Molecular Choreography: Unveiling the Conformational Dynamics of Cas9 During gRNA Binding and Target Recognition
This article provides a comprehensive analysis of the intricate structural rearrangements undergone by the CRISPR-Cas9 enzyme, from its apo state through guide RNA (gRNA) binding to ultimate target DNA recognition.
Molecular Choreography: Unveiling the Conformational Dynamics of Cas9 During gRNA Binding and Target Recognition
Abstract
This article provides a comprehensive analysis of the intricate structural rearrangements undergone by the CRISPR-Cas9 enzyme, from its apo state through guide RNA (gRNA) binding to ultimate target DNA recognition. Designed for researchers, scientists, and drug development professionals, it explores foundational structural biology, current experimental methodologies for observing these changes, common challenges in interpreting conformational data, and validation techniques. We synthesize findings from recent high-resolution studies to illustrate how these dynamic transitions underpin Cas9's precision, inform engineering efforts for improved specificity and efficiency, and open new avenues for therapeutic genome editing applications.
From Inert to Active: The Structural Journey of Cas9 from Apo State to Functional Complex
This whitepaper details the architecture of the apo, or unprimed, state of the CRISPR-associated protein 9 (Cas9) from Streptococcus pyogenes (SpCas9). It serves as a foundational reference within the broader thesis investigating the profound conformational changes Cas9 undergoes upon guide RNA (gRNA) binding and subsequent target DNA recognition—a critical process for genome editing applications and therapeutic development.
Structural Domains of Apo-Cas9
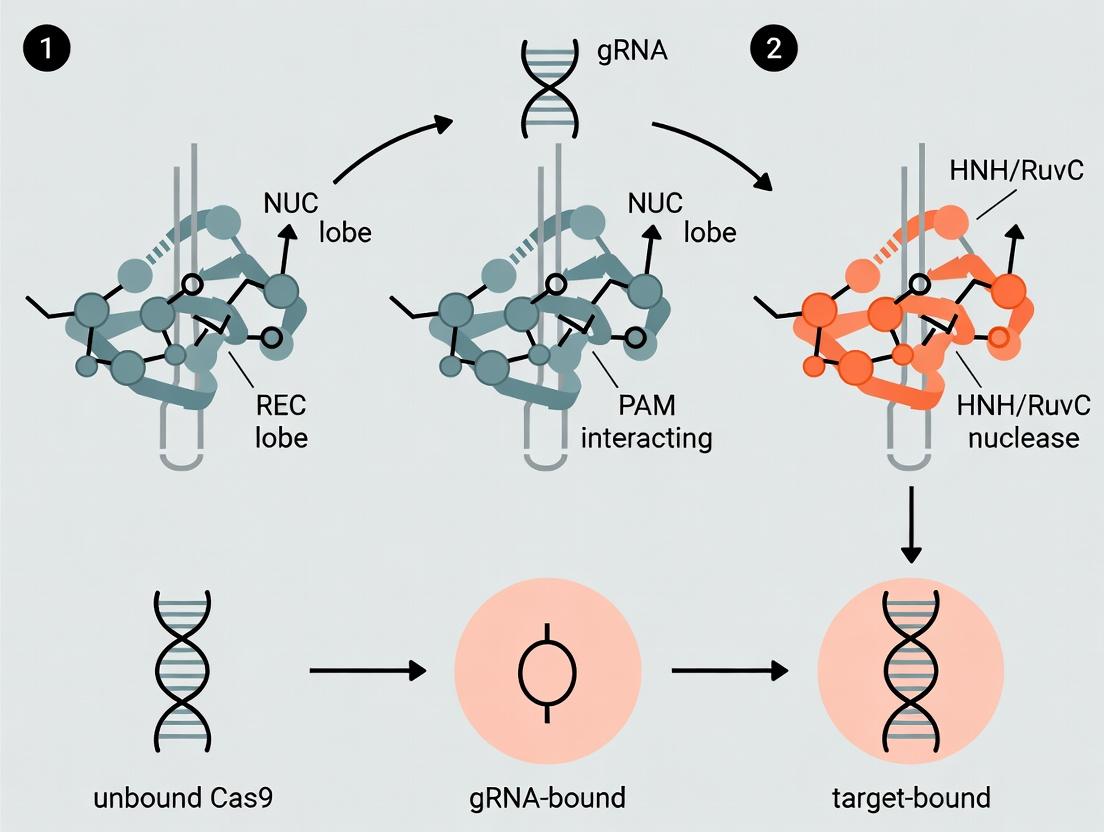
In its apo state, Cas9 exists in an autoinhibited conformation, unable to bind DNA. The structure is divided into several lobes and key domains:
- REC Lobe (Recognition Lobe): Predominantly α-helical, this lobe is responsible for gRNA and target DNA binding. In the apo state, it is disorganized and flexible.
- REC I, II, III: Subdomains involved in gRNA and DNA interactions.
- Bridge Helix (BH): A long α-helix connecting the REC and NUC lobes.
- NUC Lobe (Nuclease Lobe): Contains the catalytic domains and DNA interaction interfaces.
- RuvC Domain: Cleaves the non-target (complementary) DNA strand. In apo-Cas9, its active site is incomplete.
