviral Vector Particles | Replication-incompetent lentivirus for gene delivery; clinical-grade for GMP. | Vector production via contract manufacturers (e.g., Oxford Biomedica). |
| LAM-PCR Kit | All necessary primers, linkers, and enzymes for linear amplification-mediated PCR. | Published protocol; reagents from Thermo Fisher, NEB. |
| Next-Generation Sequencing (NGS) Platform | For high-throughput analysis of off-target sites and integration sites. | Illumina MiSeq/NovaSeq. |
| Murine Myeloid Proliferation Assay | In vitro assay to assess potential for vector-induced clonal expansion. | Custom assay using primary murine bone marrow cells. |
| ddPCR Assay for VCN | Absolute quantification of vector copies per genome in transduced cells. | Bio-Rad, ddPCR LV Copy Number Assays. |
Comparative Analysis with Other Gene Editing Platforms (ZFNs, TALENs) for SCD
Within the broader thesis on CRISPR-Cas9 mechanisms for Sickle Cell Disease (SCD) research, a critical evaluation of alternative nuclease platforms is essential. This whitepaper provides a technical comparison between Zinc Finger Nucleases (ZFNs), Transcription Activator-Like Effector Nucleases (TALENs), and CRISPR-Cas9, focusing on their application in correcting the sickle β-globin (HBB) mutation. Each platform induces double-strand breaks (DSBs) to facilitate gene correction via Homology-Directed Repair (HDR) or targeted gene disruption via Non-Homologous End Joining (NHEJ).
Platform Mechanism & Design
- CRISPR-Cas9: Utilizes a guide RNA (gRNA) complementary to a ~20 bp target DNA sequence to direct the Cas9 nuclease. The Protospacer Adjacent Motif (PAM, e.g., 5'-NGG-3' for SpCas9) is a key targeting constraint.
- TALENs: Engineered fusion proteins. The DNA-binding domain consists of TALE repeats, each recognizing a single nucleotide (via Repeat Variable Diresidues, RVDs). This domain is fused to the FokI nuclease dimer.
- ZFNs: Engineered fusion proteins. The DNA-binding domain consists of zinc finger motifs, each typically recognizing a 3-bp triplet. This domain is fused to the FokI nuclease dimer, which requires paired binding sites in opposite orientations for dimerization and cleavage.
Diagram 1: Gene Editing Platform DNA Recognition & Cleavage (89 chars)
Quantitative Comparison for SCD Applications
Table 1: Platform Comparison for SCD Gene Editing
| Parameter | CRISPR-Cas9 | TALENs | ZFNs |
|---|---|---|---|
| Targeting Constraint | Requires PAM (e.g., NGG) near target. | Requires 5'-T at start of each binding site. | Requires G-rich triplet sequences; context-dependent recognition. |
| Design & Cloning | Simple, rapid (gRNA synthesis). | Complex, repetitive TALE assembly. | Complex, context-dependent zinc finger engineering. |
| Targeting Specificity | Moderate; potential for off-targets due to gRNA tolerance. | High; longer, more specific binding domain. | High; but can be limited by cross-reactivity of zinc fingers. |
| Efficiency (HDR in HSPCs) | Typically 20-50% for HBB correction. | Typically 10-30% for HBB correction. | Typically 5-20% for HBB correction. |
| Delivery to HSPCs | High (RNP electroporation). | Moderate (mRNA or protein electroporation). | Low-Moderate (often viral delivery required). |
| Immunogenicity Risk | Moderate-High (pre-existing anti-Cas9 antibodies). | Low (bacterial-derived protein). | Moderate (potential immune response to FokI domain). |
| Clinical Status (SCD) | Approved (exa-cel, lovo-cel). | Preclinical research. | Phase 1/2 trials (SB-318 for related disease). |
Experimental Protocol: Ex Vivo HDR in CD34+ HSPCs
This protocol outlines the core methodology for gene correction of the HBB E6V mutation using CRISPR-Cas9 HDR, adaptable for TALEN/ZFN protein delivery.
