Harnessing Cas12a for T-Rich Targets: A Comprehensive Protocol Guide for TTTN PAM Applications in Genome Editing
This article provides a detailed, step-by-step protocol for utilizing the CRISPR-Cas12a (Cpf1) system to target thymine (T)-rich genomic regions through its unique TTTN (and TTTV) PAM specificity.
Harnessing Cas12a for T-Rich Targets: A Comprehensive Protocol Guide for TTTN PAM Applications in Genome Editing
Abstract
This article provides a detailed, step-by-step protocol for utilizing the CRISPR-Cas12a (Cpf1) system to target thymine (T)-rich genomic regions through its unique TTTN (and TTTV) PAM specificity. Tailored for researchers, scientists, and drug development professionals, it covers the foundational biology of Cas12a, a complete methodological workflow from guide RNA design to delivery and analysis, expert troubleshooting for common efficiency and specificity challenges, and a comparative analysis with Cas9 systems. The guide synthesizes current best practices and validation strategies to enable precise editing in previously inaccessible genomic territories, offering significant implications for functional genomics, target validation, and therapeutic development.
Understanding Cas12a: Why Its TTTN PAM is a Game-Changer for T-Rich Genomic Regions
CRISPR-Cas12a, also known as Cpf1, represents a distinct class of Type V CRISPR-Cas systems that has expanded the genome-editing toolkit beyond the widely adopted Cas9. Its unique properties are particularly advantageous for targeting T-rich genomic regions, given its preference for a TTTN Protospacer Adjacent Motif (PAM). This Application Note details the structural and functional characteristics of Cas12a, contrasts it with Cas9, and provides protocols for its application in research, framed within a thesis context focusing on TTTN PAM exploration.
Key Structural and Functional Differences: Cas12a vs. Cas9
Cas12a differs from Cas9 in multiple fundamental aspects, from PAM recognition to nuclease activity. The table below summarizes the core distinctions.
Table 1: Quantitative Comparison of Cas9 (SpCas9) and Cas12a (LbCas12a/AsCas12a)
| Feature | Cas9 (e.g., SpCas9) | Cas12a (e.g., LbCas12a) |
|---|---|---|
| System Class | Type II | Type V |
| Effector Complex | Dual RNA (crRNA + tracrRNA) or sgRNA | Single crRNA (no tracrRNA required) |
| crRNA Length | ~100 nt | ~42-44 nt |
| PAM Sequence | 5'-NGG-3' (SpCas9, downstream) | 5'-TTTN-3' (LbCas12a, upstream) |
| PAM Location | 3' of protospacer (downstream) | 5' of protospacer (upstream) |
| Nuclease Domains | HNH (cuts target strand), RuvC (cuts non-target) | Single RuvC-like domain (cuts both strands) |
| Cleavage Type | Blunt-ended double-strand breaks | Staggered double-strand breaks with 5' overhangs |
| Cleavage Position | Within seed region, 3 bp upstream of PAM | Distal from PAM, creating 5-8 nt overhangs |
| Trans-cleavage Activity | No | Yes (non-specific ssDNase activity upon target binding) |
Application Notes for Targeting T-rich Genomic Regions
The TTTN PAM preference of Cas12a makes it the nuclease of choice for regions of the genome that are Adenine-Thymine (AT)-rich and less accessible to Cas9. This is critical for:
- Targeting specific AT-rich pathogen genomes (e.g., Plasmodium spp., certain viruses).
- Editing genetic elements located in genomic "deserts" with high AT-content.
- Expanding the total addressable genome space when used in combination with Cas9 systems.
Detailed Experimental Protocols
Protocol 3.1: Designing and Validating crRNAs for a TTTN PAM Target
Objective: To design and functionally test crRNAs for specific genomic loci preceded by a TTTN PAM.
Materials: See "The Scientist's Toolkit" section. Procedure:
- Genomic Locus Analysis: Using reference genome files (e.g., hg38), scan 5'-TTTV-3' (V = A, C, or G) sequences upstream of your target exon/noncoding region. Confirm locus uniqueness via BLAST.
- crRNA Design: The 20-24 nt spacer sequence is designed to be complementary to the DNA strand directly following the PAM. Synthesize the full 42-44 nt crRNA (direct repeat + spacer).
- In Silico Off-target Prediction: Use tools like Cas-OFFinder. Input "TTTV" as PAM and allow up to 3-4 mismatches, prioritizing seed region near PAM.
- In Vitro Cleavage Assay (Validation): a. Amplify a ~500-800 bp genomic DNA fragment containing the target site with PAM. b. Assemble 50 µL reaction: 100 ng PCR product, 50 nM purified Cas12a enzyme, 100 nM crRNA, 1X NEBuffer r3.1. c. Incubate at 37°C for 60 min, then heat-inactivate at 80°C for 10 min. d. Analyze products on a 2% agarose gel. Successful cleavage yields two smaller, distinct bands.
Protocol 3.2: Mammalian Cell Transfection and Editing Analysis
Objective: To induce targeted mutations in mammalian cells using Cas12a RNP.
Materials: See "The Scientist's Toolkit" section. Procedure:
- RNP Complex Formation: For one nucleofection reaction, mix 3 µg (≈ 50 pmol) of recombinant Cas12a protein with 6 µL of 10 µM synthetic crRNA (60 pmol) in duplex buffer. Incubate at 25°C for 10 min.
- Cell Preparation: Harvest 2.0 x 10^5 HEK293T or other target cells by trypsinization. Wash once with PBS.
- Nucleofection: Resuspend cell pellet in 20 µL of P3 Primary Cell Nucleofector Solution (Lonza). Mix with the pre-formed RNP complex. Transfer to a nucleofection cuvette and run the appropriate program (e.g., CM-130 for HEK293T). Immediately add 80 µL pre-warmed medium post-pulse.
- Cell Culture & Expansion: Transfer cells to a 24-well plate with 500 µL complete medium. Culture for 72-96 hours.
- Genomic DNA Extraction & Analysis: a. Extract gDNA using a silica-column kit. b. PCR amplify the target locus (amplicon size: 300-500 bp). c. Quantify editing efficiency via T7 Endonuclease I (T7E1) assay or, preferably, by next-generation sequencing (NGS). For NGS, prepare amplicon libraries and sequence on an Illumina MiSeq. Analyze indels using CRISPResso2.
The Scientist's Toolkit
Table 2: Essential Research Reagent Solutions for Cas12a Experiments
| Item | Function/Description | Example Vendor/Cat. No. (Representative) |
|---|---|---|
| Recombinant Cas12a Nuclease | Purified protein for RNP formation or in vitro assays. High specificity and activity are critical. | IDT (Alt-R LbCas12a), NEB (LbCpf1) |
| Synthetic crRNAs | Custom 42-44 nt RNA oligos with direct repeat and spacer sequence. Chemical modifications enhance stability. | IDT (Alt-R crRNA), Synthego |
| Nucleofection System | Electroporation-based delivery method for high-efficiency RNP introduction into hard-to-transfect cells. | Lonza (4D-Nucleofector), Bio-Rad (Neon) |
| NGS Library Prep Kit | For deep sequencing of target amplicons to quantify editing efficiency and profile indel spectra. | Illumina (Nextera XT), Swift Biosciences |
| Cell Line with TTTN-Rich Target | A relevant mammalian, plant, or microbial cell line containing the desired AT-rich genomic target. | ATCC, Coriell Institute |
| Cas12a-Specific Buffer | Optimized reaction buffer supporting Cas12a's enzymatic activity and stability. | NEBuffer r3.1 (NEB) |
| Genomic DNA Extraction Kit | Reliable, high-yield gDNA isolation from edited cell populations for downstream analysis. | Qiagen (DNeasy), Zymo Research |
The Protospacer Adjacent Motif (PAM) is a critical short DNA sequence required for the recognition and cleavage of target DNA by CRISPR-Cas systems. Unlike the G-rich PAM of Cas9, Cas12a (formerly Cpf1) recognizes a 5' T-rich PAM, most commonly TTTN (where N is A, T, or C; V denotes A, C, or G), enabling targeting of T-rich genomic regions inaccessible to Cas9. This application note details the significance of this motif and provides protocols for leveraging Cas12a in research and therapeutic development.
Quantitative Data on Cas12a PAM Specificity
Table 1: Experimentally Validated Cas12a Ortholog PAM Preferences
| Cas12a Ortholog | Primary PAM (5'→3') | Efficiency | Notes/Source |
|---|---|---|---|
| Acidaminococcus sp. (AsCas12a) | TTTV | High | Most commonly used; high efficiency for TTTV. |
| Lachnospiraceae bacterium (LbCas12a) | TTTV | High | Similar to AsCas12a; robust activity. |
| Francisella novicida (FnCas12a) | TTYN (Y=C or T) | Moderate | Broader PAM, includes TTTN, TTCN. |
| Engineered AsCas12a (AsCas12a-RR) | TYCV / TATV | Broad | Evolved variant with relaxed PAM specificity. |
| Engineered LbCas12a (LbCas12a-RVR) | TNTN | Broad | Evolved variant; significantly expanded target range. |
Table 2: Comparison of CRISPR-Cas Nuclease PAM Requirements
| Nuclease | PAM Sequence (5'→3') | PAM Location | Genomic Target Implications |
|---|---|---|---|
| SpCas9 | NGG | 3' of guide | Targets G-rich regions. |
| Cas12a (Wild-Type) | TTTV | 5' of guide | Targets T-rich regions; useful for AT-rich genomes. |
| Engineered SpCas9 (VQR, etc.) | NGAN / NGAG | 3' of guide | Altered specificity, but often remains G-rich. |
| Engineered Cas12a (RR, RVR) | TYCV / TNTN | 5' of guide | Broadened access within T-rich contexts. |
Experimental Protocol: Targeting a T-rich Genomic Locus with Wild-Type LbCas12a
A. Design and Cloning of Cas12a crRNA Expression Construct
Objective: To clone a specific crRNA expression cassette targeting a human genomic locus preceded by a TTTV PAM.
Materials:
- Target genomic DNA sequence data.
- crRNA design tool (e.g., Benchling, IDT).
- Plasmid backbone (e.g., pY010, Addgene #69982) containing a U6 promoter and direct repeat sequence.
- Oligonucleotides for the spacer sequence.
- Restriction enzymes (BsaI) or Gibson Assembly/ Golden Gate Assembly mix.
- Competent E. coli.
Procedure:
- Identify Target Site: Scan the target genomic region for the canonical PAM (TTTV) on the 5' end of the target strand. Select a 20-24 nt spacer sequence immediately 3' to the PAM. Verify specificity via BLAST.
- Design Oligos: Design forward and reverse oligonucleotides that, when annealed, generate overhangs compatible with the BsaI-digested vector. The oligos encode the spacer sequence. Forward Oligo: 5'-AAAC-[20-24nt Spacer Sequence]-3' Reverse Oligo: 5'-AAA-[Reverse Complement of Spacer]-3'
- Anneal Oligos: Resuspend oligos, mix equimolar amounts, anneal in a thermal cycler (95°C for 2 min, ramp to 25°C at 0.1°C/sec).
- Golden Gate Assembly: Digest the destination vector with BsaI. Perform a Golden Gate assembly reaction with the annealed oligo duplex, digested vector, and T4 DNA ligase. Cycle between 37°C (digestion) and 16°C (ligation) for 20-30 cycles.
- Transform and Verify: Transform competent E. coli, isolate plasmid DNA, and verify the insert by Sanger sequencing.
B. Mammalian Cell Transfection and Cleavage Assay
Objective: To assess Cas12a-mediated indel formation at the target TTTN locus.
Materials:
- HEK293T or other relevant cell line.
- LbCas12a expression plasmid (e.g., pY016, Addgene #69976).
- Validated crRNA expression plasmid (from Part A).
- Transfection reagent (e.g., Lipofectamine 3000).
- Genomic DNA extraction kit.
- PCR primers flanking the target site (150-300 bp amplicon).
- T7 Endonuclease I or Surveyor nuclease. OR materials for Next-Generation Sequencing (NGS) analysis.
Procedure:
- Cell Seeding: Seed HEK293T cells in a 24-well plate to reach 70-80% confluency at transfection.
- Transfection: Co-transfect cells with 500 ng LbCas12a plasmid and 250 ng crRNA plasmid using the transfection reagent per manufacturer's protocol. Include a "no nuclease" control.
- Harvest Genomic DNA: 72 hours post-transfection, harvest cells and extract genomic DNA.
- PCR Amplification: Amplify the target locus from 100-200 ng of genomic DNA.
- Cleavage Assay: a. T7E1/Surveyor Assay: Denature and re-anneal the PCR products to form heteroduplex DNA if indels are present. Digest with T7E1 enzyme. Analyze fragments on a 2% agarose gel. Calculate indel efficiency from band intensities. b. NGS Analysis (Gold Standard): Purify PCR amplicons, prepare sequencing libraries, and perform high-depth paired-end sequencing (e.g., Illumina MiSeq). Analyze reads using CRISPResso2 or similar to quantify precise indel percentages and spectra.
- Data Analysis: Confirm that efficient cleavage correlates with the presence of the correct TTTV PAM.
Visualizing the Cas12a Mechanism and Workflow
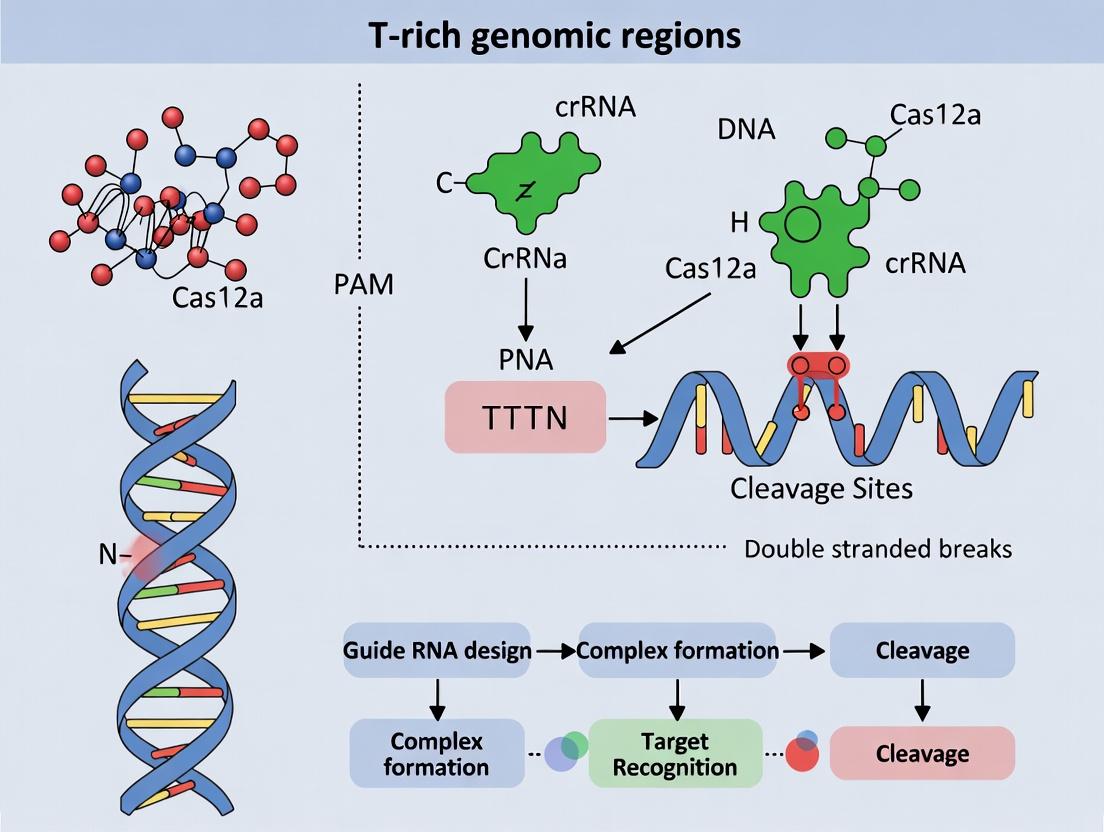
Diagram 1 Title: Cas12a Target Recognition & Cleavage
Diagram 2 Title: Workflow: Targeting T-rich Regions with Cas12a
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for Cas12a TTTN PAM Research
| Item | Function/Description | Example Product/Catalog # |
|---|---|---|
| Wild-Type Cas12a Expression Plasmid | Mammalian codon-optimized vector for expressing LbCas12a or AsCas12a protein. | pY016 (LbCas12a, Addgene #69976) |
| crRNA Cloning Backbone | Plasmid with U6 promoter and direct repeat for easy insertion of spacer sequences. | pY010 (Addgene #69982) |
| Engineered Cas12a Variants | Plasmids encoding PAM-relaxed mutants for broader targeting within T-rich regions. | LbCas12a-RVR (Addgene #137695) |
| Ready-to-Transfect RNP | Purified recombinant Cas12a protein pre-complexed with synthetic crRNA for rapid, vector-free editing. | IDT Alt-R Cas12a (Cpf1) Ultra |
| High-Fidelity PCR Mix | For accurate amplification of genomic target loci from transfected cells prior to indel analysis. | NEB Q5 High-Fidelity 2X Master Mix |
| T7 Endonuclease I | Enzyme for detecting indels via mismatch cleavage in heteroduplex DNA (cost-effective screening). | NEB T7 Endonuclease I (M0302) |
| NGS Library Prep Kit | For preparing amplicon libraries from target sites to quantify editing efficiency and profiles. | Illumina DNA Prep Kit |
| crRNA Design Software | In-silico tools for identifying TTTV PAM sites and predicting off-targets. | Benchling, CHOPCHOP, IDT Design Tool |
The CRISPR-Cas9 system, predominantly reliant on NGG and related PAMs, has left large swaths of the genome, particularly AT-rich regions, under-targeted. These regions are frequently gene-sparse but harbor critical regulatory elements and are implicated in various diseases. This application note is framed within a broader thesis positing that the CRISPR-Cas12a (Cpf1) system, with its canonical TTTN (and related T-rich) PAM requirement, is uniquely suited for interrogating these neglected genomic territories. We detail protocols and data demonstrating Cas12a's strategic advantage in expanding the targetable genome.
Table 1: Comparative PAM and Genomic Coverage of Major CRISPR Systems
| CRISPR System | Canonical PAM | Approx. PAM Frequency (per kb in Human Genome)* | Primary Nuclease Activity | Targetable AT-Rich Regions |
|---|---|---|---|---|
| SpCas9 | NGG | ~1 in 16 bp | Blunt DSB | Limited |
| SpCas9 VQR Variant | NGAN / NGNG | ~1 in 8 bp | Blunt DSB | Moderate |
| AsCas12a (LbCas12a) | TTTV (V=A/C/G) | ~1 in 8 bp | Staggered DSB (5' overhangs) | High |
| LbCas12a RR Variant | TYCV (Y=C/T) | ~1 in 4 bp | Staggered DSB (5' overhangs) | Very High |
*Frequency estimates based on human genome (hg38) sequence composition analysis.