- HNH Domain: Cleaves the target DNA strand. In the apo state, it is positioned distantly from the RuvC domain and the DNA-binding channel.
- PI (PAM-Interacting) Domain: Critical for recognizing the protospacer adjacent motif (PAM). It is partially occluded in the apo conformation.
- Linker Regions: Flexible segments that allow large-scale domain movements upon activation.
Key Quantitative Structural Parameters of Apo-SpCas9
Table 1: Primary structural features of apo-Cas9 (PDB ID: 4CMP)
| Parameter | Value / State | Functional Implication |
|---|---|---|
| Overall Conformation | Closed, autoinhibited | DNA binding cleft is inaccessible |
| HNH Domain Position | ~40 Å from catalytic site | Catalytically inactive; requires large rotation |
| RuvC Active Site | Disordered/incomplete | Unable to coordinate catalytic metal ions |
| PAM Interaction Interface | Partially obstructed | Low affinity for DNA; cannot scan for PAM |
| gRNA Binding Channel | Collapsed | Very low affinity for nucleic acids |
| Bridge Helix Conformation | Kinked and rigid | Cannot facilitate DNA strand separation |
Experimental Protocols for Apo-Cas9 Structure Determination
Protocol 1: X-ray Crystallography of Apo-SpCas9
This protocol was used to solve the first high-resolution structure (4CMP).
Protein Expression & Purification:
- Express N-terminal 6xHis-tagged SpCas9 in E. coli BL21(DE3) cells.
- Induce expression with 0.5 mM IPTG at 18°C for 16 hours.
- Lyse cells in buffer (20 mM HEPES pH 7.5, 500 mM KCl, 10% glycerol, 20 mM imidazole).
- Purify via Ni-NTA affinity chromatography, followed by heparin affinity and size-exclusion chromatography (Superdex 200).
Crystallization:
- Concentrate purified apo-Cas9 to 10 mg/mL in 20 mM HEPES pH 7.5, 150 mM KCl, 1 mM TCEP.
- Use the sitting-drop vapor diffusion method at 20°C.
- Mix 0.2 μL protein with 0.2 μL reservoir solution (0.1 M Tris pH 8.5, 21% PEG 3350).
- Crystals appear within 3-7 days.
Data Collection & Structure Solution:
- Cryo-protect crystals in reservoir solution supplemented with 20% ethylene glycol.
- Collect X-ray diffraction data at a synchrotron source (e.g., 0.979 Å wavelength).
- Solve the structure by molecular replacement using a homologous Cas9 model (e.g., from Francisella novicida).
Protocol 2: Cryo-Electron Microscopy (Cryo-EM) for Conformational Analysis
Used to capture the dynamic flexibility of apo-Cas9.
Sample Preparation:
- Apply 3 μL of 0.5 mg/mL apo-Cas9 to a freshly glow-discharged Quantifoil grid.
- Blot for 3-4 seconds and plunge-freeze in liquid ethane using a Vitrobot (4°C, 100% humidity).
Data Acquisition:
- Collect micrographs on a 300 keV cryo-TEM with a K3 direct electron detector.
- Use a nominal magnification of 81,000x (pixel size 1.06 Å).
- Collect a dose-fractionated dataset with a total dose of ~50 e⁻/Ų over 40 frames.
Image Processing:
- Perform motion correction and dose-weighting (e.g., using MotionCor2).
- Generate initial models via ab initio reconstruction in cryoSPARC.
- Conduct heterogeneous refinement to separate and classify distinct conformational states of the flexible REC and NUC lobes.
Conformational Change Pathway from Apo to Holoenzyme
Diagram Title: Conformational Activation Pathway of Cas9
The Scientist's Toolkit: Essential Research Reagents
Table 2: Key reagents and materials for studying apo-Cas9 architecture
| Reagent / Material | Function / Purpose | Example Product (Supplier) |
|---|---|---|
| Recombinant SpCas9 Protein | High-purity, nuclease-free protein for structural and biochemical studies. | SpyCas9 Nuclease (Thermo Fisher), Purified SpCas9 (NEB) |
| Size-Exclusion Chromatography (SEC) Column | Final polishing step to obtain monodisperse, aggregation-free protein for crystallization/cryo-EM. | Superdex 200 Increase 10/300 GL (Cytiva) |
| Cryo-EM Grids | Specimen support for vitrified sample preparation in cryo-EM. | Quantifoil R 1.2/1.3 300 mesh Au grids |
| Heparin Sepharose Resin | Affinity purification step that exploits Cas9's positive charge and nucleic acid-mimetic properties. | Heparin Sepharose 6 Fast Flow (Cytiva) |
| TEV Protease | For cleaving affinity tags (e.g., His-tag) after purification to avoid interference with structural studies. | AcTEV Protease (Thermo Fisher) |
| PEG 3350 | Common precipitant in crystallization screens for large, multi-domain proteins like Cas9. | Polyethylene glycol 3350 (Hampton Research) |
| TCEP-HCl | Reducing agent to maintain cysteines in reduced state and prevent disulfide-mediated aggregation. | Tris(2-carboxyethyl)phosphine hydrochloride (Sigma-Aldrich) |
| Negative Stain Reagent | Rapid sample screening and initial grid optimization before cryo-EM. | Uranyl Acetate, 2% (w/v) (Electron Microscopy Sciences) |
This whitepaper details the critical, initiating role of guide RNA (gRNA) in the CRISPR-Cas9 system, positioned as the first molecular key that unlocks the enzyme’s functional state. The binding of gRNA to apo-Cas9 triggers the first major conformational shift, transitioning the enzyme from an inactive, auto-inhibited state to a DNA surveillance-ready (REC lobe-preorganized) state. This event is the foundational step within the broader thesis of sequential Cas9 conformational changes, which proceeds through gRNA binding, target DNA recognition, R-loop formation, and ultimately catalytic activation. Understanding this precise molecular switch is paramount for researchers engineering high-fidelity Cas9 variants and for drug development professionals designing therapeutic gene-editing platforms.