A. Materials: CD34+ HSPC Isolation & Culture
- Mobilized peripheral blood or cord blood.
- Clinical-grade cytokines (SCF, TPO, FLT3L).
- STEMCELL Technologies CD34+ isolation kit.
- Serum-free expansion medium (e.g., StemSpan).
B. Design & Preparation of Editing Components
- CRISPR-Cas9: Synthesize high-fidelity Cas9 mRNA and chemically modified sgRNA targeting near the HBB E6V codon. Prepare a single-stranded oligodeoxynucleotide (ssODN) HDR template containing the corrective sequence (A to T) and silent restriction site for screening.
- TALENs/ZFNs: Obtain validated mRNA sequences encoding the paired TALEN or ZFN proteins targeting the HBB locus. Use the same ssODN HDR template as for CRISPR.
C. Electroporation & Editing
- Pre-stimulate purified CD34+ cells for 24-48 hours in cytokine-rich medium.
- For CRISPR-Cas9: Combine Cas9 protein (RNP complex) with sgRNA and HDR template. For TALENs/ZFNs: Combine paired mRNAs with HDR template.
- Electroporate cells using a optimized system (e.g., Lonza 4D-Nucleofector, P3 Primary Cell Kit, program DZ-100).
- Immediately transfer cells to pre-warmed recovery medium, then culture in expansion medium with cytokines.
D. Analysis of Editing Outcomes
- Genomic DNA Extraction: Harvest cells at 72h (for indel analysis) and day 7-14 (for HDR analysis).
- Next-Generation Sequencing (NGS): Amplify the target locus via PCR. Use NGS to quantify total indels (NHEJ) and precise HDR events.
- Functional Assay: Differentiate edited cells into erythroid lineage and perform HPLC to measure fetal (HbF) and adult hemoglobin, assessing correction of sickle phenotype.
Diagram 2: Ex Vivo Gene Editing Workflow for SCD (62 chars)
The Scientist's Toolkit: Key Reagents for SCD Gene Editing
Table 2: Essential Research Reagents & Materials
| Reagent/Material | Function in SCD Experiment | Example Vendor/Product |
|---|---|---|
| Human CD34+ HSPCs | Primary target cell for ex vivo editing and transplantation models. | AllCells, StemCell Technologies. |
| Cas9 Nuclease (HiFi) | High-fidelity nuclease enzyme for reduced off-target cleavage. | Integrated DNA Technologies (IDT), Aldevron. |
| Chemically-modified sgRNA | Enhances stability and reduces immune activation in primary cells. | Synthego, Trilink BioTechnologies. |
| TALEN mRNA Pair | Encodes the TALEN proteins for HBB targeting; requires in vitro transcription. | GeneArt (Thermo Fisher), custom synthesis. |
| ssODN HDR Template | ~100-200 nt DNA donor template with corrective sequence for precise repair. | IDT Ultramer, Eurofins Genomics. |
| Nucleofector System | High-efficiency electroporation device for hard-to-transfect HSPCs. | Lonza 4D-Nucleofector X Unit. |
| Cytokine Cocktail | Supports survival, proliferation, and stemness of HSPCs during editing. | PeproTech (SCF, TPO, FLT3L). |
| NGS Amplicon-EZ Kit | For preparation of target amplicon libraries for deep sequencing analysis. | Azenta/Genewiz, Illumina. |
| Erythroid Differentiation Kit | Drives edited HSPCs into red blood cell lineage for functional validation. | STEMdiff Erythroid Kit. |
While ZFNs and TALENs demonstrated the feasibility of targeted HBB editing, CRISPR-Cas9's unparalleled design simplicity, high efficiency, and multiplexing capability have accelerated its translation into approved therapies for SCD. The choice of platform hinges on the specific application: TALENs may offer advantages where supreme specificity is critical, but CRISPR-Cas9 currently provides the most effective and tractable path for complex ex vivo autologous cell therapies. Future work will focus on refining the safety and delivery of all platforms to broaden therapeutic access.