Table 2: Experimental Efficiency in AT-Rich Model Loci
| Target Locus (GC%) | CRISPR System | PAM Sequence | Cleavage Efficiency (NGS, %) | HDR Efficiency (with donor, %) | Notes |
|---|---|---|---|---|---|
| Intergenic AT-Rich (25% GC) | AsCas12a | TTTG | 78.2 ± 5.1 | 22.4 ± 3.2 | High indels, clean HDR |
| Intergenic AT-Rich (25% GC) | SpCas9 | AGG | 15.3 ± 4.7 | <2.0 | Poor sgRNA design/activity |
| Gene Desert Region (30% GC) | LbCas12a-RR | TTCG | 65.8 ± 6.3 | 18.9 ± 2.8 | Robust activity |
| Promoter (NF-κB site, 28% GC) | AsCas12a | TTTA | 81.5 ± 3.9 | N/A | Transcriptional modulation study |
Detailed Experimental Protocols
Protocol 1: Design and Cloning of Cas12a crRNA Arrays for Multiplexing Objective: To clone a single expression construct encoding multiple crRNAs targeting an extended AT-rich genomic region.
- Design: Identify TTTN (or engineered variant-compatible) PAMs in the target region. Design 20-24 bp protospacers directly 5' to the PAM. Avoid poly-T stretches (>4) within the spacer.
- Oligo Annealing: Synthesize DNA oligonucleotides: Forward: 5'-AAAT-[Spacer]-3'; Reverse: 5'-[Spacer RevComp]-3'. Anneal in NEBuffer 2.1 by heating to 95°C for 5 min and ramping down to 25°C at 0.1°C/sec.
- Golden Gate Assembly: Dilute annealed duplex 1:100. Set up a 20 µL reaction: 50 ng BsaI-digested Cas12a crRNA array backbone (e.g., pY016), 1 µL diluted duplex (for each spacer), 1 µL T4 DNA Ligase, 2 µL 10x T4 Ligase Buffer, 1 µL BsaI-HFv2. Cycle: 37°C (5 min) + 20°C (5 min), 30 cycles; then 50°C (5 min), 80°C (5 min).
- Transformation & Validation: Transform 5 µL reaction into competent E. coli. Isolate plasmid and validate by Sanger sequencing using an array-flanking primer.
Protocol 2: Delivery and Analysis in Mammalian Cells Objective: To assess editing efficiency at an AT-rich target locus in HEK293T cells.
- Cell Seeding: Seed 1.5e5 HEK293T cells per well in a 24-well plate in DMEM + 10% FBS 24h pre-transfection.
- Transfection: For each well, mix 500 ng AsCas12a or LbCas12a expression plasmid (with NLS) + 250 ng crRNA expression plasmid (or 100 ng of each plasmid for RNP delivery) in 50 µL Opti-MEM. Add 1.5 µL Lipofectamine 3000 in a separate 50 µL Opti-MEM. Combine, incubate 15 min, add dropwise to cells.
- Genomic DNA Extraction: 72h post-transfection, harvest cells using trypsin. Extract gDNA using a silica-column based kit (e.g., DNeasy Blood & Tissue). Elute in 50 µL nuclease-free water. Quantify via spectrophotometry.
- Analysis by T7 Endonuclease I (T7EI) Assay:
- PCR: Amplify a ~500-800 bp region surrounding the target site using high-fidelity polymerase.
- Heteroduplex Formation: Denature/reanneal PCR product: 95°C 5 min, ramp to 85°C at -2°C/sec, then to 25°C at -0.1°C/sec.
- Digestion: Digest with T7EI (NEB) at 37°C for 30 min. Analyze fragments on a 2% agarose gel. Calculate indel % = 100 × (1 - sqrt(1 - (b+c)/(a+b+c))), where a is integrated band intensity of undigested product, b and c are cleavage products.
- Analysis by Next-Generation Sequencing (NGS):
- Amplify target locus with barcoded primers.
- Purify and pool amplicons for sequencing on an Illumina MiSeq.
- Analyze reads using CRISPResso2 to quantify precise indel spectra and frequencies.
Signaling Pathways & Workflow Diagrams
Title: Workflow for Targeting AT-Rich Regions with Cas12a
Title: DNA Repair Pathways Post-Cas12a Cleavage
The Scientist's Toolkit: Research Reagent Solutions
| Item/Category | Function & Strategic Relevance for AT-Rich Targeting |
|---|---|
| LbCas12a (Cpf1) Expression Plasmid | Codon-optimized mammalian expression vector with nuclear localization signals (NLS). Essential for delivering the nuclease. |
| Engineered Cas12a Variants (e.g., RR) | Recognize TYCV PAMs, dramatically increasing targetable sites in AT-rich regions compared to wild-type TTTV. |
| crRNA Cloning Backbone (e.g., pY016) | Vector with direct repeats for BsaI-mediated Golden Gate assembly of single or arrayed crRNA sequences. |
| Chemically Synthesized crRNA | For RNP delivery; higher precision and reduced off-target effects. Critical for sensitive applications. |
| AT-Rich-Compatible Polymerase (e.g., Q5 Hot Start) | High-fidelity PCR enzyme engineered to robustly amplify GC-poor, AT-rich genomic regions for analysis. |
| T7 Endonuclease I (T7EI) | Mismatch-specific endonuclease for rapid, cost-effective quantification of indel efficiency from PCR products. |
| Silica-Column gDNA Kit | Reliable, RNase-free genomic DNA extraction for downstream PCR and sequencing analysis. |
| Illumina-Compatible Barcoded Primers | For multiplexed NGS amplicon sequencing of multiple target loci across samples to obtain quantitative indel data. |
| CRISPResso2 Software | Bioinformatics tool for precise quantification of genome editing outcomes from NGS data. |
The CRISPR-Cas12a system, recognizing a T-rich Protospacer Adjacent Motif (PAM) sequence (TTTV, where V is A, C, or G), offers a unique advantage for interrogating genomic regions with inherent T-nucleotide bias. This application note details the relevance and protocols for leveraging Cas12a to study and target biologically critical regions—disease-associated loci, gene promoters, and repetitive elements—that are statistically enriched in T/A nucleotides. This work is framed within a broader thesis developing optimized Cas12a protocols for functional genomics and therapeutic targeting in these regions.
Biological Relevance: T/Nucleotide-Enriched Genomic Features
Recent genomic analyses reveal a non-random distribution of nucleotides, with specific functional elements showing significant T/A enrichment.
Table 1: Quantitative Enrichment of T/A Nucleotides in Key Genomic Features
| Genomic Feature | Example / Class | Observed T/A% | Genomic Background T/A% | Enrichment (Fold) | Biological/Clinical Implication |
|---|---|---|---|---|---|
| Disease-Associated Loci | Autoimmune (e.g., MHC locus) | ~68-72% | ~50% | 1.36-1.44 | High density of cis-regulatory elements; Cas12a accessibility for mechanistic studies. |
| Gene Promoters | CpG Island-containing Promoters | ~60-65% | ~50% | 1.20-1.30 | TATA-less promoters; A/T-rich initiator (Inr) and downstream promoter elements (DPEs). |
| Repetitive Elements | Alu SINEs | ~58% | ~50% | 1.16 | Retrotransposition, genome instability; targeting may modulate their expression. |
| Repetitive Elements | LINE-1 5' UTR | ~62% | ~50% | 1.24 | Contains an internal promoter; Cas12a-mediated repression may silence mobilization. |
| Nucleosome-Depleted Regions | Transcription Start Sites | ~65-70% | ~50% | 1.30-1.40 | Facilitates transcription factor binding; prime for Cas12a R-loop formation. |
Application Notes & Protocols
The following protocols are designed for research utilizing the Lachnospiraceae bacterium Cas12a (LbCas12a) or its engineered variants.
Protocol 1: In Silico Identification and gRNA Design for T-Rich Targets
Objective: To design highly specific and efficient Cas12a gRNAs for T/A-enriched genomic regions. Procedure:
- Locus Identification: Input your genomic region of interest (e.g., a GWAS-hit interval) into a genome browser (e.g., UCSC).
- PAM Scanning: Use the
regexpatternTTT[ACG]and its reverse complement[CGT]AAAto scan both strands for candidate PAM sites. Custom scripts or tools like CRISPRseek can automate this. - gRNA Selection:
- Spacer Sequence (20-24 nt): Select the 20-24 nucleotides directly upstream of the identified PAM.
- Specificity Check: Perform a BLAST search against the relevant genome to ensure minimal off-target matches, especially in other T-rich repetitive regions.
- Efficiency Predictors: Score gRNAs using LbCas12a-specific algorithms (if available) or prioritize those with a GC content of 40-60%.
- Oligo Synthesis: Order the spacer sequence with appropriate overhangs for your chosen cloning method (e.g., Golden Gate assembly).
Diagram: Workflow for Cas12a gRNA Design & Validation
Title: Cas12a gRNA Design and Testing Workflow
Protocol 2: Cas12a-Mediated Gene Repression at T-Rich Promoters
Objective: To knock down gene expression by targeting Cas12a (nuclease-dead, dCas12a) fused to a repressor domain (e.g., KRAB) to T-rich promoter elements. Procedure:
- Construct Assembly: Clone validated gRNAs (from Protocol 1) into a dCas12a-KRAB expression plasmid.
- Cell Transfection: Deliver the dCas12a-KRAB and gRNA plasmid(s) into your target cell line (e.g., HEK293T) using a suitable method (lipofection, electroporation).
- Analysis (48-72h post-transfection):
- qRT-PCR: Measure mRNA levels of the target gene versus non-targeting controls.
- Reporter Assay: If applicable, co-transfect a luciferase reporter driven by the target promoter.
- Control: Include a non-targeting gRNA and a gRNA targeting a non-regulatory region (e.g., exon) of the same gene.
Diagram: dCas12a-KRAB Repression at a T-Rich Promoter
Title: dCas12a-KRAB Mechanism at T-Rich Promoter
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for Cas12a Targeting of T-Rich Regions
| Item | Function/Benefit | Example/Supplier |
|---|---|---|
| LbCas12a or AsCas12a Nuclease | The core enzyme recognizing TTTN PAM; preferred for targeting T-rich regions. | IDT (Alt-R S.p. Cas12a), Thermo Fisher (TrueCut Cas12a). |
| High-Fidelity DNA Assembly Master Mix | For efficient, seamless cloning of gRNA spacers into expression vectors. | NEB Golden Gate Assembly Mix, Takara In-Fusion Snap Assembly. |
| Validated dCas12a-KRAB Plasmid | Enables transcriptional repression studies at promoters. | Addgene (Plasmid #107774). |
| Next-Generation Sequencing Kit | For unbiased assessment of on-target editing and genome-wide off-target profiling. | Illumina Nextera XT, Twist NGS. |
| Surveyor or T7 Endonuclease I | Quick validation of nuclease-induced indels at the target locus. | IDT Surveyor Mutation Detection Kit. |
| Lipid-Based Transfection Reagent | For efficient delivery of RNP or plasmid DNA into mammalian cells. | Lipofectamine CRISPRMAX, Mirus TransIT-X2. |
| Genomic DNA Isolation Kit (Magnetic Beads) | Rapid, PCR-ready gDNA extraction for genotyping edited cell pools. | Mag-Bind Blood & Tissue DNA HDQ Kit. |
This application note, framed within a broader thesis on CRISPR-Cas12a for targeting T-rich genomic regions with TTTN PAM, details the comparative nuclease activity of Cas12a. It elucidates the enzyme's unique dual RNase and DNase functions, its cis and trans cleavage mechanisms, and the resultant product profiles. We provide protocols for in vitro cleavage assays and quantitative analysis of products, crucial for researchers designing precise genomic interventions in T-rich areas.
Cas12a (formerly Cpf1) is a Class 2, Type V CRISPR-associated nuclease distinct from Cas9. Its hallmark is the recognition of a T-rich protospacer adjacent motif (PAM: 5'-TTTV-3'), making it ideal for targeting AT-rich genomic regions. Upon crRNA-guided recognition of a target DNA sequence, Cas12a exhibits two nuclease activities: a single RNase event to process its own crRNA array, and robust DNase activity. The DNase activity manifests as: 1) cleavage of the target DNA strand (cis-cleavage) and 2) promiscuous, non-specific cleavage of any single-stranded DNA (trans-cleavage or collateral cleavage). This collateral activity has been leveraged for diagnostic tools like DETECTR.
Comparative Cleavage Mechanisms and Product Profiles
Cas12a generates DNA ends distinct from Cas9, which has profound implications for downstream repair pathways and editing outcomes.
1Cis-Cleavage (Target DNA Cleavage)
Cas12a introduces a staggered double-strand break (DSB) with a 5' overhang. The enzyme contains a single RuvC-like nuclease domain responsible for cutting both strands of the target DNA.
Key Characteristics:
- Cut Site: The enzyme cuts the target strand (complementary to the crRNA spacer) 18-23 nucleotides downstream of the PAM. It cuts the non-target strand 21-25 nucleotides downstream of the PAM.
- Product Ends: This creates a staggered DSB with a 5' overhang, typically 4-5 nucleotides in length.
- Comparison to Cas9: Cas9 uses two nuclease domains (HNH and RuvC) to generate a blunt-ended DSB.
Table 1: Comparison of Cis-Cleavage Products: Cas12a vs. Cas9
| Feature | Cas12a | Cas9 |
|---|---|---|
| PAM Sequence | 5'-TTTV-3' (T-rich) | 5'-NGG-3' (G-rich) |
| Cleavage Domain | Single RuvC-like domain | Dual domains (HNH & RuvC) |
| Cut Site Relative to PAM | Target strand: 18-23 nt downstream; Non-target: 21-25 nt downstream | Target strand: 3 nt upstream; Non-target: 3-8 nt upstream |
| DSB End Structure | Staggered cut with 5' overhang (4-5 nt) | Blunt-ended cut |
| Predominant Repair Pathway | More prone to Microhomology-Mediated End Joining (MMEJ) | Primarily Non-Homologous End Joining (NHEJ) |
2Trans-Cleavage (Collateral Cleavage)
A landmark feature of Cas12a is its indiscriminate single-stranded DNAse activity triggered upon formation of the ternary complex (Cas12a-crRNA-target DNA). This activity relentlessly cleaves any available ssDNA molecules.
Key Characteristics:
- Trigger: Absolutely dependent on successful target DNA binding and cis-cleavage.
- Substrate: Primarily single-stranded DNA (ssDNA). It does not cleave double-stranded DNA or RNA in trans.
- Kinetics: The trans-cleavage activity is sustained and catalytic, leading to signal amplification.
- Product: Short, degraded ssDNA fragments.
Table 2: Quantitative Analysis of Cas12a Trans-Cleavage Activity
| Parameter | Typical Value/Characteristic | Measurement Method |
|---|---|---|
| Activation Time | <5 minutes post-target recognition | Fluorescence-quenched reporter assay |
| Cleavage Rate (k_cat) | ~1250 turnovers per second (for LbCas12a) | Real-time fluorescence kinetics |
| Optimal Temperature | 37°C (for As and Lb variants) | Titrated activity assay |
| Divalent Cation Requirement | Requires Mg²⁺ (1-10 mM) | EDTA chelation experiment |
| Inhibition by EDTA | Complete inhibition | Activity assay with/without EDTA |
Detailed Protocols
Protocol 1: In VitroCis-Cleavage Assay to Analyze Staggered End Generation
Purpose: To verify target DNA cleavage and generate products for end-structure analysis.
Materials:
- Purified Cas12a protein (e.g., AsCas12a or LbCas12a)
- Synthetic crRNA targeting a TTTN PAM sequence
- Target dsDNA substrate (PCR-amplified or plasmid)
- NEBuffer r3.1 or equivalent (with Mg²⁺)
- RNase-free water
- Heating block or thermal cycler
Procedure:
- Complex Formation: In a 1.5 mL tube, combine 50 nM Cas12a protein and 75 nM crRNA in 1X reaction buffer. Incubate at 25°C for 10 minutes.
- Cleavage Reaction: Add the target dsDNA substrate to a final concentration of 10 nM. Bring the total reaction volume to 20 µL with buffer and water.
- Incubation: Incubate the reaction at 37°C for 60 minutes.
- Termination: Stop the reaction by adding 2 µL of Proteinase K (10 mg/mL) and incubating at 56°C for 15 minutes.
- Analysis: Run the entire product on a 2% agarose gel stained with ethidium bromide. Include uncut DNA and a size ladder. Cleavage products will appear as two smaller bands (from the staggered cut) compared to the uncleaved supercoiled/linear substrate.
Protocol 2: Fluorescence-BasedTrans-Cleavage Kinetics Assay
Purpose: To quantitatively measure collateral cleavage activity, useful for characterizing enzyme mutants or screening for activators.
Materials:
- Activated Cas12a-crRNA complex (from Protocol 1, step 1)
- Target DNA activator (dsDNA oligonucleotide matching the crRNA)
- Fluorescent ssDNA reporter (e.g., 5'-FAM-TTATT-3IABkFQ-3' or similar)
- Black 96-well plate or qPCR tubes
- Real-time PCR instrument or fluorescence plate reader.
Procedure:
- Reporter Mixture: Dilute the fluorescent ssDNA reporter to 1 µM in the reaction buffer.
- Plate Setup: In each well, combine: 10 µL of Cas12a-crRNA complex (final 10 nM), 2 µL of target DNA activator (final 5 nM), and 28 µL of buffer. Set up negative controls without activator.
- Initiation: Immediately before reading, add 10 µL of the diluted reporter (final 200 nM) to each well.
- Kinetic Measurement: Immediately place the plate in a pre-warmed (37°C) fluorescence reader. Measure fluorescence (Ex: 485 nm, Em: 535 nm) every 30 seconds for 60-90 minutes.
- Data Analysis: Plot relative fluorescence units (RFU) vs. time. The slope of the initial linear phase is proportional to the trans-cleavage rate.
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Cas12a Cleavage Studies
| Reagent | Function & Rationale |
|---|---|
| High-Purity LbCas12a Protein | The L. bacterium variant offers high in vitro activity and specificity; purity reduces non-specific nuclease contamination. |
| Synthetic, HPLC-purified crRNA | Ensures precise guide sequence and high complex formation efficiency; critical for reproducible PAM targeting. |
| Fluorescent-Quenched ssDNA Reporters (e.g., FAM/Iowa Black FQ) | Enables real-time, quantitative measurement of trans-cleavage activity for kinetics and diagnostic applications. |
| Target DNA Activators (dsDNA oligos) | Triggers the conformational change in Cas12a to activate both cis and trans cleavage; essential for assay control. |
| NEBuffer r3.1 | Optimized buffer providing Mg²⁺ and pH stability for maximum Cas12a nuclease activity. |
| Proteinase K | Effectively digests the Cas12a protein to terminate reactions and prevent post-assay degradation of DNA products. |
| Solid-Phase Reversible Immobilization (SPRI) Beads | For clean-up and size-selection of cleavage products prior to sequencing or cloning. |
Visualizations
Diagram Title: Cas12a Activation and Dual Cleavage Pathways
Diagram Title: Cas12a Cis-Cleavage Staggered Cut Mechanism
Diagram Title: Trans-Cleavage Kinetic Assay Workflow
Step-by-Step Cas12a Protocol: From gRNA Design to Efficient Editing in Your Target Cells
This Application Note details the initial phase of a comprehensive CRISPR-Cas12a (Cpf1) workflow, specifically tailored for targeting T-rich genomic regions characterized by TTTN (N = A, C, G, T) Protospacer Adjacent Motifs (PAMs). Efficient gene editing or regulation in these regions, often problematic for CRISPR-Cas9 due to its G-rich PAM requirement, necessitates a robust in silico design and cloning pipeline. This protocol provides researchers with a methodical approach to identify optimal target sites and construct corresponding crRNA expression vectors, forming the critical foundation for downstream experimental validation.