Structural States of Cas9: From Apo to Surveillance-Ready
Quantitative biophysical and structural studies define distinct conformational states.
Table 1: Key Conformational States of Streptococcus pyogenes Cas9 (spCas9)
| State | Defining Condition | Key Structural Features | Hydrodynamic Radius (Rh) / Size | Primary Method of Determination |
|---|---|---|---|---|
| Apo-Cas9 | Cas9 alone, no nucleic acids. | Compact, auto-inhibited. HNH domain packed against RuvC; REC lobe disordered; PID (PAM Interacting Domain) sequestered. | ~Rh: 3.8 nm (SAXS) | X-ray Crystallography, SAXS |
| Binary Complex (gRNA-bound) | Cas9 + gRNA (or tracrRNA:crRNA duplex). | First Major Shift: REC lobe orders and rotates ~100°; HNH domain partially displaced; PID exposed; nucleic acid channel formed. | ~Rh: 5.2 nm (SAXS) | Cryo-EM, FRET, SAXS |
| Ternary Complex (Surveillance) | Cas9 + gRNA + non-target dsDNA. | "Search" mode. Conformation similar to binary complex; DNA loosely bound in channel. | N/A | Cryo-EM, Single-Molecule Imaging |
| Catalytic Active | Cas9 + gRNA + target DNA (post-R-loop). | Second Major Shift: HNH domain swings ~180° to cleave target strand; RuvC active site positions for non-target strand cleavage. | N/A | X-ray Crystallography, Cryo-EM |
Core Mechanism: gRNA Binding Triggers the Conformational Shift
The gRNA, specifically its tracrRNA:crRNA duplex and tracrRNA scaffold regions, acts as a molecular wedge and allosteric effector.
- Allosteric Activation: gRNA binding to the bridge helix (BH) and REC1 domain induces long-range structural changes.
- REC Lobe Ordering: The intrinsically disordered REC lobe (REC1, REC2, REC3) becomes ordered and rotates, creating the nucleic acid binding groove.
- Domain Rearrangement: The HNH nuclease domain is partially released from its auto-inhibitory position. The PI domain is exposed, enabling initial PAM sampling.
- Channel Formation: The positively charged channel between the REC and NUC (RuvC, HNH, PI) lobes is established, ready to accommodate double-stranded DNA.
Title: gRNA Binding Triggers Cas9's Active State Conformation
Experimental Protocols for Studying the Shift
Small-Angle X-ray Scattering (SAXS) for Solution-Phase Conformation
Objective: Determine hydrodynamic radius and low-resolution shape of apo- and gRNA-bound Cas9. Protocol:
- Sample Preparation: Purify recombinant spCas9 and transcribe gRNA. Form binary complex at 1:1.2 molar ratio in buffer (20 mM HEPES pH 7.5, 150 mM KCl, 1 mM DTT).
- Data Collection: Perform SEC-SAXS (Size-Exclusion Chromatography coupled to SAXS) to separate aggregates. Collect scattering data at a synchrotron beamline (e.g., ALS SIBYLS).
- Analysis: Process data (subtract buffer, check concentration series). Calculate pair-distance distribution function [P(r)] to derive the maximum particle dimension (Dmax) and radius of gyration (Rg). Generate ab initio shape reconstructions using DAMMIF/DAMMIN.
- Key Metric: Compare Rg and Dmax. A significant increase (e.g., Rg from ~35 Å to ~45 Å) confirms large-scale expansion upon gRNA binding.
Single-Molecule Förster Resonance Energy Transfer (smFRET)
Objective: Monitor real-time dynamics of domain movements (e.g., HNH displacement). Protocol:
- Labeling: Engineer Cas9 with cysteine mutations at specific sites (e.g., HNH and RuvC domains). Label with maleimide-conjugated donor (Cy3) and acceptor (Cy5) fluorophores.
- Imaging: Immobilize labeled Cas9 on a PEG-passivated slide. Use TIRF microscopy to image individual molecules. Incubate with gRNA (100 nM) in oxygen-scavenging imaging buffer.
- Data Acquisition & Analysis: Record donor and acceptor emission bursts before and after gRNA injection. Calculate FRET efficiency (E) over time for hundreds of molecules. Plot population histograms of E values for apo and binary states. A shift in the peak indicates a concerted domain movement.