Regulatory and Commercialization Pathways for CRISPR-Based SCD Therapies
The application of CRISPR-Cas9 for sickle cell disease (SCD) represents a paradigm shift in genetic medicine, moving from symptom management towards a potential functional cure. Within the broader thesis on CRISPR-Cas9 mechanisms, SCD serves as a pioneering model due to its well-characterized monogenic etiology—primarily the HBB gene mutation (HbS). The therapeutic strategy involves ex vivo editing of autologous hematopoietic stem and progenitor cells (HSPCs) to either reactivate fetal hemoglobin (HbF) via BCL11A suppression or directly correct the HbS point mutation. This whitepaper delineates the technical, regulatory, and commercial frameworks essential for translating this mechanistic research into an approved therapy.
Technical Guide: Core Editing Strategies & Protocols
Key Therapeutic Approaches
2.1.1 BCL11A Erythroid Enhancer Disruption
- Mechanism: CRISPR-Cas9 generates double-strand breaks (DSBs) in the +58 BCL11A erythroid-specific enhancer region in the BCL11A gene. This disrupts BCL11A expression in erythroid lineage cells, relieving its repression of the HBG1/HBG2 genes, thereby inducing HbF production.
- Rationale: HbF inhibits the polymerization of HbS, preventing sickling of red blood cells.
2.1.2 Direct HbS Correction (Gene Conversion)
- Mechanism: Utilizes CRISPR-Cas9 with a donor DNA template to facilitate homology-directed repair (HDR). This converts the pathogenic A•T nucleotide pair at codon 6 of HBB back to the wild-type G•C, restoring normal adult hemoglobin (HbA) production.
Detailed Experimental Protocol for Ex Vivo HSPC Editing
This protocol outlines the core process used in clinical trials like those for exagamglogene autotemcel (exa-cel).
Protocol: CRISPR-Cas9 Editing of Human CD34+ HSPCs for BCL11A Enhancer Disruption
I. HSPC Mobilization, Apheresis, and Isolation
- Mobilization: Administer granulocyte colony-stimulating factor (G-CSF) and plerixafor to the patient to mobilize HSPCs from bone marrow to peripheral blood.
- Apheresis: Collect mobilized cells via leukapheresis. Target collection: ≥ 4.0 x 10^6 CD34+ cells/kg patient weight.
- CD34+ Selection: Isulate CD34+ HSPCs using clinical-grade immunomagnetic selection (e.g., CliniMACS system). Assess viability (must be >95%) and purity (typically >90% CD34+).
II. Electroporation and RNP Delivery
- RNP Complex Formation: Complex high-fidelity Cas9 protein (e.g., SpyFi Cas9) with synthetic single-guide RNA (sgRNA) targeting the BCL11A +58 enhancer. Use a molar ratio of 1:2.5 (Cas9:sgRNA). Incubate at room temperature for 10-20 minutes to form the ribonucleoprotein (RNP).
- Electroporation: Wash and resuspend CD34+ cells in electroporation buffer at a concentration of 1-2 x 10^8 cells/mL. Mix cells with RNP complex (typical dose: 100-200 nM RNP). Electroporate using a clinical-grade system (e.g., MaxCyte GT or Lonza 4D-Nucleofector) with an optimized pulse code (e.g., EO-115 for MaxCyte).
- Post-Electroporation Recovery: Immediately transfer cells to pre-warmed, serum-free culture medium supplemented with cytokines (SCF, TPO, FLT3L). Incubate at 37°C, 5% CO2 for 1-3 hours before proceeding.
III. Cell Expansion and Quality Control
- Culture: Maintain cells in gas-permeable cell culture bags in a closed, automated bioreactor system (e.g., Miltenyi Prodigy) for 5-7 days. Medium is supplemented with cytokines to support maintenance and modest expansion.
- QC Testing:
- Editing Efficiency: Assess INDEL frequency at the target site via next-generation sequencing (NGS) of bulk cells. Target range: 70-90%.
- Viability and Yield: Confirm cell count and viability (>70%) meet release specifications.