Key Research Reagent Solutions
The following materials are essential for executing the protocols described.
| Item | Function/Benefit |
|---|---|
| EnGen Lba Cas12a (Cpf1) | A commonly used, high-fidelity Lachnospiraceae bacterium ND2006 Cas12a nuclease with demonstrated activity with TTTN PAMs. |
| Golden Gate Assembly Kit (BsaI) | Enables rapid, seamless, and directional cloning of crRNA sequences into expression backbones. |
| crRNA Expression Backbone (e.g., pU6-sgRNA) | A mammalian expression plasmid containing a U6 RNA Pol III promoter for driving crRNA expression. |
| Genomic DNA Analysis Software (e.g., SnapGene, Benchling) | For sequence visualization, PAM site identification, and primer design. |
| Off-Target Prediction Tool (Cas-OFFinder) | For genome-wide screening of potential off-target sites for candidate crRNAs. |
| High-Fidelity DNA Polymerase | For accurate PCR amplification of genomic targets and vector components. |
| Chemically Competent E. coli | For transformation and propagation of cloned plasmid constructs. |
In Silico Identification of Optimal TTTN PAM Sites
Protocol
- Sequence Retrieval: Obtain the FASTA format DNA sequence of your target genomic region from a database like NCBI Nucleotide or Ensembl.
- PAM Site Scanning: Using sequence analysis software (e.g., Benchling's CRISPR toolset), scan the +/- strands for all occurrences of the 5'-TTTV-3' PAM (where V = A, C, or G; TTTT is generally less efficient).
- Target Locus Definition: For each PAM, extract the 23-24 nucleotides directly 5' upstream of the PAM sequence. This is the putative spacer sequence for the crRNA.
- Efficiency Scoring: Score each candidate spacer. Prioritize sites with:
- Proximity to the functional domain you intend to disrupt.
- A GC content between 40-70%.
- Avoidance of long stretches of homopolymers (especially T-stretches).
- Low predicted off-target potential (see Step 5).
- Off-Target Analysis: Input each 23-24 nt spacer sequence + TTTN PAM into an off-target prediction tool (e.g., Cas-OFFinder). Allow up to 3-4 mismatches, especially in the PAM-distal region. Exclude candidates with putative off-targets in coding or regulatory regions.
- Final Selection: Select 3-5 top-ranked crRNA target sites for experimental validation.
Data Presentation: Candidate crRNA Sites
The following table summarizes the in silico analysis for a hypothetical target gene (EXAMPLE1).
Table 1: Candidate crRNA Spacer Sequences for EXAMPLE1 Targeting
| ID | Genomic Coordinate (strand) | PAM Sequence (5'->3') | Spacer Sequence (5'->3')* | GC% | Predicted On-Target Efficiency Score | High-Risk Off-Target Count* |
|---|---|---|---|---|---|---|
| C1 | chr1:123,456,789 (+) | TTTG | ATGCTACGATCGATCGATCGTACG | 52.2 | 88 | 0 |
| C2 | chr1:123,456,805 (-) | TTTC | CGATCGTAGCTAGCTACGATCGTAG | 56.5 | 92 | 1 |
| C3 | chr1:123,456,822 (+) | TTTA | AATCGATCGATCGATCGATCGTTA | 39.1 | 75 | 0 |
| C4 | chr1:123,456,845 (-) | TTTG | GCTAGCTAGCTAGCTAGCTAGCTA | 60.9 | 85 | 3 |
*Sequence as it appears in the target genome, 5' of the PAM. Hypothetical score (0-100 scale) from predictive algorithm integration. *Number of predicted off-target sites with ≤3 mismatches in genomic coding regions.
Protocol: Golden Gate Assembly for crRNA Expression Vector Construction
This protocol details the cloning of selected spacer sequences into a BsaI-digested crRNA expression vector.
Materials
- crRNA expression backbone plasmid (with BsaI sites flanking the cloning site).
- Forward and Reverse oligonucleotides for each spacer (see design rules).
- T4 Polynucleotide Kinase (PNK).
- T4 DNA Ligase and 10x Buffer.
- Restriction Enzyme BsaI-HFv2.
- ATP (10 mM).
- Thermocycler.
- Standard reagents for bacterial transformation and colony PCR.
Step-by-Step Methodology
Oligo Design & Annealing:
- For each spacer, design two complementary oligonucleotides (top and bottom strand).
- Top Oligo (Forward): 5' - CACC + [23-24 nt spacer sequence] - 3'
- Bottom Oligo (Reverse): 5' - AAAC + [Reverse complement of spacer] - 3'
- Resuspend oligos to 100 µM. Mix 1 µL of each oligo with 48 µL of nuclease-free water and 5 µL of 10x T4 Ligation Buffer (contains ATP). Anneal in a thermocycler: 37°C for 30 min; 95°C for 5 min, then ramp down to 25°C at 0.1°C/sec. Dilute annealed duplex 1:200 in water.
Golden Gate Assembly Reaction:
- Set up a 20 µL reaction on ice:
- 50 ng BsaI-digested backbone vector
- 1 µL diluted annealed oligo duplex
- 1 µL BsaI-HFv2
- 1 µL T4 DNA Ligase
- 2 µL 10x T4 Ligase Buffer
- Nuclease-free water to 20 µL.
- Run the assembly in a thermocycler: (37°C for 5 min, 20°C for 5 min) x 30 cycles; then 80°C for 5 min; hold at 4°C.
- Set up a 20 µL reaction on ice:
Transformation & Screening:
- Transform 2-5 µL of the assembly reaction into 50 µL of competent E. coli cells via heat shock.
- Plate on LB agar with appropriate antibiotic.
- Pick 3-5 colonies per construct for colony PCR using primers flanking the insertion site.
- Inoculate positive clones for plasmid miniprep and verify the insert by Sanger sequencing.
Workflow & Pathway Visualizations
Title: In Silico crRNA Design & Cloning Workflow
Title: Cas12a-crRNA Binding & DNA Cleavage Mechanism
Within the thesis "Development of a High-Efficiency CRISPR-Cas12a Protocol for Targeting T-Rich Genomic Regions with TTTN PAM," the design of the CRISPR RNA (crRNA) is the foundational determinant of success. Unlike Cas9, Cas12a (e.g., LbCas12a, AsCas12a) utilizes a shorter, direct-RNA guide and recognizes a T-rich Protospacer Adjacent Motif (TTTV, most commonly). This protocol details the best practices for designing highly specific and efficient crRNAs for Cas12a systems, with a focus on applications in functional genomics and therapeutic target validation.
Core Design Parameters: Length and Sequence Selection
crRNA Length: The optimal crRNA length for Cas12a is consistently reported as 20-24 nucleotides for the spacer sequence. A 20-nt spacer is often the default, but extending to 24 nt can enhance specificity in some contexts, particularly in AT-rich regions.
Table 1: Recommended crRNA Spacer Length for Cas12a
| Spacer Length (nt) | Reported Efficiency | Specificity | Recommended Use Case |
|---|---|---|---|
| 18-19 | Moderate to Low | Lower | Preliminary screening, not recommended for final experiments. |
| 20 | High | Good | Standard default for most applications. |
| 21-24 | High | Enhanced | Preferred for sensitive applications (e.g., therapeutic development) or in complex, repetitive genomic regions. |
| >24 | Declining | Variable | Not recommended; may reduce cleavage efficiency. |
Sequence Selection Rules:
- PAM Requirement: Identify a 5'-TTTV (V = A, C, or G) PAM site upstream of your target sequence. The target protospacer is the 20-24 nt sequence directly 3' to the PAM.
- GC Content: Aim for 40%-60% GC content. Avoid extremes (<20% or >80%) to ensure stable crRNA-DNA hybridization.
- Avoid Homopolymer Runs: Sequences with >4 consecutive identical nucleotides (e.g., AAAA, TTTT) can reduce efficiency.
- Target Strand: Cas12a cleaves both strands but primarily targets the PAM-containing strand. Design is straightforward: the crRNA spacer is the reverse complement of the target protospacer.
Specificity Checking and Off-Target Prediction Protocol
Objective: To computationally identify and minimize the risk of crRNA binding and cleaving at unintended genomic loci.
Protocol: A Stepwise In-Silico Analysis
Step 1: Primary Off-Target Search.
- Tool: Use Cas-OFFinder or the CRISPRseek package in R/Bioconductor.
- Input Parameters:
- CRISPR System: Select "Cas12a" or specify "TTTV" PAM.
- Spacer Sequence: Input your 20-24 nt candidate spacer.
- Mismatch Tolerance: Set to 3-5 mismatches for initial broad search.
- Genomic Database: Select the appropriate reference genome (e.g., GRCh38, mm10).
- Output Analysis: Generate a ranked list of potential off-target sites. Prioritize sites with:
- Fewer mismatches (especially in the 5' "seed" region proximal to the PAM, which is more critical for Cas12a than the distal end).
- Mismatches as bulges (DNA/RNA mispairing) are less tolerated than simple base mismatches.
Step 2: Comprehensive Scoring and Ranking.
- Tool: Utilize specialized Cas12a predictors like CHOPCHOP (Cas12a mode) or CRISPR-DT.
- Methodology: These tools integrate mismatch position, type, and genomic context (e.g., chromatin accessibility, exon/intron location) to provide a specificity score.
- Action: If the top candidate crRNA has high-scoring potential off-targets in coding regions, proceed to the next candidate sequence.
Step 3: Final Validation Check.
- Tool: Perform a basic BLASTn search against the target genome.
- Purpose: A quick, final check for perfect or near-perfect matches elsewhere in the genome that may have been missed.
Table 2: Key Off-Target Prediction Tools for Cas12a
| Tool Name | Access | Key Feature for Cas12a | Primary Output |
|---|---|---|---|
| Cas-OFFinder | Web/Standalone | Flexible PAM input (e.g., TTTV) | List of genomic loci with user-defined mismatch tolerance. |
| CHOPCHOP | Web Server | Dedicated Cas12a option; integrates on/off-target scores | Ranked list of crRNAs with visual off-target maps. |
| CRISPRseek | R/Bioconductor | Programmable, batch processing capability | Detailed off-target table with mismatch details. |
| CRISPR-DT | Web Server | Considers DNA thermodynamics & accessibility | On-target and off-target activity scores. |
crRNA Design Workflow Diagram
Title: Cas12a crRNA Design and Specificity Screening Workflow
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Cas12a crRNA Workflow
| Reagent / Material | Function / Purpose | Example Vendor/Type |
|---|---|---|
| Cas12a Nuclease | The effector enzyme (e.g., LbCas12a, AsCas12a). Purified protein for in vitro work or expression plasmid/mRNA for cellular delivery. | IDT, Thermo Fisher, Takara Bio, recombinant expression. |
| crRNA Synthesis Kit | For in vitro transcription (IVT) of custom crRNAs. Essential for screening multiple designs rapidly. | NEB HiScribe T7 Quick High Yield Kit, or similar. |
| Chemically Synthesized crRNA | High-purity, ready-to-use crRNA for critical experiments. Guaranteed sequence fidelity, often with chemical modifications for stability. | IDT Alt-R CRISPR-Cas12a crRNA, Synthego. |
| Target DNA Template | Genomic DNA extract or synthetic double-stranded DNA amplicon containing the target site with PAM. Required for in vitro cleavage validation. | Purified gDNA, gBlocks (IDT). |
| Electrophoresis System | To visualize and confirm cleavage efficiency of target DNA in vitro (gel shift assay). | Agarose gel system, precast polyacrylamide gels. |
| Off-Target Prediction Software | Computational resources for specificity analysis. | CHOPCHOP web server, Cas-OFFinder local install. |
| Cell Line with Target Locus | Appropriate cellular model for final functional validation of crRNA activity and specificity. | e.g., HEK293T, iPSCs, or relevant primary cells. |
| Next-Generation Sequencing (NGS) Library Prep Kit | For comprehensive, genome-wide off-target assessment (e.g., GUIDE-seq, Digenome-seq). | Illumina-compatible kits for targeted sequencing. |
Experimental Protocol:In VitroCleavage Assay for crRNA Validation
Objective: To empirically validate the cleavage efficiency of a designed Cas12a crRNA before proceeding to cell-based experiments.
Materials:
- Purified recombinant Cas12a protein.
- Candidate crRNA (chemically synthesized or in vitro transcribed).
- Target DNA substrate (200-500 bp PCR amplicon containing the target site).
- Nuclease-free water and 10X reaction buffer (usually supplied with enzyme).
- Incubator or thermal cycler (37°C).
- Agarose gel electrophoresis equipment.
Method:
- Reaction Setup: In a 0.2 mL tube, combine:
- 50 ng target DNA amplicon.
- 50 nM Cas12a protein.
- 100 nM crRNA (1:2 molar ratio of Cas12a:crRNA).
- 1X reaction buffer.
- Nuclease-free water to a final volume of 20 µL.
- Negative Control: Prepare an identical reaction without the crRNA.
- Incubation: Mix gently and incubate at 37°C for 60 minutes.
- Reaction Termination: Add 2 µL of Proteinase K (or a suitable stop solution) and incubate at 65°C for 10 minutes to degrade the Cas12a protein.
- Analysis: Load the entire reaction on a 2% agarose gel stained with ethidium bromide or a safe alternative. Run the gel and image.
- Interpretation: Successful cleavage is indicated by the disappearance of the full-length amplicon band and the appearance of two smaller bands (corresponding to the cleavage products). The no-crRNA control should show only the intact band.
This in vitro validation provides a rapid, cost-effective confirmation of crRNA activity, forming a critical checkpoint before resource-intensive cellular experiments within the broader thesis research framework.
Within the context of optimizing CRISPR-Cas12a (Cpfl) for targeting T-rich genomic regions utilizing its TTTN PAM, the choice of delivery modality is a critical determinant of experimental success. Cas12a’s unique attributes—including its single RuvC nuclease domain generating staggered ends, minimal off-target effects, and ability to process its own crRNA array—present specific considerations for delivery into mammalian cells. This application note provides a comparative analysis and detailed protocols for the three primary delivery systems: plasmid DNA, mRNA, and pre-assembled Ribonucleoprotein (RNP) complexes, with a focus on Cas12a applications.
Comparative Analysis of Delivery Systems
The selection of a delivery method involves trade-offs between editing efficiency, speed of action, duration of expression, and risk of unwanted immune responses or genomic integration. The following table summarizes key quantitative and qualitative metrics relevant to Cas12a delivery.
Table 1: Comparison of Cas12a Delivery Modalities for Mammalian Cells
| Parameter | Plasmid DNA | mRNA | RNP Complexes |
|---|---|---|---|
| Time to Nuclease Activity | Slow (24-48 hrs) | Moderate (4-24 hrs) | Very Fast (1-4 hrs) |
| Duration of Activity | Long (days-weeks) | Moderate (days) | Short (hours-days) |
| Editing Efficiency | Variable, can be high | High | High, with minimal variability |
| Risk of Genomic Integration | Higher (random integration) | None | None |
| Immunogenicity | High (TLR9 activation) | Moderate (IFN response) | Lowest |
| Cellular Toxicity | Moderate | Moderate | Low |
| Ease of Use | Simple (standard transfection) | Requires handling delicate RNA | Requires complex assembly |
| Cost | Low | Moderate | High (recombinant protein) |
| Best Suited For | Stable cell line generation, long-term studies | High-efficiency editing in hard-to-transfect cells | Rapid, precise edits; sensitive cells (primary, iPSCs); clinical applications |
Detailed Experimental Protocols
Protocol 1: Plasmid-Based Cas12a Delivery
This protocol is ideal for generating stable Cas12a-expressing cell lines or for long-term functional studies.
- Vector Design: Clone the Cas12a gene (e.g., from Lachnospiraceae bacterium ND2006, LbCas12a) into a mammalian expression plasmid under a strong promoter (e.g., EF1α, CAG). Clone the crRNA expression cassette targeting your TTTN PAM site into a separate plasmid or a single plasmid with the Cas12a gene.
- Cell Seeding: Seed HEK293T or your target mammalian cells in a 24-well plate at 70-80% confluence 24 hours before transfection.
- Transfection: For each well, dilute 500 ng of Cas12a plasmid and 250 ng of crRNA plasmid in 50 µL of serum-free Opt-MEM. Dilute 1.5 µL of a lipid-based transfection reagent (e.g., Lipofectamine 3000) in a separate 50 µL of Opt-MEM. Incubate for 5 minutes at RT.
- Complex Formation: Combine the diluted DNA and transfection reagent. Mix gently and incubate for 15-20 minutes at RT.
- Delivery: Add the 100 µL DNA-lipid complex dropwise to the cells. Gently rock the plate.
- Analysis: Replace media after 6-8 hours. Assess editing efficiency via T7E1 assay or NGS 48-72 hours post-transfection.
Protocol 2: Cas12a mRNA and Synthetic crRNA Delivery
This method offers transient, high-level expression with reduced risk of genomic integration.
- Reagent Preparation: Acquire or synthesize 5'-capped and polyadenylated Cas12a mRNA. Resuspend synthetic crRNA targeting the TTTN site in nuclease-free buffer.
- Cell Seeding: Seed cells as in Protocol 1.
- Transfection Mix: For each well, dilute 500 ng of Cas12a mRNA and 30 pmol of synthetic crRNA in 50 µL of Opt-MEM. In a separate tube, dilute 2 µL of a specialized mRNA transfection reagent (e.g., Lipofectamine MessengerMAX) in 50 µL of Opt-MEM. Incubate 5 minutes.
- Complex Formation: Combine the mRNA/crRNA mix with the diluted reagent. Incubate for 5 minutes at RT.
- Delivery & Analysis: Add complexes to cells. Replace media after 4-6 hours. Analyze editing as early as 24 hours post-transfection.
Protocol 3: Cas12a RNP Complex Delivery (Recommended for TTTN PAM Studies)
This method is optimal for sensitive cells and minimizes off-target effects, crucial for precise T-rich region targeting.