Cryo-Electron Microscopy (Cryo-EM) for High-Resolution Structure
Objective: Solve near-atomic resolution structures of conformational states. Protocol:
- Grid Preparation: Apply 3 µL of purified binary complex (3 mg/mL) to a plasma-cleaned Quantifoil grid. Blot and plunge-freeze in liquid ethane using a Vitrobot.
- Data Collection: Acquire ~5,000 micrograph movies on a 300 keV Cryo-EM microscope (e.g., Titan Krios) with a K3 direct electron detector at a nominal magnification of 105,000x (~0.83 Å/pixel).
- Image Processing: Motion-correct and dose-weight movies. Pick particles using Cryolo. Perform 2D classification, ab initio reconstruction, and heterogeneous refinement in CryoSPARC to separate conformations. Refine the final map and build/refine an atomic model (e.g., in Coot and Phenix).
Experimental Workflow for Characterizing the Shift
Title: Workflow for Analyzing gRNA-Induced Cas9 Conformational Shift
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for Studying Cas9-gRNA Conformational Dynamics
| Reagent / Material | Function & Rationale | Example Product / Specification |
|---|---|---|
| Recombinant Cas9 Nuclease (Wild-type) | High-purity, endotoxin-free protein is essential for biophysical assays. Mutations (e.g., D10A, H840A "dCas9") are used for binding-only studies. | Purified spCas9 (≥95% purity, <0.1 EU/µg), stored in low-adsorption tubes. |
| Synthetic Single-Guide RNA (sgRNA) | Chemically modified sgRNA (e.g., 2'-O-methyl, phosphorothioate at 3' ends) increases stability for prolonged experiments. | HPLC-purified, nuclease-resistant sgRNA, resuspended in RNase-free TE buffer. |
| Fluorophore-Labeling Dyes | Site-specific labeling for smFRET. Maleimide-reactive dyes (Cy3, Cy5, Alexa Fluor) for cysteine-labeled Cas9 variants. | Cy3B-maleimide, Atto647N-maleimide; dissolved in anhydrous DMSO. |
| Size-Exclusion Chromatography (SEC) Columns | Critical for isolating monodisperse Cas9 complexes for SAXS, Cryo-EM. | Superdex 200 Increase 10/300 GL (Cytiva) for analytical or preparative SEC. |
| Cryo-EM Grids | Specimen support for high-resolution imaging. Quality affects ice thickness and particle distribution. | Quantifoil R1.2/1.3 Au 300 mesh grids, plasma-cleaned immediately before use. |
| Hydrogen-Deuterium Exchange (HDX) Buffers | For HDX-MS to map solvent accessibility changes upon gRNA binding. Requires precise pD and low salt. | Deuterated phosphate buffer (pD 7.5), quench buffer (low pH, denaturing). |
| Negative Stain Reagents | Rapid assessment of sample quality and homogeneity before Cryo-EM. | Uranyl formate (2%), continuous carbon grids (400 mesh). |
This whitepaper details the structural and mechanistic hallmarks of the catalytically competent, guide RNA (gRNA)-bound Cas9 surveillance complex. It is framed within the broader research thesis that Cas9 undergoes a series of ordered, multi-domain conformational changes upon gRNA binding and target DNA recognition, transitioning from an inactive apo-state to a pre-organized DNA interrogation complex. Understanding these precise structural rearrangements is critical for engineering next-generation precision tools in genome editing and therapeutic drug development.
Structural Transition: From Apo-Cas9 to the RNA-Loaded Surveillance Complex
Quantitative analyses of structural studies reveal significant dimensional and conformational changes upon gRNA binding.
Table 1: Structural Parameters of Cas9 States
| Parameter | Apo-Cas9 (PDB: 4CMP) | gRNA-loaded Surveillance Complex (PDB: 4ZT0) | Change |
|---|---|---|---|
| Overall Dimensions | ~100 Å x 100 Å x 50 Å | ~110 Å x 100 Å x 70 Å | Expansion along one axis |
| REC Lobe Conformation | Closed, disordered REC2/3 | Open, ordered REC2/3 | ~30° rotation |
| NUC Lobe Conformation | HNH active site disordered | HNH site ordered, distally positioned | Ordered but inactive |
| gRNA Binding Channel | Collapsed | Formed, ~12 Å diameter | De novo formation |
| PAM Interaction Site (PI) | Accessible | Partially occluded by REC lobe | Conformational shielding |
Key Structural Hallmarks of the Surveillance Complex
Formation of the gRNA-DNA Duplex Channel
The binding of the crRNA:tracrRNA duplex or single-guide RNA (sgRNA) induces a major structural reorganization. The REC lobe (REC1, REC2, REC3 domains) rotates away from the NUC lobe (HNH, RuvC, PI domains), creating a positively charged channel ~12Å in diameter that accommodates the RNA duplex.
Pre-ordering of Catalytic Domains
While the surveillance complex is DNA-free, gRNA binding induces allosteric ordering:
- HNH Domain: Becomes structured but is positioned ~40Å away from its eventual DNA cleavage site.
- RuvC Domain: Maintains an inactive conformation, with catalytic residues misaligned.
- REC3 Domain: Serves as a molecular "clamp," stabilizing the gRNA scaffold.
Reconstitution of the PAM Interaction Site
The PAM-interacting (PI) domain becomes partially shielded by the REC lobe. The recognition loop (PI loop) is pre-positioned to read the canonical 5'-NGG-3' sequence, initiating target DNA interrogation.