- Sterility: Perform endotoxin, mycoplasma, and bacterial/fungal culture tests.
- Vector Copy Number: Confirm absence of recombinant lentivirus if used in research comparators.
- Karyotype: Perform G-band analysis to rule out chromosomal abnormalities.
IV. Myeloablative Conditioning and Reinfusion
- Patient Conditioning: After apheresis, the patient undergoes myeloablative chemotherapy (busulfan) to clear residual bone marrow and enable engraftment of edited cells.
- Product Wash and Formulation: Harvest edited cells, wash, and formulate in infusion medium containing cryoprotectant (DMSO).
- Cryopreservation and Release: Cryopreserve the final drug product (DP) in a controlled-rate freezer. Final DP release requires passing all QC specifications.
- Reinfusion: Thaw the DP at the patient's bedside and administer via intravenous infusion.
Table 1: Clinical Outcomes from Pivotal CRISPR-SCD Trials (as of 2024)
| Therapy (Sponsor) | Target | Trial Phase | Patients Treated (n) | HbF Induction (Mean) | VOC-Free Survival | Key Regulatory Designation |
|---|---|---|---|---|---|---|
| exa-cel (Vertex/CRISPR Tx) | BCL11A Enhancer | Phase 3 (CLIMB SCD-121) | 30+ | >40% of total Hb | 100% at 24 mos (n=30) | FDA Approved (12/2023), EMA PRIME |
| OTQ923 (Novartis/Intellia) | BCL11A Enhancer | Phase 1/2 | 10 | ~25% F-cells | Data pending | Fast Track (FDA) |
| BCH-BB694 (Boston Children's) | BCL11A +58 & -123 | Phase 1/2 | 5 | >30% HbF | 100% at 18 mos | Orphan Drug |
Table 2: Key Analytical QC Benchmarks for Drug Product Release
| Parameter | Acceptance Criterion | Analytical Method |
|---|---|---|
| Viability | ≥ 70% | Flow cytometry (7-AAD) |
| Purity (CD34+) | ≥ 70% | Flow cytometry |
| Editing Efficiency (INDEL%) | ≥ 60% | NGS (amplicon-based) |
| Sterility (Bacterial/Fungal) | No growth | BacT/ALERT culture |
| Mycoplasma | Not Detected | PCR-based assay |
| Endotoxin | ≤ 5 EU/kg/hr | LAL test |
| Potency | Engraftment in NSG mice or HbF induction in erythroid differentiation assay | In vivo or in vitro functional assay |
Regulatory Pathways
FDA Framework: CBER/OTAT
CRISPR-SCD therapies are regulated as gene therapy products under the Biologics License Application (BLA) pathway (21 CFR 601). They are classified as more-than-minimally manipulated cellular therapies.
Critical Regulatory Milestones:
- Pre-IND Meeting: Align on chemistry, manufacturing, and controls (CMC), preclinical, and clinical plans.
- Investigational New Drug (IND) Application: Requires robust data on:
- Manufacturing: Master Cell Bank characterization, process validation, DP release criteria.
- Preclinical: Proof-of-concept and biodistribution in murine models (e.g., Townes SCD mice), off-target analysis (e.g., GUIDE-seq, CIRCLE-seq), and tumorigenicity risk assessment.
- Clinical: Detailed protocol for Phase 1/2 study.
- Accelerated Pathways: Utilize Regenerative Medicine Advanced Therapy (RMAT), Breakthrough Therapy, and Fast Track designations for intensive FDA interaction.
- BLA Submission & Review: Requires complete data from pivotal trials demonstrating substantial evidence of efficacy and safety. Priority Review is typical.
EMA Framework: CAT/CHMP
Governed under Advanced Therapy Medicinal Products (ATMP) regulation (EC No 1394/2007).
Key Elements:
- Scientific Advice & PRIME: Early interaction via PRIority MEdicines scheme for enhanced guidance.
- Marketing Authorization Application (MAA): Submitted to the Committee for Medicinal Products for Human Use (CHMP) with full risk-benefit assessment.