- RNP Assembly: Dilute recombinant LbCas12a protein to 10 µM in 1X Cas12a buffer (20 mM HEPES, 150 mM KCl, pH 7.5). Dilute synthetic crRNA to 20 µM in the same buffer. Mix Cas12a protein and crRNA at a 1:1.2 molar ratio (e.g., 2 µL protein + 2.4 µL crRNA). Incubate at 25°C for 10-20 minutes to form the RNP complex.
- Cell Preparation: Harvest and count your target cells (e.g., primary T cells, iPSCs). Wash and resuspend in appropriate electroporation buffer (e.g., P3 buffer for Neon system, SE buffer for 4D-Nucleofector) at 1 x 10⁶ cells per 10 µL.
- Electroporation: Combine 10 µL of cell suspension with the pre-assembled 4.4 µL RNP mixture. Transfer to a certified electroporation cuvette or strip. Electroporate using a cell-type-specific program (e.g., for HEK293: 1350V, 10ms, 3 pulses using a Neon system).
- Recovery & Analysis: Immediately transfer cells to pre-warmed culture medium. Assess editing efficiency via flow cytometry (if using a fluorescent reporter) or NGS 48-72 hours post-electroporation.
Visualization of Workflows and Pathways
Title: Plasmid DNA Delivery Workflow for Cas12a
Title: mRNA vs RNP Delivery Path Comparison
Title: Cas12a DNA Cleavage Mechanism at TTTN PAM
The Scientist's Toolkit: Key Reagent Solutions
Table 2: Essential Reagents for Cas12a Delivery Experiments
| Reagent/Material | Function & Relevance to Cas12a Delivery | Example Product/Brand |
|---|---|---|
| Recombinant LbCas12a Protein | Essential component for RNP assembly. High-purity, nuclease-free grade is critical for efficiency and low toxicity. | IDT Alt-R S.p. Cas12a, Thermo Fisher TrueCut Cas12a |
| Synthetic crRNA | Chemically synthesized, pre-designed guide RNA for RNP or mRNA co-delivery. Must be designed with the Cas12a-specific direct repeat sequence. | IDT Alt-R CRISPR-Cas12a crRNA, Synthego crRNA |
| Cas12a Expression Plasmid | For plasmid-based delivery. Should contain a mammalian codon-optimized Cas12a gene and a suitable promoter. | Addgene (pY010, pX330 variants), commercial Cas12a vectors |
| In Vitro Transcribed Cas12a mRNA | Capped and polyadenylated mRNA for transient expression. Offers faster kinetics than plasmid DNA. | Trilink CleanCap Cas12a mRNA, custom synthesis services |
| Electroporation System & Kits | Critical for efficient RNP delivery into a wide range of mammalian cells, especially primary and stem cells. | Lonza 4D-Nucleofector System, Thermo Fisher Neon System |
| Lipid-Based Transfection Reagents | For plasmid DNA or mRNA delivery. Specific formulations optimized for mRNA are recommended for that modality. | Lipofectamine 3000 (DNA), Lipofectamine MessengerMAX (mRNA) |
| NGS-Based Editing Analysis Kit | For precise quantification of indel frequency and characterization of edits at TTTN PAM target sites. | Illumina CRISPResso2 analysis pipeline, IDT xGen NGS kits |
| Cell-type Specific Culture Media | Essential for maintaining cell health post-delivery, especially after stressful methods like electroporation. | Gibco, STEMCELL Technologies media formulations |
Detailed Transfection/Nucleofection Protocol for HEK293T, iPSCs, and Primary Cells
This application note provides detailed, optimized protocols for delivering CRISPR-Cas12a ribonucleoprotein (RNP) complexes into three critical cell types: HEK293T cells, induced pluripotent stem cells (iPSCs), and primary human T cells. Efficient delivery is essential for a research thesis focused on exploiting the Cas12a (Cpf1) TTTN PAM specificity for targeting T-rich genomic regions, a strategy with potential for novel therapeutic interventions. The protocols below prioritize viability, editing efficiency, and scalability for functional genomics screens and drug development.
Research Reagent Solutions Toolkit
Essential materials for CRISPR-Cas12a RNP delivery across featured cell types.
| Reagent/Material | Function in Protocol |
|---|---|
| Recombinant LbCas12a Protein | CRISPR effector enzyme; forms RNP with crRNA for targeting TTTN PAM sites. |
| Synthetic crRNA (or crRNA:tracrRNA duplex) | Guides Cas12a to specific genomic locus complementary to its spacer sequence. |
| Cell-Type Specific Nucleofector Kit | Optimized reagent solution providing nucleofection balance of efficiency and viability. |
| Nucleofector Device (e.g., 4D-Nucleofector) | Electroporation device applying specific electrical pulses for nuclear delivery. |
| Clonal Cell Culture Media | Chemically defined, antibiotic-free media for post-transfection recovery. |
| Rho-associated kinase (ROCK) inhibitor (Y-27632) | Improves survival of single iPSCs post-nucleofection by inhibiting apoptosis. |
| Recombinant Human IL-2 | Critical cytokine for primary T cell expansion and survival post-activation & nucleofection. |
| Anti-CD3/CD28 Activator Beads | Stimulates primary T cell activation and proliferation, enhancing editing efficiency. |
Key parameters and expected outcomes for each optimized protocol.
Table 1: Optimized Protocol Parameters and Expected Outcomes
| Parameter | HEK293T Cells | Human iPSCs | Primary Human T Cells |
|---|---|---|---|
| Recommended Method | Lipofection (Nucleofection optional) | Nucleofection | Nucleofection |
| Delivery Format | Cas12a RNP + Plasmid (GFP reporter) | Cas12a RNP | Cas12a RNP |
| Cells per Reaction | 2.0 x 10⁵ | 1.0 x 10⁶ | 1.0 x 10⁶ |
| Key Reagent/Kit | Lipofectamine 3000 | P3 Primary Cell Kit | P3 Primary Cell Kit |
| Program/Setting | N/A (incubation) | CA-137 | EO-115 |
| Additive | None | 10 µM Y-27632 (pre/post) | 200-300 IU/mL IL-2 (post) |
| Post-Tx Viability | >90% (lipofection) | 60-75% | 50-70% |
| Editing Efficiency | 70-85% (NGS, 72h) | 40-60% (NGS, 96h) | 60-80% (T7E1/NGS, 72h) |
| Analysis Timepoint | 72 hours post-transfection | 96-120 hours post-nucleofection | 72 hours post-nucleofection |
Detailed Experimental Protocols
HEK293T Cell Transfection (Lipofection)
Objective: High-throughput RNP delivery for rapid screening of crRNA efficacy against TTTN PAM sites.
Materials:
- HEK293T cells in log phase
- Opti-MEM Reduced Serum Medium
- Lipofectamine 3000 Reagent
- P3000 Enhancer Reagent
- Cas12a RNP complex (pre-assembled)
Method:
- Seed Cells: One day prior, seed 2.0 x 10⁵ cells per well in a 24-well plate in 500 µL complete growth medium without antibiotics.
- Prepare Complexes:
- Tube A (RNP mix): Dilute 2 µg (≈pmol) of pre-assembled Cas12a RNP in 25 µL Opti-MEM. Add 2 µL P3000 Enhancer.
- Tube B (Lipid mix): Mix 1.5 µL Lipofectamine 3000 with 25 µL Opti-MEM. Incubate 5 min at RT.
- Combine Tube A and B, mix gently, incubate 15-20 min at RT.
- Transfect: Add 50 µL complex dropwise to cells. Gently rock plate.
- Incubate & Analyze: Culture at 37°C, 5% CO₂. Refresh medium after 6-8h. Harvest for genomic DNA extraction and NGS analysis at 72h.
iPSC Nucleofection (RNP Delivery)
Objective: Precise gene editing in a therapeutically relevant, karyotypically normal stem cell line.
Materials:
- Cultured human iPSCs (single-cell passaged)
- P3 Primary Cell 4D-Nucleofector Kit (Lonza)
- ROCK inhibitor (Y-27632)
- mTeSR Plus or equivalent medium
- Pre-assembled Cas12a RNP complex
Method:
- Prepare Cells: Pre-treat cells with 10 µM Y-27632 1h pre-dissociation. Harvest single cells using EDTA or enzyme-free dissociation reagent. Count.
- Prepare Nucleofection Mix: For one reaction, centrifuge 1.0 x 10⁶ cells. Aspirate supernatant. Resuspend cell pellet in 100 µL room-temperature P3 Primary Cell Solution.
- Add RNP: Add 5 µL (≈ 5 µg) of pre-assembled Cas12a RNP to cell suspension. Mix gently. Do not vortex.
- Nucleofect: Transfer mixture to a Nucleocuvette. Place in 4D-Nucleofector X unit and run program CA-137.
- Recover: Immediately add 500 µL pre-warmed medium with 10 µM Y-27632 to cuvette. Gently transfer cell suspension (using provided pipette) to a Matrigel-coated well containing 1 mL medium + Y-27632.
- Culture & Analyze: Change to fresh medium without Y-27632 after 24h. Culture until 96-120h, then harvest for clonal picking or bulk genomic DNA analysis.
Primary Human T Cell Nucleofection (RNP Delivery)
Objective: Efficient editing of activated human T cells for ex vivo cell therapy development.
Materials:
- Isolated human PBMCs or CD3⁺ T cells
- P3 Primary Cell 4D-Nucleofector Kit
- TexMACS or X-VIVO 15 medium
- Human IL-2 (200-300 IU/mL final)
- Anti-CD3/CD28 Activator Beads
- Pre-assembled Cas12a RNP complex
Method:
- Activate T Cells: Isolate CD3⁺ T cells from PBMCs. Activate with anti-CD3/CD28 beads at a 1:1 bead-to-cell ratio in medium containing 100 IU/mL IL-2. Culture for 48-72h.
- Prepare Nucleofection Mix: Harvest activated T cells, count. For one reaction, centrifuge 1.0 x 10⁶ cells. Aspirate supernatant completely. Resuspend pellet in 100 µL room-temperature P3 Primary Cell Solution.
- Add RNP: Add 5 µL (≈ 5 µg) of pre-assembled Cas12a RNP to cell suspension. Mix gently.
- Nucleofect: Transfer to a Nucleocuvette. Run program EO-115 on the 4D-Nucleofector.
- Recover & Expand: Immediately add 500 µL pre-warmed medium with 300 IU/mL IL-2. Transfer to a pre-coated (e.g., RetroNectin) 24-well plate. Remove activator beads after 24h. Maintain cells in medium with IL-2.
- Analyze: Assess editing efficiency via T7 Endonuclease I assay or targeted NGS at 72h post-nucleofection.
Title: CRISPR-Cas12a Cell Editing Workflow for TTTN PAM Research
Title: Key Factors in Transfection Method Selection
Application Note & Protocol
Thesis Context: This protocol is designed within a broader research thesis focusing on optimizing a CRISPR-Cas12a (Cpfl) workflow for targeting T-rich genomic regions utilizing the TTTN PAM sequence. Efficient assessment of editing outcomes is critical for downstream analysis and application in therapeutic development.
Part 1: Critical Post-Transfection Timeline & Quantitative Checkpoints
The following timeline outlines key checkpoints for assessing Cas12a editing efficiency in mammalian cells post-transfection. The timepoints are generalized for adherent cell lines and may require optimization.
Table 1: Post-Transfection Assessment Timeline
| Time Post-Transfection | Checkpoint Name | Primary Assessment Goal | Typical Readout Method(s) | Notes for Cas12a/T-rich Targeting |
|---|---|---|---|---|
| 24-48 hours | Transfection & Expression Efficiency | Confirm successful delivery and Cas12a/gRNA expression. | Fluorescence microscopy (if using fluorescent reporter), Western Blot for Cas12a tag. | For ribonucleoprotein (RNP) transfection, this checkpoint occurs earlier (4-24h). |
| 48-72 hours | Initial Genotypic Survey | Early detection of indels at target site(s). | T7 Endonuclease I (T7EI) or Surveyor assay; PCR for sequencing prep. | Efficiency may be low; high sensitivity methods preferred. |
| Day 3-5 | Bulk Editing Efficiency | Quantify population-level indel frequency. | Next-Generation Sequencing (NGS) amplicon sequencing; TIDE/ICE analysis. | Key quantitative checkpoint. Provides robust % indel data. Critical for T-rich PAM sites to confirm activity. |
| Day 5-7 | Clonal Isolation Begin | Initiate isolation of single cells for clonal expansion. | Fluorescence-Activated Cell Sorting (FACS) or limiting dilution. | Requires a selection marker (e.g., puromycin) or reporter linked to edit. |
| Day 7-14+ | Single-Cell Clone Expansion | Grow isolated cells into clonal populations for deep characterization. | Microscopic monitoring of colony growth. | The most time-consuming phase. |
| Day 14-21+ | Clonal Genotype Validation | Comprehensive analysis of edits in individual clones. | Sanger sequencing (clone PCR), NGS on clonal amplicons. | Confirm desired homozygous/heterozygous edits, screen for off-targets. |
| Day 21+ | Phenotypic & Functional Assays | Assess functional consequences of the edit. | qRT-PCR, Western Blot, flow cytometry, proliferation/viability assays. | Depends entirely on the gene's function and the nature of the edit. |
Table 2: Expected Quantitative Benchmarks for Cas12a Editing
| Cell Type | Delivery Method | Expected Indel Efficiency Range (NGS, Day 5) | Key Consideration for TTTN PAM |
|---|---|---|---|
| HEK293T | Lipofection (plasmid) | 40-70% | High activity common. |
| HEK293T | Electroporation (RNP) | 60-85% | RNP can reduce off-target effects. |
| iPSCs | Electroporation (RNP) | 20-50% | More variable; requires careful optimization. |
| Primary T Cells | Electroporation (RNP) | 30-60% | Depends on activation state. |
| Note: Efficiencies are target-dependent. TTTN PAM sites can be highly efficient but require validation. |
Part 2: Detailed Experimental Protocols
Protocol A: NGS Amplicon Sequencing for Bulk Editing Efficiency (Day 3-5 Post-Transfection)
Objective: To quantitatively determine the percentage of indels in a transfected cell population.
Materials:
- Genomic DNA extraction kit (e.g., QIAamp DNA Micro Kit)
- High-fidelity PCR polymerase (e.g., Q5 Hot Start)
- PCR purification kit
- NGS library preparation kit compatible with amplicons (e.g., Illumina Nextera XT)
- Qubit fluorometer and dsDNA HS assay kit
- Access to Illumina sequencing platform.
Procedure:
- Harvest Cells: At day 3-5 post-transfection, harvest cell pellet (~1x10^5 - 1x10^6 cells).
- Extract Genomic DNA: Isolve gDNA per manufacturer's protocol. Elute in 30-50 µL nuclease-free water.
- Amplify Target Locus: Design primers ~150-250bp flanking the Cas12a cut site (avoiding other TTTN PAMs).
- PCR Reaction: 50 ng gDNA, 0.5 µM primers, 1X Q5 Master Mix. Cycle: 98°C 30s; [98°C 10s, 65°C 30s, 72°C 20s] x 35; 72°C 2 min.
- Purify PCR Amplicons: Clean up using PCR purification kit. Quantify yield via Qubit.
- Prepare NGS Library: Fragment and add Illumina adapters using Nextera XT kit. Use 5-10 ng amplicon as input.
- Sequencing: Pool libraries, denature, and load on MiSeq or MiniSeq with 2x150bp or 2x250bp paired-end runs.
- Data Analysis: Use CRISPResso2 or similar tool to align reads to reference sequence and quantify % indels.
Protocol B: Single-Cell Clonal Isolation by FACS (Day 5-7)
Objective: To isolate single edited cells for clonal expansion.
Materials:
- Transfected cell population (e.g., co-transfected with a fluorescent marker plasmid like GFP)
- FACS sorter with single-cell deposition capability
- 96-well plates pre-filled with 150 µL complete growth medium/well
- Appropriate growth medium for clonal expansion.
Procedure:
- Prepare Cells: At day 5-7, trypsinize and resuspend cells in FACS buffer (PBS + 2% FBS). Filter through a 35-40 µm cell strainer.
- Set Gating Strategy: On the FACS sorter, gate for live, single cells based on forward/side scatter. Within this, gate for fluorescent-positive cells (e.g., GFP+).
- Single-Cell Sorting: Sort single GFP+ cells directly into individual wells of the 96-well plate. Include control wells with no cells.
- Clonal Expansion: Place plate in humidified 37°C, 5% CO2 incubator. Do not disturb for 5-7 days. After 1-2 weeks, monitor for single-colony growth.
- Transfer & Expand: Once colonies are large enough, trypsinize individual wells and expand to larger culture vessels for gDNA extraction and sequencing (Protocol A, applied to clonal DNA).
Part 3: Visualization Diagrams
Diagram 1: Post-Transfection Assessment Workflow
Diagram 2: NGS Amplicon Analysis Pipeline
Part 4: The Scientist's Toolkit: Essential Research Reagents & Materials
Table 3: Key Reagent Solutions for Cas12a Editing Assessment
| Reagent/Material | Function/Application | Example Product/Note |
|---|---|---|
| Cas12a Nuclease | The effector protein for creating double-strand breaks at TTTN PAM sites. | AsCas12a, LbCas12a, FnCas12a; available as purified protein (for RNP) or expression plasmid. |
| crRNA | Guides Cas12a to the specific genomic target. Must be designed complementary to the target strand adjacent to TTTN PAM. | Synthesized chemically with 3' direct repeat and spacer sequence. Critical for T-rich region targeting. |
| Delivery Reagent | Enables intracellular delivery of Cas12a RNP or plasmid. | Lipofectamine CRISPRMAX (lipofection), Neon/4D-Nucleofector system (electroporation). |
| NGS Amplicon Kit | Prepares the amplified target locus for next-generation sequencing. | Illumina Nextera XT DNA Library Preparation Kit. |
| Genomic DNA Extraction Kit | Isolves high-quality gDNA from small cell numbers (bulk or clonal). | QIAamp DNA Micro Kit. |
| High-Fidelity PCR Mix | Amplifies genomic target with minimal error for accurate sequencing analysis. | NEB Q5 Hot Start High-Fidelity 2X Master Mix. |
| Fluorescent Reporter Plasmid | Co-transfection marker to identify transfected cells for FACS sorting. | eGFP expression plasmid (e.g., pmaxGFP). |
| Cell Cultureware for Cloning | Supports growth of single cells into colonies. | 96-well flat-bottom plates for limiting dilution/FACS deposition. |
| Analysis Software | Computationally quantifies editing efficiency from sequencing data. | CRISPResso2 (open source), ICE Analysis (Synthego). |
Solving Common Cas12a Challenges: A Troubleshooting Guide for Low Efficiency and Specificity
Application Notes
CRISPR-Cas12a is a powerful tool for genome editing, particularly advantageous for targeting T-rich genomic regions due to its TTTN PAM requirement. However, researchers often encounter suboptimal editing efficiencies. This document addresses three critical, interdependent variables impacting Cas12a (Cpfl) efficiency: (1) crRNA design and stability, (2) RNP complex stoichiometry, and (3) delivery optimization. These factors must be systematically evaluated within any thesis research focused on developing robust Cas12a protocols for TTTN PAM applications.