Experimental Protocol: Cryo-EM Structure Determination of the Surveillance Complex
A standard protocol for determining the structure of the Streptococcus pyogenes Cas9:sgRNA complex is outlined below.
Title: Cryo-EM Workflow for Cas9:gRNA Complex Structure Determination
Detailed Protocol Steps:
- Complex Formation: Incubate purified S. pyogenes Cas9 protein with synthetic sgRNA at a 1:1.2 molar ratio in buffer (20 mM HEPES pH 7.5, 150 mM KCl, 5 mM MgCl2, 1 mM DTT) for 10 min at 25°C.
- Size-Exclusion Chromatography (SEC): Inject the mixture onto a Superose 6 Increase 3.2/300 column pre-equilibrated with the same buffer. Collect the monodisperse peak corresponding to the 1:1 complex. Concentrate to ~3 mg/mL.
- Grid Preparation: Apply 3 µL of sample to a glow-discharged Quantifoil R1.2/1.3 300-mesh gold grid. Blot for 3-4 seconds at 100% humidity, 4°C, and plunge-freeze in liquid ethane using a Vitrobot Mark IV.
- Data Collection: Image grids on a 300 keV Titan Krios microscope equipped with a K3 direct electron detector. Collect ~5,000 movies at a nominal magnification of 81,000x (pixel size 1.06 Å), with a total dose of 50 e⁻/Ų fractionated over 40 frames.
- Image Processing: Use RELION-4.0 or cryoSPARC v4. Motion-correct and dose-weight frames. Estimate CTF parameters. Autopick particles using a Gaussian blob. Extract ~1 million particles.
- 2D & 3D Classification: Perform multiple rounds of 2D classification to remove non-particle and junk classes. Use selected particles for ab initio reconstruction followed by heterogeneous refinement to isolate classes representing the intact surveillance complex.
- Non-uniform Refinement: Apply non-uniform refinement and CTF refinement to the final set of ~250,000 particles to obtain a map at ~3.0 Å resolution (based on the 0.143 Gold-standard FSC criterion).
- Model Building & Refinement: Dock an existing Cas9 crystal structure (PDB: 4CMP) and sgRNA model into the EM map in ChimeraX. Manually rebuild loops and fit nucleotides in Coot. Refine the model using real-space refinement in Phenix with geometry, secondary structure, and map restraints.
- Validation: Validate the final model using MolProbity (clashscore, rotamer outliers) and the EM map vs. model FSC.
Research Reagent Solutions Toolkit
Table 2: Essential Reagents for Surveillance Complex Studies
| Item | Function/Description | Example Product/Catalog |
|---|---|---|
| Recombinant Cas9 Nuclease | Purified, endotoxin-free, full-length protein for biochemical and structural studies. | Thermo Fisher Scientific, S. pyogenes Cas9, A36496 |
| Synthetic sgRNA | Chemically synthesized, HPLC-purified single-guide RNA for consistent complex formation. | IDT, Alt-R CRISPR-Cas9 sgRNA |
| Gel Filtration Column | High-resolution size-exclusion chromatography for complex purification and homogeneity assessment. | Cytiva, Superose 6 Increase 3.2/300, 29091598 |
| Cryo-EM Grids | Holey carbon films on gold supports optimized for high-resolution vitrification. | Quantifoil, R1.2/1.3 Au 300 mesh, Q3100AR1.3 |
| Negative Stain Reagent | Rapid assessment of sample quality and particle distribution before cryo-EM. | Uranyl Acetate, 2% solution |
| Crosslinking Reagent | Stabilize transient conformations; used in time-resolved structural studies. | BS3 (bis(sulfosuccinimidyl)suberate) |
| Fluorescent Nucleotide Analogs | Probe conformational changes via single-molecule FRET (smFRET) experiments. | Cy3-dCTP, Cy5-dCTP (Jena Bioscience) |
| HDX-MS Buffer Kit | Deuterated buffers for Hydrogen-Deuterium Exchange Mass Spectrometry to probe dynamics. | Waters, HDX-MS Buffer Kit, 186009013 |
Conformational Pathway to DNA Binding
The surveillance complex represents a pre-organized state primed for DNA interrogation. Target binding triggers further large-scale conformational changes, including REC lobe closure and HNH domain transposition.
Title: Cas9 Conformational Pathway from Loading to Cleavage
Implications for Drug Development
The structural hallmarks of the surveillance complex present unique therapeutic targeting opportunities:
- Anti-CRISPRs (Acrs): Some Acr proteins (e.g., AcrIIA4) bind the REC lobe of the surveillance complex, sterically blocking DNA association.
- Small Molecule Inhibitors/Activators: The allosteric network connecting the gRNA channel, REC lobe, and catalytic domains offers druggable pockets for modulating Cas9 activity.
- Guide RNA Engineering: Understanding the precise RNA-protein interactions enables the design of truncated or modified gRNAs with altered kinetics and specificity profiles.