- Hospital Exemption: National pathway for non-routine use of non-authorized ATMPs, but not a substitute for MAA.
Commercialization & Market Access
4.1 Manufacturing & Supply Chain
- Model: Decentralized or centralized. Patient-specific apheresis shipped to a central GMP facility for editing, then shipped back.
- Challenges: Scalability, vein-to-vein time, cost of goods (dominated by cytokines, single-use materials, and quality control).
- Automation: Adoption of closed, automated systems (e.g., Miltenyi Prodigy, Lonza Cocoon) to improve robustness and scalability.
4.2 Pricing, Reimbursement, and Market Shaping
- Pricing: Exa-cel (Casgevy) list price: ~$2.2 million in the US. Justified by high initial efficacy, potential lifelong benefit, and cost-offset from avoided VOC crises, strokes, and chronic care.
- Reimbursement Models: Outcomes-based/annuity-based payments negotiated with payers to mitigate risk and spread cost over time.
- Market Access Barriers: Need for specialized Treatment Centers with expertise in stem cell transplantation, gene therapy administration, and SCD management. Significant upfront capital investment required.
Visualizations
(CRISPR Disruption of BCL11A Enhancer Activates HbF)
(From R&D to Market: CRISPR-SCD Therapy Pathway)
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Key Research Reagents for CRISPR-SCD Experiments
| Reagent / Material | Function / Role | Example Vendor/Product |
|---|---|---|
| High-Fidelity Cas9 Nuclease | Minimizes off-target editing while maintaining on-target efficiency. Essential for translational research. | IDT Alt-R S.p. HiFi Cas9, Thermo Fisher TrueCut Cas9 Protein v2 |
| Synthetic sgRNA | Guides Cas9 to the specific genomic target (e.g., BCL11A enhancer). Chemically modified for stability. | Synthego sgRNA, IDT Alt-R CRISPR-Cas9 sgRNA |
| CD34+ HSPC Isolation Kit | Immunomagnetic positive selection of human CD34+ cells from mobilized peripheral blood or cord blood. | Miltenyi Biotec CD34 MicroBead Kit, Stemcell Technologies EasySep |
| Electroporation System & Buffer | Enables efficient, non-viral delivery of RNP complexes into sensitive HSPCs. | Lonza P3 Primary Cell 4D-Nucleofector X Kit, MaxCyte Electroporation Buffer |
| StemSpan SFEM II Media | Serum-free, cytokine-supplemented medium optimized for ex vivo HSPC culture and maintenance. | Stemcell Technologies (Catalog #09605) |
| Recombinant Cytokines (SCF, TPO, FLT3L) | Supports HSPC survival, prevents differentiation, and enables limited expansion during ex vivo culture. | PeproTech, R&D Systems |
| NGS Amplicon Sequencing Kit | For quantifying on-target editing efficiency (INDEL%) and assessing potential off-target sites. | Illumina MiSeq, IDT xGen Amplicon Panel |
| SCD Disease Model (e.g., Townes Mouse) | In vivo model expressing human HbS for preclinical efficacy and safety testing. | The Jackson Laboratory (Stock #013071) |
| Erythroid Differentiation Kit | In vitro assay to differentiate edited HSPCs into erythroid lineage and measure HbF expression (HPLC/Flow). | Stemcell Technologies Erythroid Expansion Kit |
Conclusion
CRISPR-Cas9 gene editing represents a paradigm shift in the potential cure for sickle cell disease, moving from symptom management to definitive genetic correction. Foundational understanding of the HBB/BCL11A targets enables precise therapeutic strategies, while methodological advances in HSC editing are yielding robust clinical results. However, challenges in optimization, such as off-target effects and engraftment efficiency, require continued innovation. Validation through clinical trials demonstrates remarkable efficacy, positioning CRISPR-based therapies at the forefront of the SCD treatment landscape. Future directions will focus on enhancing safety profiles, developing accessible in vivo delivery methods, and expanding this powerful platform to other monogenic hematologic disorders, solidifying gene editing's role in next-generation biomedicine.