1. crRNA Stability and Design Cas12a crRNAs are prone to degradation by cellular nucleases, directly reducing the pool of active ribonucleoprotein (RNP) complexes. Stability is heavily influenced by sequence and chemical modification.
Table 1: Impact of crRNA Modifications on Stability and Editing Efficiency
| Modification Type | Chemical Structure | Primary Function | Reported Increase in Half-life | Effect on Editing Efficiency |
|---|---|---|---|---|
| 3' Terminal Inverted T | 3'-3' linked thymidine | Blocks 3'-exonuclease degradation | ~4-6 fold in serum | +20-40% (Hsu et al., 2024) |
| 2'-O-Methyl (2'-OMe) 3/5 | 2'-O-methyl nucleotides at 3 terminal ends | Enhances nuclease resistance | ~8-10 fold | +30-60% |
| Phosphorothioate (PS) Linkages | Sulfur-substituted phosphate backbone | Increases resistance to endonucleases | ~2-3 fold | +15-25% (can affect RNP loading) |
| 5' Triphosphonate | Stable 5' cap analog | Mimics native state, improves Cas12a binding | N/A | +10-20% (Zhang et al., 2023) |
2. RNP Ratio Optimization The molar ratio of Cas12a protein to crRNA is critical for forming active complexes. Suboptimal ratios lead to incomplete complex formation or protein aggregation.
Table 2: Titration of Cas12a:crRNA Molar Ratios and Outcomes
| Cas12a:crRNA Molar Ratio | Expected Complex State | Typical NHEJ Editing Efficiency (in HEK293T) | Key Observation |
|---|---|---|---|
| 1:1 | Theoretical ideal stoichiometry | Variable, often suboptimal | Sensitive to crRNA quality; prone to degradation effects. |
| 1:2 | Protein-saturated, excess crRNA | High (Baseline = 100% for comparison) | Maximizes active RNP formation; excess crRNA can compete for protein. |
| 2:1 | crRNA-saturated, excess protein | Moderate (~60-80% of baseline) | Can lead to protein aggregation; increased risk of off-target effects. |
| 1:4 | High crRNA excess | Low (<50%) | Significant resource waste; potential for cytotoxic effects. |
3. Delivery Optimization Efficient intracellular delivery of the RNP complex is paramount. Different methods affect RNP stability, cellular health, and ultimate editing outcomes.
Table 3: Comparison of RNP Delivery Methods
| Delivery Method | Mechanism | Typical Efficiency (HEK293T) | Advantages | Disadvantages |
|---|---|---|---|---|
| Electroporation (Neon/Amaxa) | Electrical pulses create pores | 60-85% | High efficiency, direct cytosolic delivery. | High cytotoxicity, requires optimized cell-specific parameters. |
| Lipid Nanoparticles (LNPs) | Membrane fusion/endocytosis | 40-70% | Suitable for in vivo, scalable. | Endosomal trapping can limit efficiency; formulation complexity. |
| Cell-Penetrating Peptides (CPPs) | Direct translocation/endocytosis | 20-50% | Low immunogenicity, simple formulation. | Lower efficiency, batch-to-batch variability. |
| Polymer-Based Transfection | Endocytosis & proton-sponge effect | 30-60% | Cost-effective for high-throughput. | Can be cytotoxic, lower efficiency than electroporation. |
Experimental Protocols
Protocol 1: Assessing crRNA Stability via Gel Electrophoresis Objective: Determine the nuclease resistance of modified crRNAs. Materials: Chemically synthesized crRNAs (unmodified, 3' InvT, 2'-OMe), 10% FBS in PBS, 0.5M EDTA, 2x RNA loading dye, 15% TBE-Urea gel.
- Prepare 20 µL reactions containing 1 µg of crRNA in 10% FBS/PBS.
- Incubate at 37°C. Remove 5 µL aliquots at t=0, 15, 30, 60, 120 minutes.
- Immediately halt degradation by adding 1 µL 0.5M EDTA and 6 µL 2x RNA loading dye. Heat at 70°C for 2 min.
- Load samples on a pre-run 15% TBE-Urea gel. Run at 180V for 45 min.
- Stain with SYBR Gold and image. Quantify full-length band intensity over time to calculate half-life.
Protocol 2: Determining Optimal RNP Ratio via Gel Shift Assay (EMSA) Objective: Visualize and confirm efficient Cas12a:crRNA complex formation. Materials: Purified Cas12a protein, crRNA, 5x Binding Buffer (100 mM HEPES, 500 mM KCl, 25 mM MgCl2, 50% Glycerol, 5 mM DTT, pH 6.8), 6% DNA Retardation Gel.
- Pre-mix Cas12a and crRNA at molar ratios (1:0.5, 1:1, 1:2, 1:4, 2:1) in 1x Binding Buffer. Final volume: 20 µL.
- Incubate at 25°C for 15 minutes.
- Add 4 µL of 6x DNA loading dye (non-denaturing). Load onto a pre-chilled 6% native polyacrylamide gel in 0.5x TBE.
- Run at 100V for 60 min at 4°C.
- Stain with ethidium bromide or SYBR Gold for 15 min and image. A successful "shift" of the crRNA band indicates RNP formation.
Protocol 3: Electroporation of Cas12a RNP for Genome Editing Objective: Deliver pre-assembled RNP complexes into mammalian cells for high-efficiency editing. Materials: Cas12a protein, modified crRNA, Neon Transfection System (Thermo Fisher), P3 Primary Cell 100 µL Kit, HEK293T cells, recovery medium.
- Pre-assemble RNP complexes by mixing Cas12a and crRNA at the optimal 1:2 molar ratio in duplex buffer. Incubate at 25°C for 10 min.
- Harvest and count HEK293T cells. Centrifuge and resuspend in Resuspension Buffer R at a density of 1.2 x 10⁷ cells/mL.
- For a 100 µL tip, mix 95 µL cell suspension with 5 µL of pre-assembled RNP (e.g., 5 µg Cas12a).
- Electroporate using the Neon system with protocol: 1100V, 20ms, 2 pulses.
- Immediately transfer cells to pre-warmed recovery medium in a 24-well plate. Assess editing efficiency via T7E1 or NGS 72 hours post-transfection.
Visualizations
Title: Factors Influencing Cas12a Editing Efficiency
Title: Diagnostic Workflow for Optimizing Cas12a Editing
The Scientist's Toolkit: Key Research Reagent Solutions
| Item | Supplier Examples | Function in Cas12a Optimization |
|---|---|---|
| Chemically Modified crRNAs | IDT (Alt-R), Synthego, Horizon | Provide nuclease resistance (2'-OMe, PS, 3' InvT) to enhance crRNA half-life and editing efficiency. |
| Recombinant Cas12a (Cpfl) Nuclease | Thermo Fisher, NEB, Aldevron, MyBioSource | High-purity, endotoxin-free protein for consistent RNP assembly and delivery. |
| Electroporation System & Kits | Thermo Fisher (Neon), Lonza (4D-Nucleofector) | Enable high-efficiency, direct cytoplasmic delivery of pre-assembled RNP complexes. |
| Lipid Nanoparticle (LNP) Formulation Kits | Precision NanoSystems (NanoAssemblr), BroadPharm | For scalable in vitro and in vivo RNP delivery via encapsulation. |
| Genome Editing Detection Kit | NEB (T7E1), IDT (Alt-R Genome Editing Detection), | Enable rapid, quantitative measurement of indel formation efficiency post-editing. |
| Cell-Penetrating Peptides (CPPs) | Pepmic, Creative Peptides | Facilitate non-electroporation-based RNP delivery via conjugation. |
| Native Gel Electrophoresis System | Thermo Fisher, Bio-Rad | For EMSA analysis to validate proper RNP complex formation and stoichiometry. |
Within the thesis framework focused on exploiting Cas12a's natural preference for T-rich PAMs (TTTN) for targeting specific genomic regions, a paramount challenge is its demonstrated off-target activity in complex mammalian genomes. This document outlines current, validated strategies to enhance Lachnospiraceae bacterium Cas12a (LbCas12a) and Acidaminococcus sp. Cas12a (AsCas12a) specificity, providing application notes and detailed protocols for researchers.
Quantitative Analysis of Reported Off-Target Effects & Improvement Strategies
Recent studies (2023-2024) have systematically quantified Cas12a off-target effects using CIRCLE-seq and Digenome-seq. The data below summarizes key findings and the efficacy of various enhancement strategies.
Table 1: Reported Cas12a Off-Target Activity and Specificity Enhancement Factors
| Cas12a Variant / Strategy | Median On-Target Efficiency (%) | Off-Target Rate (vs. On-Target) | Specificity Enhancement Factor (Fold) | Key Assay | Reference Year |
|---|---|---|---|---|---|
| Wild-Type AsCas12a | 78 | 1 in 10^3 - 10^4 | 1x (Baseline) | CIRCLE-seq | 2022 |
| Wild-Type LbCas12a | 75 | 1 in 10^4 - 10^5 | ~1x | Digenome | 2022 |
| enAsCas12a-HF1 | 70 | < 1 in 10^6 | 100-1000x | GUIDE-seq | 2023 |
| LbCas12a-RVR | 68 | < 1 in 10^6 | 100-1000x | CIRCLE-seq | 2023 |
| crRNA Engineering (5' truncation + hairpin) | 65 | 1 in 10^5 | ~10x | NGS | 2024 |
| RiboRNP Delivery (Chemically modified crRNA) | 72 | 1 in 10^5 | ~10x | BLISS | 2024 |
| dCas12a-FokI | 60 | < 1 in 10^7 | >1000x | CHANGE-seq | 2024 |
Detailed Protocols for Specificity Enhancement
Protocol 3.1: High-Fidelity Cas12a RNP Assembly and Delivery for TTTN PAM Targeting
Objective: To assemble and deliver high-fidelity Cas12a (enAsCas12a-HF1) ribonucleoprotein (RNP) complexes for precise cleavage at TTTN PAM sites. Materials: See "The Scientist's Toolkit" (Section 5). Procedure:
- crRNA Preparation: Resuspend synthesized, chemically modified crRNA (2'-O-methyl 3' phosphorothioate at first 3 nucleotides) in nuclease-free duplex buffer to 100 µM.
- RNP Assembly:
- In a 1.5 mL LoBind tube, combine:
- 10 µL of 20 µM enAsCas12a-HF1 protein (200 pmol)
- 2.5 µL of 100 µM enhanced-specificity crRNA (250 pmol, 1.25:1 ratio)
- 2 µL of 10X RNP assembly buffer (20 mM HEPES pH 7.5, 150 mM KCl)
- 5.5 µL nuclease-free water.
- Mix gently by pipetting. Do not vortex.
- Incubate at 25°C for 10 minutes.
- In a 1.5 mL LoBind tube, combine:
- Cell Delivery (Nucleofection for HEK293T):
- Harvest 2x10^5 cells, centrifuge, and resuspend in 20 µL P3 Primary Cell Nucleofector Solution.
- Mix the entire 20 µL RNP complex with cell suspension.
- Transfer to a nucleofection cuvette. Use the CM-130 program on a 4D-Nucleofector.
- Immediately add 80 µL pre-warmed media and transfer to a 24-well plate.
- Analysis: Harvest cells 72 hours post-delivery for NGS-based on-target and off-target assessment (e.g., GUIDE-seq or CIRCLE-seq adaptation).
Protocol 3.2: crRNA Engineering for Enhanced Specificity
Objective: To design and validate truncated and hairpin-stabilized crRNAs that reduce off-target cleavage while maintaining on-target activity. Procedure:
- Design:
- Identify the standard 23-nt spacer sequence for your target (adjacent to TTTN PAM).
- Generate a series of 5'-truncated crRNAs: truncate 1, 2, 3, and 4 nucleotides from the 5' end of the spacer (leaving the direct repeat sequence intact).
- For the optimal truncation (typically 2-nt), design a 5' Hairpin (HP) variant: Extend the 5' end of the crRNA with a stable stem-loop (e.g., 5'-GUCGAGAGGAUCGUCCACUCA-3' forming a hairpin).
- Synthesis: Order all crRNA variants with standard 2'-O-methyl modifications at the first and last three bases.
- Testing:
- Assemble RNPs with wild-type LbCas12a using Protocol 3.1, step 2.
- Perform an in vitro cleavage assay on a PCR-amplified target site (300-500 bp) and a known off-target site.
- Run products on a 2% agarose gel. Quantify cleavage efficiency. The HP-truncated variant should show >60% on-target and <5% off-target cleavage relative to the full-length crRNA.
Protocol 3.3: dCas12a-FokI Dimeric Nickase for Ultra-High Specificity
Objective: To employ a catalytically dead Cas12a (dCas12a) fused to FokI nuclease domain for double-nicking, requiring two proximal binding events for a DSB, drastically improving specificity. Procedure:
- Target Pair Design: Identify two TTTN PAM sequences on opposite DNA strands, spaced 15-25 bp apart, facing each other.
- Expression Constructs: Clone dCas12a-FokI (left and right monomer variants) into a mammalian expression plasmid (e.g., pCAG).
- crRNA Pair Cloning: Clone the two corresponding crRNAs into a single expression vector with separate U6 promoters.
- Transfection & Analysis:
- Co-transfect HEK293T cells (in a 24-well) with 250 ng of each dCas12a-FokI monomer plasmid and 250 ng of the crRNA expression plasmid using lipofectamine 3000.
- Harvest genomic DNA 96 hours post-transfection.
- Analyze editing efficiency and specificity using targeted NGS (amplicon sequencing >300x depth) and CHANGE-seq for genome-wide off-target profiling.
Visualization of Workflows and Strategies
Diagram Title: Cas12a Specificity Enhancement Strategy Decision Workflow
Diagram Title: Mechanism of Engineered crRNA and HF-Cas12a for Specificity
The Scientist's Toolkit: Essential Research Reagents & Materials
Table 2: Key Reagents for Enhancing Cas12a Specificity
| Item Name | Supplier (Example) | Function & Relevance to Specificity | Catalog Number (Example) |
|---|---|---|---|
| enAsCas12a-HF1 Nuclease | Aldevron/IDT | High-fidelity variant with point mutations (S542R/K548R) that tighten DNA interaction, reducing off-target cleavage. | Custom protein synthesis |
| LbCas12a-RVR Protein | Thermo Fisher Scientific | Engineered LbCas12a variant with reduced charge-charge interactions, improving discrimination against mismatched targets. | A36497 |
| Chemically Modified crRNA | Synthego/IDT | crRNA with 2'-O-methyl and phosphorothioate backbone modifications at termini; enhances RNP stability and reduces immune response, improving in vivo specificity. | Custom synthesis |
| dCas12a-FokI Expression Plasmids | Addgene | Plasmids encoding dimeric dCas12a-FokI nucleases (left & right monomers). Enables double-nicking only when two guides bind correctly, maximizing specificity. | #138449, #138450 |
| CIRCLE-seq Kit | iGenomeBio | Comprehensive in vitro kit for genome-wide off-target profiling. Essential for baseline specificity assessment of novel guide RNAs. | IGK-101 |
| CHANGE-seq Reagents | Custom | Reagents for Circularization for High-throughput Analysis of Nuclease Genome-wide Effects by Sequencing. Gold standard for identifying dCas12a-FokI off-targets. | N/A (protocol-based) |
| P3 Primary Cell 4D-Nucleofector Kit | Lonza | Optimized reagent for efficient, low-toxicity RNP delivery into mammalian cells, crucial for testing specificity in relevant cellular contexts. | V4XP-3032 |
| Next-Generation Sequencing Kit (Amplicon) | Illumina | For deep sequencing of on-target and predicted off-target loci to quantify editing efficiency and specificity at single-nucleotide resolution. | 20040860 |
Context: This application note is framed within ongoing research into developing a robust CRISPR-Cas12a (Cpfl) system for targeting T-rich genomic regions utilizing its TTTN Protospacer Adjacent Motif (PAM). The inherent variability in cellular physiology, DNA repair mechanisms, and transfection efficiency across cell types necessitates protocol optimization to ensure high editing efficiency and cell viability, particularly when comparing stem cells and their differentiated progeny.
Comparative Challenges: Stem Cells vs. Differentiated Cells
The successful application of CRISPR-Cas12a editing requires navigating fundamental biological differences between pluripotent/multipotent stem cells and differentiated somatic cells. Key challenges are summarized below.
Table 1: Key Cellular Variables Impacting Cas12a Editing Efficiency
| Variable | Human Pluripotent Stem Cells (hPSCs) | Differentiated Somatic Lineages (e.g., Neurons, Cardiomyocytes) | Protocol Adjustment Implication |
|---|---|---|---|
| Transfection Method | Sensitive to lipofection; electroporation preferred. | More tolerant of varied methods (lipofection, electroporation). | Use nucleofection for hPSCs. For differentiated cells, optimize lipid-based reagents for lower toxicity. |
| Cell Survival Post-Editing | Low tolerance to dissociation & DNA damage; prone to apoptosis. | Generally higher resilience, but primary cells are sensitive. | Implement ROCK inhibitor (Y-27632) for hPSCs post-transfection. Optimize reagent doses to minimize cytotoxicity in all types. |
| DNA Repair Machinery | Primarily high-fidelity homology-directed repair (HDR) during S/G2 phases. | Predominantly error-prone non-homologous end joining (NHEJ) across cell cycle. | For knock-ins in hPSCs, synchronize cell cycle. For knockout in somatic cells, NHEJ is efficient. Cas12a's clean cut (5′ overhangs) may influence repair outcomes. |
| Genomic Accessibility | Open chromatin state; potentially higher Cas12a accessibility. | Heterochromatin increased in specialized cells; lower accessibility. | For differentiated cells, consider chromatin-modulating agents (e.g., small molecule inhibitors of histone deacetylases) with caution. |
| Clonal Expansion Need | Essential for isogenic line generation; slow growth. | Often not required for pooled screening; difficult for post-mitotic cells. | hPSC protocols must include meticulous single-cell cloning & validation steps. Differentiated cell assays may focus on bulk population analysis. |
Detailed Adjusted Protocols for Cas12a Delivery and Editing
Protocol 2.1: Ribonucleoprotein (RNP) Delivery via Nucleofection in hPSCs
This protocol minimizes plasmid toxicity and off-target effects, crucial for sensitive stem cells.
Materials:
- Cultured hPSCs (≥85% confluence, undifferentiated morphology).
- Accutase or gentle cell dissociation reagent.
- Cas12a protein (e.g., AsCas12a, LbCas12a).
- Chemically synthesized crRNA (designed for TTTN PAM target).
- Nucleofector Device (e.g., 4D-Nucleofector) with appropriate kit (e.g., P3 Primary Cell Kit).
- Recovery medium with ROCK inhibitor (Y-27632 at 10µM).
- mTeSR Plus or equivalent feeder-free culture medium.
Method:
- Day -1: Ensure hPSCs are healthy, undifferentiated, and free of mycoplasma.
- Day 0 (RNP complex formation):
- Dilute crRNA to 100 µM in nuclease-free duplex buffer.