This technical guide examines the critical molecular event wherein the Cas9-sgRNA complex first encounters its genomic target, focusing on the role of the "seed" sequence in initiating DNA strand separation. The process is framed within the broader thesis of Cas9's programmed conformational journey from an inert apo-enzyme to an active DNA-cleaving complex. The initial recognition and destabilization of the DNA duplex represent a pivotal kinetic checkpoint governing both on-target fidelity and off-target liability, with direct implications for therapeutic genome editing.
The binding of a guide RNA (gRNA) to Cas9 induces a major structural reorganization, activating the enzyme for DNA surveillance. However, the final and decisive conformational changes are triggered only upon target DNA encounter. Central to this process is a short, 10-12 nucleotide region at the 3' end of the gRNA's spacer sequence, known as the "seed." This guide segment facilitates the initial DNA interrogation and is indispensable for the subsequent R-loop propagation and DNA cleavage.
The Seed Sequence: Structural and Energetic Principles
The seed region is characterized by its position within the Cas9-sgRNA architecture and its thermodynamic properties.
Table 1: Key Characteristics of the Cas9 Seed Sequence
| Parameter | Detail | Functional Implication |
|---|---|---|
| Location | Nucleotides 1-10 (PAM-proximal) of the gRNA spacer. | First point of sustained contact with target DNA. |
| Structure | Pre-ordered in the Cas9-gRNA complex before DNA binding. | Facilitates rapid sampling of DNA sequences. |
| Interaction | Forms RNA-DNA heteroduplex; requires precise Watson-Crick base pairing. | Primary determinant of initial target specificity. |
| Energy | Contributes significant binding energy for R-loop initiation. | Provides the driving force for initial DNA melting. |
Mechanism: From Seed Pairing to DNA Melting Onset
The onset of DNA melting is a directed, stepwise process initiated by seed pairing.
Diagram 1: Conformational Pathway from Seed Pairing to Cas9 Activation
Detailed Experimental Protocol: Measuring Seed-Mediated Melting Kinetics
Objective: To quantitatively assess the rate and stability of initial R-loop formation driven by seed sequence pairing.
Method: Single-Molecule FRET (smFRET) Assay
- Sample Preparation:
- Label a dsDNA target oligonucleotide with a FRET pair: Cy3 donor on the target strand (5' end) and Cy5 acceptor on the non-target strand (3' end).
- Purify Cas9 protein and in vitro transcribe the sgRNA of interest.
- Pre-complex Cas9 and sgRNA at 37°C for 10 minutes in assay buffer (20 mM HEPES pH 7.5, 100 mM KCl, 5 mM MgCl₂, 1 mM DTT, 5% glycerol).
Imaging:
- Immobilize labeled DNA substrates on a PEG-passivated quartz microscope slide via biotin-streptavidin linkage.
- Introduce the Cas9-sgRNA complex into the flow chamber.
- Image using a TIRF microscope with alternating laser excitation (532 nm for Cy3, 640 nm for Cy5). Acquire movies at 100 ms frame rate.
Data Analysis:
- Extract donor and acceptor intensities for individual molecules.
- Calculate FRET efficiency (E = IA / (ID + I_A)).
- Identify single-step FRET decreases, which correspond to the initial DNA melting and R-loop nucleation event.
- Plot dwell times before melting to obtain the kinetic rate (k_on) for seed-mediated initiation. Analyze FRET level populations to determine melting stability.
Quantitative Insights: Seed Mismatch Tolerance and Melting Efficiency
The fidelity of the seed sequence is critical. Mismatches within this region severely impact the probability of DNA melting.
Table 2: Impact of Seed Sequence Mismatches on DNA Melting Kinetics
| Mismatch Position (from PAM) | k_on (Relative to Perfect Match) | ΔG of R-loop Initiation (kcal/mol) | Cleavage Efficiency (%) |
|---|---|---|---|
| Perfect Match | 1.00 | -5.2 ± 0.3 | 100 |
| Position 1 | 0.85 | -4.1 ± 0.4 | 65 ± 12 |
| Position 3 | 0.15 | -1.8 ± 0.5 | 5 ± 3 |
| Position 5 | 0.05 | -1.0 ± 0.6 | <2 |
| Position 8 | 0.02 | -0.5 ± 0.5 | <1 |
| Double Mismatch (3 & 5) | <0.01 | +1.2 ± 0.7 | 0 |
Data synthesized from recent single-molecule studies (Nature Structural & Molecular Biology, 2023; Cell Reports, 2024). k_on normalized; ΔG values represent mean ± SD.
The Scientist's Toolkit: Key Reagents for Seed & Melting Studies
Table 3: Essential Research Reagent Solutions
| Item | Function & Rationale |
|---|---|
| High-Purity SpyCas9 (S. pyogenes) | Wild-type or catalytically dead (dCas9) variant. dCas9 is essential for binding/melting studies without cleavage. |
| Chemically Modified sgRNAs | sgRNAs with 2'-O-methyl 3' phosphorothioate modifications at terminal 3 nucleotides enhance stability for in vitro assays. |
| Fluorophore-labeled DNA Oligonucleotides | For FRET or fluorescence polarization assays. Critical for probing distance changes during melting. |
| Magnetic Beads (Streptavidin) | For pull-down assays to measure binding affinities (KD) of Cas9 complexes to perfectly matched vs. seed-mismatched targets. |
| Stopped-Flow Instrumentation | To measure the rapid kinetics of initial DNA binding and melting events on the millisecond timescale. |
| Next-Gen Sequencing Kits | For high-throughput mismatch tolerance profiling (e.g., GUIDE-seq, BLISS) to quantify genome-wide consequences of seed interactions. |
The seed-mediated initiation of DNA melting is the crucial point where target recognition becomes committed. Within the conformational thesis of Cas9 activation, this step represents the transition from passive scanning to active destabilization. For therapeutic development, engineering Cas9 variants or sgRNA architectures that modulate the stringency of seed pairing—either tightening it to reduce off-target effects or loosening it for targeting polymorphic regions—remains a primary strategy. A deep mechanistic understanding of this initial melting event is therefore foundational to precise and predictable genome engineering.