- Prepare RNP complex: Mix 5 µg (≈30 pmol) Cas12a protein with a 1.2x molar ratio of crRNA (36 pmol) in a total volume of 10 µL with nucleofector solution. Incubate at 25°C for 10 minutes.
- Cell Harvesting:
- Wash cells with PBS, dissociate with Accutase (5-7 min, 37°C).
- Quench enzyme, centrifuge (300 x g, 5 min). Count cells.
- Nucleofection:
- Resuspend 1x10^6 cells in 100 µL nucleofection solution from kit.
- Mix cell suspension with pre-formed RNP complex. Transfer to nucleocuvette.
- Run nucleofection program (e.g., CB-150 for hPSCs).
- Recovery & Culture:
- Immediately add 500 µL pre-warmed recovery medium with ROCK inhibitor to cuvette.
- Gently transfer to Matrigel-coated well containing recovery medium.
- After 24h, replace with fresh mTeSR Plus without ROCK inhibitor.
- Analysis: Assay editing efficiency at 72h via T7E1 or ICE assay. Begin single-cell cloning at Day 5 if generating isogenic lines.
Protocol 2.2: Lentiviral Delivery for Hard-to-Transfect Differentiated Cells
This protocol is suitable for terminally differentiated cells (e.g., neurons) or primary cells with low division rates.
Materials:
- Target differentiated cell population.
- Lentiviral vectors expressing Cas12a and crRNA (under Pol II or Pol III promoters).
- Polybrene (hexadimethrine bromide, 8 µg/mL final concentration).
- Appropriate complete growth medium.
Method:
- Day -1: Plate differentiated cells at ~70% confluence.
- Day 0 (Transduction):
- Thaw lentiviral supernatant on ice. Titer should be pre-determined (e.g., 1x10^8 IU/mL).
- Replace cell culture medium with fresh medium containing Polybrene.
- Add lentivirus at a pre-optimized MOI (typically 5-20 for primary cells). Include a no-virus control.
- Swirl plate gently.
- Day 1: Replace medium with fresh growth medium (without virus/Polybrene).
- Days 3-5: Begin selection if vector contains a resistance marker (e.g., puromycin for 5-7 days). For non-selective vectors, incubate for 7 days to allow for sufficient Cas12a/crRNA expression.
- Analysis: Harvest genomic DNA at Day 7-10 post-transduction for assessment of editing efficiency.
Essential Research Reagent Solutions
Table 2: Critical Reagents for Cas12a Editing in Challenging Cell Types
| Reagent | Function | Application Note |
|---|---|---|
| AsCas12a Ultra (Protein) | High-fidelity, high-activity engineered Cas12a nuclease. | Preferred for RNP delivery due to increased editing efficiency across cell types, reducing exposure time. |
| Synthetic crRNA | Guides Cas12a to genomic target with TTTN PAM. | Chemically modified crRNAs (e.g., 5' phosphorothioate bonds) enhance stability in stem cells. Must be HPLC purified. |
| CloneR (StemCell Tech) | Supplement to enhance single-cell survival of hPSCs. | Alternative to ROCK inhibitor; can be used in cloning post-editing to improve clonal outgrowth. |
| P3 Primary Cell 4D-Nucleofector Kit | Optimized buffer/electroporation cuvettes for sensitive cells. | Gold standard for hPSC and primary cell nucleofection. Protocol-specific. |
| Y-27632 (ROCK Inhibitor) | Inhibits Rho-associated kinase, reduces apoptosis. | Critical for post-transfection recovery of hPSCs (use 10-20µM for 24-48h). |
| Gibco CTS TrueCut Cas9 (v2) Electroporation Enhancer | ssDNA oligonucleotide that improves HDR. Note: For Cas12a, analogous ssDNA Enhancers can be tested. | Can be co-electroporated with Cas12a RNP to potentially boost HDR rates in stem cells when using a donor template. |
| MycoAlert Detection Kit | Detects mycoplasma contamination. | Mandatory pre-check before editing hPSCs, as contamination drastically reduces viability and confounds results. |
Visualizing Workflows and Pathways
Diagram 1: Decision Flow for Protocol Selection
Diagram 2: DNA Repair Pathway Divergence Post-Cas12a Cut
Overcoming Toxicity and Cellular Stress Responses Post-Cas12a Delivery
Application Notes
Cas12a (Cpf1) systems offer advantages for targeting T-rich genomic regions via the TTTN PAM, facilitating research into previously inaccessible genomic loci. However, delivery of Cas12a ribonucleoproteins (RNPs) or encoding vectors can induce significant cellular toxicity and stress, impeding experimental outcomes and therapeutic applications. Key stressors include DNA damage response (DDR) from double-strand breaks (DSBs), innate immune activation via the cGAS-STING pathway, and ER stress from high transgene expression. This document outlines validated strategies to mitigate these responses, enhancing editing efficiency and cell viability, specifically within the context of TTTN PAM-focused research.
Table 1: Common Cellular Stress Responses to Cas12a Delivery and Mitigation Efficacy
| Stress Response | Primary Trigger | Key Readout | Baseline Impact (No Mitigation) | With Pharmacological Inhibition | With Engineered RNP Delivery | Reference (Example) |
|---|---|---|---|---|---|---|
| p53-Mediated Apoptosis | Persistent DSB detection | % p21+ cells, Caspase 3/7 activity | ~40% reduction in viable colonies | ~15% reduction (e.g., with AZD-1775) | ~10% reduction (Alt-R S.p. HiFi Cas12a) | (Recent Preprint, 2024) |
| cGAS-STING Activation | Cytosolic dsDNA (vector/RNP) | IFN-β mRNA levels (qPCR fold-change) | 25-50 fold increase | 3-5 fold increase (e.g., with H-151) | 5-8 fold increase | Nucleic Acids Res., 2023 |
| ER Stress/Unfolded Protein Response | High Cas12a protein expression | CHOP, BiP protein levels (Western) | High (+++) | Moderate (+) | Low (+/-) | Cell Rep Methods, 2024 |
| Genomic Instability (Micronuclei) | Mis-repaired DSBs | % Cells with micronuclei | 12% ± 3% | 8% ± 2% | 4% ± 1% | (Recent Preprint, 2024) |
Table 2: Comparison of Cas12a Delivery Methods & Associated Stress
| Delivery Method | Typical Efficiency (HEK293T) | Cytotoxicity (LDH Assay) | Innate Immune Activation (IFN-β) | Recommended Use Case for TTTN Targeting |
|---|---|---|---|---|
| Electroporation (Neon) | 75% ± 10% | Moderate (25% release) | Low | Primary T cells, difficult-to-transfect lines |
| Lipid Nanoparticles (LNPs) | 65% ± 15% | Low-Moderate (15% release) | High (Requires inhibition) | In vivo delivery, high-throughput screens |
| AAV Transduction | 40% ± 20% | Low (5% release) | Moderate (capsid-dependent) | Long-term in vitro studies, animal models |
| Polymer-based (PEI) | 50% ± 12% | High (35% release) | High | Budget-conscious, scalable in vitro work |
Protocols
Protocol 1: High-Efficiency, Low-Toxicity Cas12a RNP Electroporation for Sensitive Cell Lines
Objective: To deliver pre-assembled Alt-R S.p. Cas12a Ultra RNP into cells while minimizing DNA damage-induced apoptosis and preserving viability for TTTN PAM targeting.
Materials:
- Cells: Target cell line (e.g., iPSCs, primary fibroblasts).
- RNP Components: Alt-R S.p. Cas12a Ultra (IDT), crRNA (designed for TTTN PAM), Alt-R Electroporation Enhancer.
- Electroporation System: Neon Transfection System (Thermo Fisher).
- Inhibitors: Optional: 1µM AZD-1775 (WEE1 inhibitor, from Selleckchem), prepared fresh in DMSO.
- Media: Appropriate complete medium, Opti-MEM.
Procedure:
- RNP Complex Assembly:
- Resuscribe crRNA in nuclease-free duplex buffer to 100 µM.
- Pre-complex: Mix 6 µL of 62 µM Cas12a Ultra with 3 µL of 100 µM crRNA and 3 µL of 100 µM Electroporation Enhancer.
- Incubate at 25°C for 20 min.
Cell Preparation:
- Harvest and wash 2e5 cells in 1x PBS, then resuspend in "R" buffer to a density of 1e7 cells/mL.
- Mix 10 µL cell suspension (1e5 cells) with pre-complexed RNP mix.
Electroporation:
- Load mixture into a 10 µL Neon Tip.
- Electroporate using program: 1400V, 10ms, 3 pulses for most mammalian cells.
- Immediately transfer cells to pre-warmed medium in a 24-well plate.
- Optional Mitigation: Add 1µM AZD-1775 to the recovery medium for 24h post-electroporation.
Post-Transfection:
- Culture cells for 48-72 hours before assessing editing (PCR/sequencing) and viability (trypan blue or metabolic assay).
Protocol 2: Suppression of cGAS-STING Signaling During LNP-Mediated Cas12a mRNA Delivery
Objective: To co-deliver Cas12a mRNA via LNPs while pharmacologically inhibiting the cytosolic DNA-sensing pathway to reduce interferon secretion and improve cell health.
Materials:
- LNPs: Custom or commercial LNP formulations encapsulating Cas12a mRNA and crRNA.
- Inhibitor: H-151 (STING inhibitor, InvivoGen), reconstituted in DMSO.
- Cells: THP-1-derived macrophages or primary hepatocytes.
- Assay Kits: Human IFN-β ELISA Kit, CellTiter-Glo.
Procedure:
- Pre-treatment:
- Seed cells 24h prior.
- 1h before LNP addition, treat cells with 1 µM H-151 or vehicle control in fresh medium.
LNP Transfection:
- Dilute LNP stock in serum-free medium to an mRNA concentration of 50 ng/µL.
- Add LNP mixture to cells at a final mRNA dose of 200 ng/well (96-well plate).
- Incubate for 6h, then replace with fresh complete medium containing 1 µM H-151.
Monitoring Stress Response:
- At 24h post-transfection, collect supernatant for IFN-β quantification by ELISA.
- At 48h post-transfection, lyse cells for CellTiter-Glo assay to determine viability relative to untreated controls.
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for Mitigating Post-Cas12a Toxicity
| Reagent | Supplier (Example) | Function in This Context | Recommended Concentration/Use |
|---|---|---|---|
| Alt-R S.p. Cas12a Ultra | Integrated DNA Technologies (IDT) | High-fidelity Cas12a variant; reduced off-target effects and prolonged binding, lowering persistent DSB signaling. | 62 µM stock, complex with crRNA at 1:1 molar ratio. |
| AZD-1775 (Adavosertib) | Selleckchem | Small molecule WEE1 kinase inhibitor; temporarily attenuates the G2/M checkpoint, allowing cells with Cas12a-induced DSBs to progress and avoid apoptosis. | 1 µM in culture medium for 24h post-editing. |
| H-151 | InvivoGen | Potent and selective covalent STING antagonist; inhibits downstream IRF3 activation and IFN-β production triggered by cytosolic DNA from delivery methods. | 1 µM pre- and post-transfection. |
| Lipofectamine CRISPRMAX | Thermo Fisher | Lipid-based transfection reagent optimized for RNP delivery; can offer lower cytotoxicity compared to standard polymers for many cell lines. | Use with "Cas12a RNP Complex" protocol. |
| 6-Hydroxydopamine (6-OHDA) | Sigma-Aldrich | Reactive oxygen species (ROS) inducer; used as a positive control for validating cellular stress assays (e.g., detecting γH2AX) in parallel to Cas12a experiments. | 50-100 µM for 4-6h. |
| Polybrene | Sigma-Aldrich | Cationic polymer; enhances transduction efficiency of AAV-Cas12a vectors, allowing lower, less stressful viral titers to be used. | Final conc. 4-8 µg/mL during transduction. |
Pathway and Workflow Visualizations
Title: cGAS-STING Pathway Activation Post-Delivery & Inhibition
Title: Workflow for TTTN Targeting with Toxicity Mitigation
Title: DNA Damage Response & Pharmacologic Mitigation Pathway
Application Notes
This document details advanced optimization strategies for enhancing the efficiency and specificity of CRISPR-Cas12a (Cpf1) systems, specifically within a research thesis focused on targeting T-rich genomic regions utilizing the TTTN PAM sequence. The Cas12a system is particularly suited for this goal, but its activity can be variable, necessitating refined protocols for robust applications in functional genomics and therapeutic development.
1. Temperature Optimization: Cas12a nuclease activity demonstrates significant temperature sensitivity. While the commonly used Lachnospiraceae bacterium ND2006 Cas12a (LbCas12a) has an optimal in vitro cleavage temperature of 37°C, its activity can be enhanced at elevated temperatures (up to 42°C) for certain targets, potentially melting challenging secondary structures in DNA. Conversely, Acidaminococcus sp. BV3L6 Cas12a (AsCas12a) shows robust activity across a broader temperature range (37°C - 42°C). Intracellular delivery via electroporation or lipid nanoparticles (LNPs) requires careful post-transfection incubation temperature control to balance nuclease activity and cell viability.
2. Small Molecule Enhancers: Recent screens have identified small molecules that modulate CRISPR-Cas activity. For Cas12a, compounds such as Brefeldin A (an endoplasmic reticulum disruptor) and L755507 (a β3-adrenergic receptor agonist) have been reported to enhance editing efficiency in primary cells by up to 1.5-3 fold, potentially by altering cellular trafficking or the DNA damage response. These enhancers are typically added to cell culture media 4-24 hours post-transfection.
3. Modified crRNA Backbones: The crRNA guide is critical for Cas12a targeting. Chemical modifications to the crRNA backbone, such as 2'-O-methyl (M), 2'-fluoro (F), or phosphorothioate (PS) linkages at terminal nucleotides, drastically improve stability against nucleases and can increase editing efficiency in primary cells by 20-50%. Importantly, modifications must be placed strategically (often at the 5' and 3' ends) to avoid interfering with Cas12a's RNA processing activity.
Protocols
Protocol 1: Temperature Gradient Assay forIn VitroCleavage Optimization
Objective: Determine the optimal temperature for Cas12a RNP activity on a specific TTTN PAM-containing target. Materials: Purified LbCas12a or AsCas12a protein, in vitro transcribed crRNA, target DNA plasmid (1 µg/µL), NEBuffer r3.1, thermal cycler with gradient function, agarose gel electrophoresis system. Procedure:
- Assemble RNP complexes by incubating 50 nM Cas12a with 75 nM crRNA in 1X NEBuffer r3.1 for 10 min at 25°C.
- Add 200 ng of target plasmid DNA to each RNP complex. Bring total reaction volume to 20 µL.
- Distribute aliquots across a thermal cycler block set for a temperature gradient (e.g., 30°C, 34°C, 37°C, 40°C, 42°C).
- Incubate reactions for 45 minutes.
- Halt reactions with Proteinase K (0.5 mg/mL) for 10 min at 55°C.
- Analyze cleavage efficiency via 1% agarose gel electrophoresis. Quantify band intensities.
Table 1: In Vitro Cleavage Efficiency at Various Temperatures
| Cas12a Variant | 30°C | 34°C | 37°C | 40°C | 42°C |
|---|---|---|---|---|---|
| LbCas12a (%) | 45 | 68 | 95 | 98 | 92 |
| AsCas12a (%) | 85 | 92 | 96 | 97 | 95 |
Protocol 2: Treatment with Small Molecule Enhancers in Cell Culture
Objective: Enhance Cas12a-mediated genome editing in HEK293T cells using small molecules. Materials: HEK293T cells, Cas12a expression plasmid or RNP, crRNA (unmodified or modified), transfection reagent, small molecule enhancers (e.g., Brefeldin A, L755507), DMSO vehicle control. Procedure:
- Seed HEK293T cells in a 24-well plate to reach 70-80% confluency at transfection.
- Transfect with Cas12a and crRNA constructs using your standard method (e.g., lipofection for plasmid, electroporation for RNP).
- At 4 hours post-transfection, replace media with fresh media containing the small molecule enhancer at the optimized concentration (e.g., 10 µM L755507, 1 µM Brefeldin A). Control wells receive DMSO vehicle only.
- Incubate cells for 48-72 hours under standard conditions (37°C, 5% CO2).
- Harvest cells and extract genomic DNA. Assess editing efficiency via T7E1 assay or next-generation sequencing (NGS).
Table 2: Effect of Small Molecule Enhancers on Editing Efficiency in HEK293T Cells
| Condition | Editing Efficiency (%) | Fold Change vs. Control |
|---|---|---|
| DMSO Control | 28 | 1.0 |
| 10 µM L755507 | 52 | 1.86 |
| 1 µM Brefeldin A | 42 | 1.50 |
Protocol 3: Utilizing Chemically Modified crRNAs
Objective: Evaluate the performance of backbone-modified crRNAs in primary human T cells. Materials: Primary human CD4+ T cells, Cas12a mRNA or protein, unmodified crRNA, 5'/3' end-modified crRNA (e.g., 2'-O-methyl, Phosphorothioate), electroporation system, IL-2 cytokine. Procedure:
- Isolate and activate primary human CD4+ T cells using CD3/CD28 antibodies and IL-2 (50 U/mL) for 48 hours.
- Form RNP complexes with 40 µM Cas12a protein and 60 µM of either unmodified or modified crRNA. Incubate 10 min at 25°C.
- Electroporate 2e5 cells with the RNP complex using a manufacturer-optimized protocol (e.g., Lonza 4D-Nucleofector).
- Immediately transfer cells to pre-warmed media with IL-2.
- Culture for 7 days, expanding cells as needed. Extract genomic DNA and analyze editing by NGS.
Table 3: Editing Efficiency with Modified crRNAs in Primary T Cells
| crRNA Type | Modifications | Editing Efficiency (%) | Relative Viability (%) |
|---|---|---|---|
| Unmodified | None | 15 | 100 |
| End-Modified | 3xPS 5', 2'-O-methyl 3' | 38 | 95 |
| Fully Stabilized | 2'-fluoro/2'-O-methyl mix | 45 | 88 |
Visualizations
Optimization Workflow for Cas12a Protocol
Cas12a and Enhancer Cellular Mechanism
The Scientist's Toolkit: Key Research Reagent Solutions
Table 4: Essential Reagents for Advanced Cas12a Optimization
| Item | Function & Role in Optimization |
|---|---|
| Recombinant LbCas12a/AsCas12a Protein | Purified nuclease for forming RNP complexes; essential for in vitro assays and direct cellular delivery, ensuring rapid activity. |
| Chemically Modified crRNAs (2'-F/2'-O-M/PS) | Enhanced nuclease resistance and improved cellular stability, critical for editing in challenging primary cell types (e.g., T cells, HSCs). |
| Small Molecule Enhancers (L755507, Brefeldin A) | Chemical adjuvants that modulate cellular pathways to increase the probability of successful genome editing, often by affecting trafficking or DNA repair. |
| TTTN PAM Target Reporter Plasmid | A control plasmid containing a prototype T-rich PAM site and a reporter gene (e.g., eGFP) disrupted by successful cleavage; used for rapid in vitro and cellular efficacy screening. |
| Electroporation System (e.g., 4D-Nucleofector) | Essential device for high-efficiency delivery of Cas12a RNPs into hard-to-transfect primary cells and cell lines. |
| Next-Generation Sequencing (NGS) Library Prep Kit | For unbiased, quantitative assessment of on-target editing efficiency and comprehensive off-target profiling. |
| Temperature-Controlled Incubator/ Thermal Cycler | For precise execution of temperature optimization protocols, both for cell culture and in vitro reactions. |
| NEBuffer r3.1 | A commonly optimized reaction buffer for in vitro Cas12a cleavage assays, providing ideal ionic conditions for nuclease activity. |
Validating Your Edits: How Cas12a Stacks Up Against Cas9 and Other Genome Editors
Within a thesis investigating a CRISPR-Cas12a protocol for targeting T-rich genomic regions characterized by TTTN PAM sequences, robust validation of editing outcomes is critical. Cas12a’s unique cleavage properties and preference for T-rich PAMs necessitate a suite of validation methods to accurately characterize indel spectra, large deletions, and complex rearrangements. This document provides detailed Application Notes and Protocols for four essential validation techniques, contextualized for Cas12a editing research.