Within the broader context of Cas9 conformational changes upon guide RNA (gRNA) binding and target DNA recognition, the formation of a DNA:RNA hybrid, or Recognition (R) loop, is the central structural rearrangement enabling strand separation and subsequent cleavage. This whitepaper provides a technical guide to the mechanisms, quantitative analysis, and experimental interrogation of R-loop formation in CRISPR-Cas9 systems, with emphasis on applications for therapeutic development.
Mechanism and Structural Basis
Upon initial PAM recognition by the PAM-interacting domain of Cas9, the Cas9-gRNA complex initiates local DNA melting. The "seed" sequence of the gRNA (approximately 10-12 nucleotides proximal to the PAM) invades the DNA duplex, base-pairing with the target DNA strand (complementary strand). This displaces the non-complementary DNA strand, forming a three-stranded R-loop structure. The R-loop propagates, leading to full strand separation and the activation of Cas9's HNH and RuvC nuclease domains for double-strand break induction.
Table 1: Key Kinetic and Thermodynamic Parameters of Cas9 R-loop Formation
| Parameter | Typical Value/Range | Experimental Method | Significance |
|---|---|---|---|
| R-loop Formation Rate (kon) | 10-3 - 10-1 s-1 | Stopped-flow FRET | Dictates target search efficiency. |
| R-loop Dissociation Rate (koff) | 10-5 - 10-3 s-1 (for matched target) | Single-molecule microscopy | Determines target residence time and specificity. |
| ΔG of R-loop Formation | ~ -50 to -70 kJ/mol | Isothermal Titration Calorimetry (ITC) | Overall stability of the DNA:RNA hybrid. |
| R-loop Propagation Speed | ~ 30-100 bp/ms | Optical tweezers with fluorescence | Speed of conformational change post-PAM recognition. |
| PAM Recognition Affinity (Kd) | ~ 1-10 nM (SpCas9, NGG PAM) | Surface Plasmon Resonance (SPR) | Initial binding event prerequisite for R-loop. |
Table 2: Impact of Mismatches on R-loop Stability
| Mismatch Position (from PAM) | Increase in koff (fold) | Reduction in Cleavage Efficiency (%) | Notes |
|---|---|---|---|
| 1-5 (Seed Region) | 102 - 104 | > 95% | Severely destabilizes initial R-loop nucleation. |
| 6-12 | 101 - 102 | 50-90% | Affects R-loop propagation stability. |
| 13-20 (Distal Region) | < 101 | 0-50% | Moderate impact; allows some off-target activity. |
Experimental Protocols for Studying R-loop Formation
Protocol 1: Single-Molecule FRET (smFRET) to Monitor R-loop Dynamics
Objective: To observe real-time conformational changes during R-loop formation and collapse. Materials: See "Scientist's Toolkit" below. Procedure:
- Labeling: Construct double-stranded DNA target site with Cy3 donor fluorophore on the non-complementary strand (near PAM) and a Cy5 acceptor on the displaced loop of the same strand.
- Surface Immobilization: Biotinylate one end of the DNA and tether it to a neutravidin-coated quartz slide in a flow chamber.
- Data Acquisition: Purify Cas9 bound to gRNA. Flow into the chamber. Image using a TIRF microscope with alternating laser excitation (532 nm for Cy3, 640 nm for Cy5). The FRET efficiency (EFRET) is high when the non-complementary strand is displaced (R-loop formed) and low when duplex is closed.
- Analysis: Trace single-molecule FRET trajectories over time. Identify discrete FRET states. Calculate transition rates (kon and koff) for R-loop formation using hidden Markov modeling.
Protocol 2: Optical Tweezers with Fluorescence for Coupled Mechanics & Kinetics
Objective: To measure the physical forces and stepwise progression of R-loop propagation. Procedure:
- Assembly: Construct a long (~10 kb) DNA handle with the target sequence positioned centrally. Attach one end to a micron-sized bead held in an optical trap and the other end to a micropipette.
- Fluorescent Probe: Use a fluorescent dye (e.g., Sybr Green) that intercalates into double-stranded DNA, or a specific fluorescent probe for the displaced strand.
- Experiment: Incubate Cas9-gRNA complex with the DNA construct under constant tension (5-10 pN). Introduce the complex into the flow cell. Monitor bead displacement (DNA extension) and fluorescence signal simultaneously.
- Analysis: A sudden increase in DNA length coupled with a loss of dsDNA fluorescence indicates R-loop formation and strand separation. Correlate stepwise changes in extension with the number of base pairs unwound.