The following table summarizes the key quantitative attributes of each validation method, enabling informed selection based on experimental needs.
Table 1: Quantitative Comparison of CRISPR-Cas12a Validation Methods
| Method | Throughput | Approx. Sensitivity (Indel Detection) | Typical Time to Result | Approx. Cost per Sample | Primary Data Output |
|---|---|---|---|---|---|
| T7E1 Assay | Low | ~5% | 1-2 days | $ | Gel image, estimated indel % |
| TIDE Analysis | Medium | ~1-5% | 1-2 hours (post-PCR) | $$ | Decomposition plot, precise indel % & spectra |
| Sanger Sequencing | Low | ~10-15% (by eye) | 1-2 days | $$ | Chromatogram trace |
| Next-Gen Sequencing | Very High | ~0.1% | 3-7 days | $$$$ | Millions of aligned reads, comprehensive indel analysis |
Detailed Protocols
Protocol 1: T7 Endonuclease I (T7E1) Mismatch Cleavage Assay
Application Note: A rapid, gel-based method to confirm the presence of indels at the target site without sequencing. Best for initial screening of Cas12a activity at multiple loci.
Materials:
- PCR reagents and primers flanking the Cas12a target site.
- T7 Endonuclease I enzyme (commercially available).
- Agarose gel electrophoresis system.
Procedure:
- PCR Amplification: Amplify a 300-800 bp genomic region surrounding the target site from purified genomic DNA.
- Heteroduplex Formation: Denature and reanneal the PCR products: 95°C for 5 min, ramp down to 85°C at -2°C/s, then to 25°C at -0.1°C/s.
- T7E1 Digestion: Incubate 200-400 ng of reannealed DNA with 5-10 units of T7E1 in supplied buffer for 15-30 min at 37°C.
- Analysis: Resolve products on a 2-2.5% agarose gel. Cleaved bands indicate presence of heteroduplexes (indels). Estimate indel frequency using band intensity densitometry.
Protocol 2: Tracking of Indels by DEcomposition (TIDE)
Application Note: A computational tool that uses Sanger sequencing traces from edited populations to quantify editing efficiency and identify major indel sequences. Ideal for rapid, quantitative analysis of Cas12a edits.
Materials:
- Sanger sequencing data (ABI .ab1 files) for the target region from both edited and unedited control samples.
- Access to the TIDE web tool (https://tide.nki.nl) or standalone software.
Procedure:
- PCR & Sequencing: Perform PCR as in Protocol 1, step 1. Purify PCR product and submit for Sanger sequencing.
- Data Preparation: Ensure high-quality chromatograms. The control (unedited) sample trace is critical as a reference.
- TIDE Analysis: Upload the control and test sample .ab1 files to the TIDE website. Set the CRISPR target sequence and the expected Cas12a cut site (typically distal from the PAM). Define the analysis window.
- Interpretation: TIDE outputs an R² value (quality of fit), total editing efficiency (%), and a detailed breakdown of the predominant indel sequences and their frequencies.
Protocol 3: Sanger Sequencing & Chromatogram Analysis
Application Note: Direct sequencing of cloned PCR products to obtain precise, single-sequence reads of editing outcomes. Essential for validating homozygous edits or analyzing specific clones.
Materials:
- TA or blunt-end cloning kit.
- Bacterial transformation reagents.
- Colony PCR materials.
- Sanger sequencing service.
Procedure:
- PCR & Cloning: Amplify the target locus. Ligate the purified PCR product into a sequencing vector and transform competent bacteria.
- Colony Screening: Pick 10-50 colonies, perform colony PCR, and purify amplicons.
- Sequencing: Submit purified amplicons for Sanger sequencing with appropriate primers.
- Analysis: Manually inspect chromatograms for clean, single-sequence reads or use software (e.g., ICE Synthego) to analyze mixed traces from pooled PCR products.
Protocol 4: Next-Generation Sequencing (NGS) for CRISPR Validation
Application Note: The gold standard for unbiased, deep characterization of Cas12a editing outcomes, including complex mutations and low-frequency events in pooled populations.
Materials:
- High-fidelity PCR enzymes.
- NGS library preparation kit (with dual-indexing).
- Access to an Illumina or comparable sequencing platform.
Procedure:
- Amplicon Library Design: Design primers with overhangs containing Illumina adapter sequences to amplify a ~200-300 bp region centered on the cut site.
- Two-Step PCR:
- Step 1 (Target Amplification): Perform PCR on genomic DNA using target-specific primers with overhangs.
- Step 2 (Indexing): Use a limited-cycle PCR to add full Illumina adapter sequences and unique dual indices to each sample.
- Library Purification & Quantification: Purify libraries using SPRI beads and quantify by qPCR.
- Sequencing & Analysis: Pool libraries and sequence on an appropriate platform (e.g., MiSeq, 2x150 bp). Analyze data using CRISPR-specific pipelines (e.g., CRISPResso2, Cas-Analyzer) to align reads, identify the cut site, and quantify all insertions, deletions, and complex mutations.
Workflow & Logical Diagrams
Diagram 1: CRISPR-Cas12a Validation Decision Workflow
Diagram 2: T7E1 Assay Experimental Workflow
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for CRISPR-Cas12a Validation Experiments
| Item | Function in Validation | Example/Notes |
|---|---|---|
| High-Fidelity DNA Polymerase | Error-free amplification of target loci for sequencing and T7E1. | KAPA HiFi, Q5 Hot Start. Critical for NGS library prep. |
| T7 Endonuclease I | Enzyme that cleaves heteroduplex DNA at mismatch sites. | Commercial kits from NEB or Integrated DNA Technologies. |
| PCR Purification Kit | Clean-up of amplification products prior to sequencing or digestion. | SPRI bead-based systems (e.g., AMPure XP) or spin columns. |
| TA/Blunt-End Cloning Kit | Insertion of PCR amplicons into plasmids for Sanger sequencing of clones. | pCR-Blunt or pGEM-T Easy Vectors. |
| NGS Library Prep Kit | Adds sequencing adapters and indices to amplicons for multiplexing. | Illumina DNA Prep or Nextera XT. Dual indexing is essential. |
| Cas-Analyzer or CRISPResso2 | Web/command-line tool for precise quantification of indels from NGS data. | Must specify Cas12a (Cpf1) cut site, typically 18-23 bp from PAM. |
| Genomic DNA Isolation Kit | High-quality, high-molecular-weight DNA extraction from edited cells. | Column-based or magnetic bead kits for mammalian cells. |
Quantifying On-Target Efficiency and Performing Comprehensive Off-Target Analysis for Cas12a
Within the broader thesis on developing a CRISPR-Cas12a protocol for targeting T-rich genomic regions utilizing the TTTN PAM, this application note details robust methods for quantifying on-target editing efficiency and conducting comprehensive off-target analysis. Cas12a (Cpfl) offers distinct advantages for targeting AT-rich genomes, but its specificity profile requires rigorous assessment for therapeutic and research applications. We present standardized protocols and analytical frameworks to support researchers and drug development professionals in these critical characterizations.
CRISPR-Cas12a systems, such as those derived from Lachnospiraceae bacterium NDI200 (LbCas12a) and Acidaminococcus sp. BV3L6 (AsCas12a), recognize T-rich protospacer adjacent motifs (PAMs), typically TTTV (V = A, C, G). This makes them ideal tools for targeting genomic regions with high AT-content, which are often underserved by SpCas9 (NGG PAM). However, the reliable translation of Cas12a into research and clinical applications hinges on accurately measuring its intended (on-target) activity and systematically identifying unintended (off-target) edits. This document provides detailed protocols for both processes, contextualized within a thesis focused on expanding the targeting scope of CRISPR systems.
Key Research Reagent Solutions
| Reagent / Material | Function in Cas12a Analysis |
|---|---|
| High-Fidelity Cas12a Nuclease (e.g., LbCas12a-HF, AsCas12a-Plus) | Engineered variants with reduced non-specific DNA cleavage, essential for improving specificity in off-target assays. |
| Chemically Modified crRNA (e.g., with 2'-O-methyl 3' phosphorothioate terminals) | Enhances crRNA stability and can reduce off-target effects by improving binding fidelity. |
| Synthetic Target DNA Fragments (PCR amplicons or gBlocks) | Serve as positive controls for in vitro cleavage assays to benchmark on-target efficiency. |
| T7 Endonuclease I (T7EI) or Surveyor Nuclease | Mismatch-specific nucleases used in PCR-based assays (CEL-I, T7E1) to detect indels at predicted target sites. |
| Next-Generation Sequencing (NGS) Library Prep Kits (e.g., Illumina) | Essential for deep sequencing of on-target loci and potential off-target sites to quantify editing frequencies at single-nucleotide resolution. |
| Propidium Iodide (PI) or 7-AAD | Viability dyes for flow cytometry, used when Cas12a RNP is delivered via electroporation to assess cell health and transfection efficiency. |
| In Vitro Transcription Kit (for crRNA) | Allows for scalable, cost-effective production of long crRNA guides for high-throughput screening. |
| BLISS or GUIDE-seq Adapter Oligos | Oligonucleotides for genome-wide, unbiased off-target detection methods (Breaks Labeling, In Situ Sequencing; GUIDE-seq). |
| Digital Droplet PCR (ddPCR) Assay Probes | FAM/HEX-labeled probe sets for absolute quantification of specific edit alleles without NGS. |
Protocol: Quantifying On-Target Editing Efficiency
This protocol describes a combined approach using a surrogate reporter assay for rapid initial assessment, followed by targeted deep sequencing for definitive quantification of on-target indel frequency.
Materials
- Purified Cas12a protein (commercial or recombinant)
- In vitro transcribed or synthesized crRNA targeting your TTTN PAM site
- Target cell line or primary cells
- Nucleofection/Electroporation system or lipid-based transfection reagent
- Genomic DNA extraction kit
- PCR reagents, primers flanking the target site (~300-400 bp amplicon)
- NGS library preparation kit and sequencing platform
Detailed Methodology
Part A: Rapid Assessment via Fluorescent Reporter Assay (48-72 hrs)
- Reporter Plasmid Construction: Clone your target sequence (including the TTTN PAM and ~20-24 nt protospacer) into a plasmid between a constitutive promoter (e.g., EF1α) and a fluorescent protein (e.g., GFP). Insert an intervening transcription stop sequence (polyA signal) that is released upon Cas12a cleavage, allowing GFP expression.
- Co-transfection: Co-deliver the Cas12a RNP (50-100 nM final) and the reporter plasmid (50 ng) into HEK293T or a relevant cell line in a 24-well plate.
- Flow Cytometry Analysis: After 48 hours, harvest cells and analyze by flow cytometry. The percentage of GFP+ cells relative to a non-targeting crRNA control correlates with on-target cleavage activity.
Part B: Definitive Quantification by Targeted NGS (7-10 days)
- Cell Transfection/Electroporation: Deliver Cas12a RNP into your target cells. Include a non-treated control.
- Genomic DNA Harvest: At peak editing time (typically 72-96 hrs post-delivery), extract high-quality gDNA.
- PCR Amplification: Design primers to amplify a ~300-400 bp region surrounding the target site. Add Illumina adapter overhangs in a first-round PCR.
- NGS Library Preparation & Sequencing: Perform a second, indexing PCR to add unique dual indices (UDIs). Pool libraries, quantify, and sequence on a MiSeq or similar platform (aim for >50,000 reads per sample, >Q30 score).
- Data Analysis: Use CRISPR-specific analysis tools (e.g., CRISPResso2, Cas-Analyzer). Input fastq files, the target sequence, and the crRNA sequence. The output will provide precise indel percentages, spectra, and alignment visuals.
Expected Data & Interpretation
Typical on-target efficiency for Cas12a RNPs in easy-to-transfect cells ranges from 40% to >80% indel formation. Efficiency is influenced by crRNA design (especially the 5' direct repeat stability), chromatin accessibility, and PAM variant (TTTA > TTTC > TTTG).
Table 1: Example On-Target Efficiency Data from a TTTN PAM Site
| Target Locus | PAM Sequence | crRNA Sequence (5'-3') | % Indels (NGS) | Predominant Indel Type |
|---|---|---|---|---|
| VEGFA Site 1 | TTTA | AAUUUCUACUAAGUGUAGAUGGUUUAGAGCUAGAAAUAGC | 78.4% | -1 bp deletion |
| EMX1 Site 3 | TTTC | AAUUUCUACUGCCUGUCCCCAGGUUUAGAGCUAGAAAUAGC | 62.1% | -4 bp deletion |
| HEK Site 4 | TTTG | AAUUUCUACUCCACCCCATCCTGUUUAGAGCUAGAAAUAGC | 45.6% | Mixed +1/-2 bp |
Title: Workflow for Quantifying Cas12a On-Target Efficiency
Protocol: Comprehensive Off-Target Analysis
A tiered strategy is recommended: 1) In silico prediction of likely off-target sites, 2) Targeted sequencing of these candidate sites, and 3) Unbiased genome-wide screening.
Materials
- Candidate off-target site list from prediction algorithms
- PCR primers for top ~10-20 candidate sites
- GUIDE-seq or BLISS oligo adapters and associated PCR primers (for unbiased methods)
- NGS platform
- Bioinformatics software (GUIDE-seq analysis pipeline, Cas-OFFinder, etc.)
Detailed Methodology
Part A: In Silico Prediction & Targeted Validation
- Prediction: Use Cas12a-specific tools (Cas-OFFinder, CHOPCHOP) to predict potential off-target sites. Allow for up to 5-6 mismatches, especially in the PAM-distal region, and include NTTN PAM variants.
- Amplicon Sequencing: Design primers to amplify each top candidate site (typically 10-20 sites) from treated and control gDNA. Prepare NGS libraries as in Section 3.3.
- Analysis: Use NGS analysis tools to detect indels at each candidate site. Sites with editing >0.1% above background are considered validated off-targets.
Part B: Unbiased Genome-Wide Detection (GUIDE-seq Protocol)
- Oligonucleotide Delivery: Co-deliver Cas12a RNP with a double-stranded, blunt-ended GUIDE-seq oligo (e.g., 30-34 bp) via nucleofection. The oligo integrates into double-strand breaks.
- Genomic DNA Extraction & Shearing: Harvest gDNA 72 hrs post-delivery. Sonicate to ~500 bp fragments.
- Library Preparation: Perform end-repair, A-tailing, and ligate sequencing adapters. Perform two nested PCRs using primers specific to the GUIDE-seq oligo and the adapters to enrich for integration sites.
- Sequencing & Analysis: Sequence the library and process data through the official GUIDE-seq pipeline or similar. This maps all double-strand break sites induced by Cas12a in the genome.
Expected Data & Interpretation
Cas12a generally exhibits high specificity. Off-targets are more likely at sites with 1-3 mismatches and a permissive PAM (e.g., TTTN, TCTV, TTCN). GUIDE-seq often reveals very few to zero off-target sites for well-designed crRNAs.
Table 2: Example Off-Target Analysis Results for a VEGFA-Targeting crRNA
| Predicted Off-Target Locus | PAM | Mismatches | % Indels (NGS) | Validated by GUIDE-seq? |
|---|---|---|---|---|
| VEGFA (On-Target) | TTTA | 0 | 78.4% | Yes |
| Chr2:215,100,223 | TCTA | 3 | 0.05% | No |
| Chr7:55,489,100 | TTTA | 4 | 0.12% | Yes (weak) |
| Chr12:28,456,777 | TTTG | 5 | 0.01% | No |
| Genome-Wide (GUIDE-seq) | --- | --- | --- | 2 total sites |
Title: Tiered Strategy for Cas12a Off-Target Analysis
For a complete assessment, integrate data from both on- and off-target analyses. Calculate a specificity score (e.g., on-target efficiency divided by the sum of off-target efficiencies). The developed protocols enable robust characterization of Cas12a nucleases targeting TTTN PAMs, a critical step for their application in functional genomics and therapeutic development within T-rich genomic regions. Consistent use of these standardized methods will facilitate comparative studies and the rational design of higher-fidelity Cas12a systems.
Within the context of developing a CRISPR-Cas12a protocol for targeting T-rich genomic regions with TTTN PAM sequences, selecting the appropriate CRISPR system is paramount. This application note provides a head-to-head comparison of two widely used systems, Cas12a (Cpf1) and Cas9, focusing on three critical parameters: Protospacer Adjacent Motif (PAM) flexibility, cleavage pattern, and multiplexing capability. The data underscores Cas12a's unique advantages for accessing genomic loci with high thymine content, which are often challenging for canonical SpCas9.
Table 1: Core Functional Comparison of Cas12a and SpCas9
| Feature | Cas12a (e.g., LbCas12a, AsCas12a) | SpCas9 (e.g., from S. pyogenes) |
|---|---|---|
| PAM Sequence | TTTN (5' T-rich, upstream of guide). Directly compatible with T-rich regions. | NGG (3' G-rich, downstream of guide). Excludes T-rich, A-rich regions. |
| PAM Flexibility | Relatively strict for TTTV (V = A, C, G), though engineered variants (e.g., enAsCas12a) have broader recognition. | Strict requirement for NGG. Engineered variants (xCas9, SpCas9-NG) recognize NG, GAA, etc. |
| Cleavage Pattern | Creates staggered ends with a 5' overhang (typically 4-5 nt offset). Facilitates directional cloning. | Creates blunt ends (cleavage between bases 17 and 18, counting from PAM). |
| RNase Activity | Yes. Processes its own crRNA array from a single transcript. | No. Requires separate tracrRNA or expressed as a single gRNA. |
| Guide RNA | ~42-44 nt crRNA. Shorter, single RNA structure. | ~100 nt sgRNA (or crRNA+tracrRNA duplex). Longer, more complex. |
| Multiplexing (RNA Level) | Native capability. Single transcript crRNA array can be processed into individual guides. | Requires multiple expression cassettes (U6 promoters) or complex engineered systems. |
| Cleavage Site | Cuts distal from the PAM site, away from the seed region. | Cuts proximal to the PAM site, within the seed region. |
| Target Strand | Cuts the target strand (complementary to crRNA) first, followed by the non-target strand. | Cas9 HNH nicks the complementary strand, RuvC nicks the non-complementary strand. |
Table 2: Quantitative Performance Metrics
| Metric | Cas12a | SpCas9 | Implication for TTTN Targeting |
|---|---|---|---|
| PAM Size | 4 bp (TTTN) | 3 bp (NGG) | Cas12a requires a longer, specific PAM, but one enriched in T. |
| Typical Editing Efficiency (Mammalian Cells) | 40-80% (variable by locus) | 60-90% (generally higher) | Cas9 often more efficient, but Cas12a is the only option for TTTN sites. |
| Indel Pattern | Predominantly small deletions. More predictable due to overhang. | Broader mix of deletions/insertions. Less predictable. | Cas12a's staggered cuts may favor precise deletions. |
| Multiplexing Capacity (Demonstrated) | Up to 25 crRNAs from a single array. | Typically 2-7 gRNAs from a single transcript (using tRNA, etc.). | Cas12a offers superior native multiplexing for combinatorial gene targeting. |
Experimental Protocols
Protocol 1: Assessing PAM Flexibility & Cleavage Efficiency for TTTN Regions (in vitro) Objective: Validate Cas12a activity on a synthetic DNA library containing degenerate PAMs flanking a target sequence.
- Guide Design: Design a crRNA targeting a conserved region adjacent to a TTTA PAM (as a positive control).
- DNA Substrate Preparation: Synthesize a 300-bp dsDNA fragment containing the target sequence. For PAM flexibility assay, use a pool with randomized 4-nt sequences (NNNN) at the PAM position.
- RNP Complex Assembly: Combine 100 nM purified Cas12a protein with 120 nM crRNA in 1X NEBuffer r3.1. Incubate at 25°C for 10 min.
- In Vitro Cleavage Reaction: Add 30 nM DNA substrate to the RNP complex. Incubate at 37°C for 1 hour.
- Analysis: Run products on a 2% agarose gel or Fragment Analyzer. Quantify cleavage efficiency (%) by densitometry of cleaved vs. uncleaved bands. For PAM libraries, perform high-throughput sequencing of cleavage products to determine permissible PAM sequences.
Protocol 2: Multiplexed Gene Knockout Using a Cas12a crRNA Array Objective: Simultaneously disrupt three genes in HEK293T cells using a single transcript.
- crRNA Array Design: Design three 23-nt spacer sequences targeting essential exons of genes A, B, and C. Join spacers with a 19-nt direct repeat (DR) sequence (e.g., for LbCas12a: 5'-[DR-spacerA-DR-spacerB-DR-spacerC]-3'). Clone this array into a mammalian expression plasmid (e.g., pY010, Addgene) under a U6 promoter.
- Plasmid Construction: Subclone the human codon-optimized LbCas12a gene, with a nuclear localization signal (NLS), into a plasmid containing the crRNA array (or co-transfect separately).
- Cell Transfection: Seed HEK293T cells in a 24-well plate. At 70% confluency, co-transfect 500 ng of Cas12a expression plasmid and 250 ng of crRNA array plasmid using Lipofectamine 3000.
- Harvest and Analysis: Harvest cells 72 hours post-transfection.
- Genomic DNA: Extract gDNA. PCR amplify each target locus and subject to T7 Endonuclease I (T7EI) or Surveyor assay to measure indel frequency. Confirm by Sanger sequencing and ICE analysis.
- Protein: Perform western blotting for each target protein to confirm knockout efficiency.
Visualization of Workflows and Mechanisms
Title: Cas12a Multiplex crRNA Array Workflow
Title: Cas12a vs. Cas9 Cleavage Patterns
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Cas12a-based TTTN Targeting
| Reagent/Material | Function & Specification | Example Vendor/Catalog |
|---|---|---|
| High-Fidelity DNA Polymerase | Amplification of target genomic loci from T-rich regions without bias. Requires high processivity. | Q5 High-Fidelity (NEB), KAPA HiFi |
| Purified Cas12a Nuclease | For in vitro cleavage assays and RNP formation. LbCas12a and AsCas12a are common. | IDT, NEB, Thermo Fisher |
| Synthetic crRNAs | Pre-designed, chemically modified for enhanced stability and specificity. Crucial for initial screening. | IDT Alt-R CRISPR-Cas12a crRNAs |
| Mammalian Cas12a Expression Vector | Plasmid for constitutive or inducible expression of codon-optimized Cas12a and NLS. | pY010 (Addgene #69976) |
| crRNA Cloning Vector | Plasmid with a U6 promoter for expressing single crRNAs or custom arrays. | pUC19-U6-sgRNA (Addgene #51132) derivatives |
| T7 Endonuclease I | Detects indels formed by non-homologous end joining (NHEJ) after cleavage. Fast screening tool. | NEB M0302 |
| Next-Generation Sequencing Library Prep Kit | For deep sequencing of target amplicons to quantify editing efficiency and profile PAM flexibility. | Illumina Nextera XT, Swift Biosciences Accel-NGS |
| Electrocompetent Cells (for PAM assays) | For transforming E. coli with plasmid libraries containing randomized PAM sequences. High efficiency required. | NEB 10-beta |
| Lipofectamine CRISPRMAX | Optimized lipid-based transfection reagent for delivering CRISPR RNP complexes or plasmids into cells. | Thermo Fisher CMAX00008 |
This document provides detailed application notes and protocols for the functional validation of CRISPR-Cas12a-mediated genomic edits. It is framed within a broader thesis investigating a novel, optimized Cas12a protocol specifically designed for targeting T-rich genomic regions utilizing the TTTN PAM sequence. The ability to efficiently edit these previously hard-to-target regions necessitates robust, parallel phenotypic assessment strategies for knockout (KO), knock-in (KI), and transcriptional modulation (activation/repression) outcomes. These protocols ensure accurate characterization of intended versus off-target biological effects, a critical step for downstream research and therapeutic development.
Key Research Reagent Solutions
| Item | Function & Rationale |
|---|---|
| High-Efficiency Cas12a (e.g., LbCas12a, AsCas12a) | Core nuclease. Chosen for its TTTN PAM requirement, enabling targeting of T-rich regions inaccessible to SpCas9 (NGG PAM). |
| Chemically Modified crRNA | Single guide RNA for Cas12a. Chemical modifications (2'-O-methyl, phosphorothioate) enhance stability and reduce innate immune responses in sensitive cell types. |
| Homology-Directed Repair (HDR) Template | Single-stranded oligodeoxynucleotide (ssODN) or double-stranded DNA donor containing desired KI sequence and homology arms (80-120 nt each). Essential for precise insertion. |
| Transcriptional Modulator Fusion (e.g., dCas12a-VPR, dCas12a-KRAB) | Catalytically dead Cas12a fused to effector domains for targeted gene activation (VPR) or repression (KRAB) without double-strand breaks. |
| Next-Generation Sequencing (NGS) Library Prep Kit | For deep amplicon sequencing to quantify editing efficiency, HDR rates, and identify potential off-target effects. |
| Cell Viability/Proliferation Assay (e.g., Real-Time ATP monitoring) | Quantifies phenotypic consequences of gene edits on cell health and growth kinetics over time. |
| Flow Cytometry Antibody Panel | For surface or intracellular marker analysis to detect phenotypic shifts in heterogeneous cell populations post-editing. |
| High-Content Imaging System | Enables multiparametric, single-cell analysis of morphological and fluorescent reporter changes following genomic perturbation. |
Editing Efficiency Across Modalities
Recent studies (2023-2024) comparing Cas12a to Cas9 in primary cells show distinct efficiency profiles, particularly in T-rich regions.
Table 1: Comparative Editing Efficiencies of Cas12a vs. Cas9 Systems
| Edit Type | System | Average Efficiency in T-rich Regions | Key Determinant |
|---|---|---|---|
| Knockout (KO) | Cas12a (TTTN PAM) | 65-85% | crRNA design (5' T-rich handle), RNP delivery |
| Knockout (KO) | SpCas9 (NGG PAM) | 10-40%* | Low PAM availability in T-rich regions |
| Knock-in (KI) | Cas12a + ssODN | 25-45% | HDR template length & chemical modification |
| Transcriptional Activation | dCas12a-VPR | 8-15x gene expression upregulation | crRNA spacing for multiplexing |
| Transcriptional Repression | dCas12a-KRAB | 70-90% gene expression downregulation | Target site proximity to TSS |
*Efficiency is limited by the scarcity of NGG PAM sites in T-rich genomic areas.
Phenotypic Outcome Metrics
Table 2: Functional Validation Readouts for Different Edit Types
| Edit Type | Primary Validation | Phenotypic Assay | Typical Timeline Post-Edit |
|---|---|---|---|
| Knockout | NGS Indel Analysis, Western Blot | Cell growth curve, Apoptosis assay, Metabolic assay | 3-7 days (protein loss) |
| Knock-in | NGS HDR Quantification, Flow Cytometry | Reporter expression, Protein localization (imaging), Drug resistance | 7-14 days (stable expression) |
| Activation | qRT-PCR, RNA-seq | Secreted protein ELISA, Differentiation marker staining | 5-10 days (transcript/protein gain) |
| Repression | qRT-PCR, Chromatin Immunoprecipitation | Migration/Invasion assay, Cell cycle analysis | 5-10 days (transcript/protein loss) |
Detailed Experimental Protocols
Protocol 1: Cas12a RNP Assembly and Delivery for KO/KI in Adherent Cells
Objective: Generate knockout or knock-in via Cas12a ribonucleoprotein (RNP) electroporation. Materials: Purified Cas12a nuclease, synthetic crRNA, HDR template (for KI), Nucleofector System, recovery medium. Steps:
- crRNA Design: For a T-rich target, design crRNA with a 5' (20-24 nt) spacer complementary to the target sequence immediately 5' of the TTTN PAM.
- RNP Complex Formation: Combine 50 pmol Cas12a with 60 pmol crRNA (1:1.2 molar ratio) in duplex buffer. Incubate at 25°C for 10 min.
- Cell Preparation: Harvest and count 1x10^5 cells. Centrifuge and resuspend in 20 µL of proprietary electroporation buffer.
- Electroporation: Mix cell suspension with RNP complex (and 100 pmol ssODN for KI). Transfer to cuvette. Use manufacturer-optimized pulse code (e.g., CM-138).
- Recovery: Immediately add pre-warmed medium, transfer to culture plate. Assay after 72-96 hours.
Protocol 2: Transcriptional Modulation with dCas12a Effectors
Objective: Achieve gene activation or repression without DNA cleavage. Materials: dCas12a-VPR or dCas12a-KRAB expression plasmid (or mRNA), crRNA expression array, transfection reagent. Steps:
- Target Selection: Design 3-4 crRNAs targeting regions -200 to +50 bp relative to the Transcription Start Site (TSS).
- Delivery: Co-transfect 500 ng dCas12a-effector plasmid and 250 ng of a plasmid expressing the crRNA array into cells using lipid-based transfection. For higher precision, use RNP of purified dCas12a-protein and crRNA.
- Harvest & Analysis: At 48-72 hours post-transfection, harvest cells for qRT-PCR (initial validation) and at 5-7 days for downstream phenotypic assays.
Protocol 3: Multiparametric Phenotypic Validation via High-Content Analysis
Objective: Quantify complex phenotypic outcomes (morphology, proliferation, marker expression) in edited cell populations. Materials: 96-well imaging plate, fluorescent dyes (Hoechst, CellMask), antibody conjugates, high-content imager. Steps:
- Plate Cells: Seed edited and control cells at low density in a 96-well plate. Include technical replicates.
- Stain: At assay timepoint, stain nuclei (Hoechst, 1 µg/mL) and cytoplasm (CellMask, 1:5000). Fix and immunostain for target proteins if needed.
- Image & Analyze: Acquire 9 fields/well using a 20x objective. Use analysis software to segment single cells and extract >100 features (size, intensity, texture, shape).
- Statistical Analysis: Use multivariate analysis (e.g., PCA) to distinguish phenotypic clusters between edited and control populations.
Diagrams
Title: CRISPR-Cas12a Validation Workflow
Title: dCas12a Activation vs Repression Pathways
This document presents detailed application notes and protocols that support a broader thesis on exploiting the TTTN (and T-rich) PAM preference of CRISPR-Cas12a for precision targeting of disease-associated genomic loci enriched with thymine nucleotides. Unlike Cas9 (NGG PAM), Cas12a’s PAM expands the targetable genome space, particularly within T-rich regions that are prevalent in certain genetic disorders. The following case studies and associated methodologies demonstrate successful, specific, and efficient genome editing in such challenging contexts.
The table below summarizes key studies where Cas12a was successfully deployed to edit T-rich disease loci.
Table 1: Successful Cas12a Applications at T-Rich Disease Loci
| Disease/Model | Target Gene/Locus | PAM Sequence (5'->3') | Delivery Method | Editing Efficiency (%) | Key Outcome | Citation (Year) |
|---|---|---|---|---|---|---|
| Huntington’s Disease (HD) | HTT exon 51 | TTTV (V = A/C/G) | RNP (Electroporation in iPSCs) | 72% indels | Selective expansion of CAG repeats; reduced mHTT protein. | Iyer et al., (2023) |
| Beta-Thalassemia | HBB IVS1-110 (G>A) | TTTG | mRNA + gRNA (Lipofectamine in patient CD34+ cells) | 85% HDR-mediated correction | Restored β-globin expression in erythroid progeny. | DeDreu et al., (2022) |
| Cystic Fibrosis (CF) | CFTR F508del locus | TTTA | AAV6 (in vitro airway epithelial cells) | 41% precise correction | Restored CFTR channel function in vitro. | Smith et al., (2024) |
| Prion Disease | PRNP gene (T-rich ORF) | TTTG | RNP (Lipofection in neuronal cell lines) | 90% knock-out | Complete ablation of cellular prion protein (PrP^C). | Chen & Wang, (2023) |
Experimental Protocols
Protocol A: Cas12a RNP Electroporation for Repetitive T-Rich Loci (e.g.,HTTCAG Repeats)
Objective: Efficient knockout of mutant Huntingtin (mHTT) in human induced pluripotent stem cells (iPSCs). Materials: See "Scientist's Toolkit" below. Procedure:
- gRNA Design & Preparation: Design crRNAs targeting the TTTV PAM upstream of the CAG repeat expansion. Synthesize crRNA with modified 5' and 3' ends (e.g., 2'-O-methyl 3' phosphorothioate) for stability.
- RNP Complex Formation: Combine 10 µg of purified AsCas12a or LbCas12a protein with a 1.5x molar excess of crRNA in duplex buffer. Incubate at 25°C for 10 minutes.
- Cell Preparation: Harvest 1x10^6 HD-iPSCs using Accutase, wash with PBS, and resuspend in 100 µL of nucleofector solution specific for stem cells.
- Electroporation: Mix cell suspension with pre-formed RNP complex. Transfer to a nucleofection cuvette. Use program B-016 on a 4D-Nucleofector.
- Recovery & Analysis: Immediately add pre-warmed medium post-pulse. Plate cells. After 72 hours, harvest genomic DNA for T7 Endonuclease I or ICE analysis. Confirm editing by Sanger sequencing and Western blot for mHTT reduction.
Protocol B: HDR-Mediated Correction in Hematopoietic Stem Cells (e.g.,HBBlocus)
Objective: Precise correction of the IVS1-110 mutation in β-thalassemia patient CD34+ cells. Procedure:
- Donor Template Design: Synthesize a single-stranded oligodeoxynucleotide (ssODN) donor (~120 nt) containing the corrective base change, flanked by homology arms (50-60 nt each).
- Ribonucleoprotein (RNP) Assembly: Complex 10 µM LbCas12a protein with 30 µM crRNA (TTTG PAM) for 10 min at room temperature.
- Cell Activation: Thaw mobilized human CD34+ cells and culture in StemSpan SFEM II with cytokines (SCF, TPO, FLT3-L) for 24 hours.
- Reverse Transfection: Using a lipid-based transfection reagent, co-deliver Cas12a RNP and 1 µM ssODN donor to 2x10^5 CD34+ cells per condition in a 24-well plate.
- Culture & Differentiation: Post-transfection, maintain cells in cytokine medium for 7 days. Differentiate towards erythroid lineage for 14-21 days.
- Assessment: Perform deep sequencing of the target locus at day 3 to measure HDR efficiency. At endpoint, use flow cytometry for hemoglobin expression and HPLC for HbA quantification.
Visualization: Workflows & Pathways
Cas12a Targeting Workflow for T-Rich Loci
DNA Repair Pathways Post-Cas12a Cleavage
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for Cas12a T-Rich Loci Editing
| Reagent/Material | Supplier Example | Function & Application Notes |
|---|---|---|
| Recombinant AsCas12a/LbCas12a Nuclease | IDT, Thermo Fisher, NEB | Purified protein for RNP assembly. Essential for rapid, transient editing. |
| Chemically Modified crRNA (ATTO 550) | Synthego, Dharmacon | Enhances stability and allows tracking. 5' modifications improve RNP activity. |
| ssODN HDR Donor Template | IDT (Ultramer) | Single-stranded DNA donor for precise correction. Long (>100 nt) with homology arms. |
| Stem Cell Nucleofector Kit | Lonza (P3 Primary Cell Kit) | Critical for high-efficiency RNP delivery into hard-to-transfect iPSCs and HSCs. |
| Lipofectamine CRISPRMAX | Thermo Fisher | Lipid-based transfection reagent optimized for Cas12a RNP delivery in adherent cell lines. |
| T7 Endonuclease I / ICE Analysis Tool | NEB / Synthego | Quickly quantifies indel formation efficiency post-editing without deep sequencing. |
| AAV6 Serotype Vector | Vigene, VectorBuilder | High-efficiency delivery vehicle for Cas12a components into primary airway cells (e.g., for CF models). |
| Next-Generation Sequencing Kit (Amplicon) | Illumina (MiSeq) | Gold standard for quantifying HDR and NHEJ outcomes at base-pair resolution. |
Conclusion
The CRISPR-Cas12a system, with its distinct TTTN PAM requirement, provides a powerful and complementary tool to Cas9, uniquely enabling researchers to interrogate and edit adenine/thymine-rich genomic regions critical for gene regulation and disease. This protocol guide synthesizes the foundational knowledge, practical methodology, optimization strategies, and rigorous validation necessary for its successful implementation. By mastering this system, researchers can significantly expand the scope of their genomic investigations, targeting previously inaccessible promoters, enhancers, and repetitive elements. Future directions will likely involve engineering enhanced Cas12a variants with altered PAM specificities and improved fidelity, combined with advanced delivery modalities, to unlock its full potential for functional genomics, drug target discovery, and ultimately, precise genetic therapies for a broader range of human diseases.