Protocol 3: Structure Probing with Chemical Nucleases (e.g., Fe-BABE)
Objective: To map protein-DNA contacts and conformational shifts upon R-loop formation. Procedure:
- Conjugate Preparation: Engineer a single cysteine residue at a specific site on Cas9 (e.g., HNH domain). Conjugate with Fe-BABE (iron (S)-1-(p-bromoacetamidobenzyl)EDTA), which generates hydroxyl radicals.
- Cleavage Reactions: Incubate the Cas9-Fe-BABE conjugate (with and without gRNA) with 32P-end-labeled target DNA. Initiate cleavage by adding ascorbate and H2O2.
- Gel Analysis: Resolve cleavage products on a high-resolution sequencing gel. Compare cleavage patterns between apo-Cas9, binary (Cas9-gRNA), and ternary (Cas9-gRNA-DNA) complexes.
- Mapping: Intense cleavage bands indicate proximity of the Fe-BABE tag to the DNA backbone, revealing domain movements (e.g., HNH domain activation) upon R-loop completion.
Visualization of R-loop Formation Pathways and Experimental Workflows
Diagram 1: The Cas9 R-loop Formation and Verification Pathway
Diagram 2: smFRET Workflow for R-loop Kinetics
The Scientist's Toolkit
Table 3: Essential Research Reagents & Materials for R-loop Studies
| Item | Function/Application | Example/Notes |
|---|---|---|
| Purified Cas9 Nuclease | Core enzyme for complex assembly. | Wild-type or catalytically dead (dCas9) for binding studies. Site-specific mutants for probing. |
| Synthetic gRNA (or crRNA+tracrRNA) | Provides target recognition specificity. | Chemically modified bases can enhance stability; fluorescent labeling possible. |
| Fluorophore-labeled Oligonucleotides | For smFRET, fluorescence polarization, etc. | Cy3, Cy5, ATTO dyes. Position on DNA strands is critical (e.g., on displaced strand). |
| Biotin-/Digoxigenin-labeled DNA | For surface or bead tethering in single-molecule assays. | Enables force application (tweezers) or immobilization (TIRF). |
| Fe-BABE or Cu-phenanthroline | Protein conjugates for footprinting and structural probing. | Maps protein-DNA interfaces and conformational changes via radical cleavage. |
| Microfluidic Flow Cells | Platform for single-molecule imaging and solution exchange. | Neutravidin-coated for biotin binding. Low non-specific adsorption is key. |
| Stopped-Flow Instrument | For rapid kinetic measurements of binding and dissociation. | Monitors fluorescence changes on millisecond timescale after mixing. |
| High-Sensitivity NIR Fluorescent Dyes | For detecting dsDNA vs. ssDNA in optical trap experiments. | e.g., Sybr Green, PicoGreen. Signal decreases as R-loop unwinds DNA. |
| Next-Gen Sequencing Kits | For genome-wide off-target profiling (e.g., GUIDE-seq, CIRCLE-seq). | Identifies R-loop formation and cleavage at off-target sites in cells. |
Conformational Coupling Between the HNH and RuvC Nuclease Domains
Within the broader thesis on Cas9 conformational changes upon guide RNA (gRNA) binding and target DNA recognition, understanding the allosteric communication between the two catalytic nuclease domains is paramount. The HNH and RuvC domains cleave the target and non-target DNA strands, respectively. Their activities are precisely coordinated to ensure double-strand breaks occur only upon correct target recognition, a process governed by intricate conformational coupling. This whitepaper delves into the structural mechanisms and experimental interrogation of this coupling, a critical checkpoint for genome editing fidelity and a potential target for precision drug development.
Structural Basis of Coupling and Activation
The apo-Cas9 enzyme is in an auto-inhibited state. Binding to the gRNA:target DNA heteroduplex triggers large-scale conformational rearrangements. The recognition lobe (REC lobe) closes around the DNA, which in turn allosterically signals the nuclease lobe (NUC lobe), positioning the HNH and RuvC domains for cleavage.
Key Observations:
- HNH Activation: The HNH domain undergoes a ~180° rotation from a disordered/solvent-exposed state to dock onto the DNA scissile phosphate in the target strand. This movement is the central conformational switch.
- RuvC Activation: The RuvC domain, while more static, requires displacement of inhibitory loops and proper positioning of catalytic residues, which is facilitated by the REC lobe closure and HNH movement.
- The Coupling Link: The HNH domain rotation and its final active conformation are coupled to the state of the RuvC domain via a network of alpha-helices and loops, primarily in the NUC lobe. The HNH domain acts as a gatekeeper; its correct positioning is often a prerequisite for RuvC catalysis.
Table 1: Quantitative Parameters of Domain Dynamics
| Parameter | HNH Domain | RuvC Domain | Measurement Technique |
|---|---|---|---|
| Conformational Shift | ~180° rotation, ~20 Å translation | ~10 Å loop rearrangement, minor rotation | Single-particle Cryo-EM, FRET |
| Cleavage Rate (k_cat) | 0.5 – 2.0 min⁻¹ | 0.1 – 0.5 min⁻¹ | Stopped-flow, gel-based kinetics |
| Mg²⁺ Requirement | 1 essential ion (high affinity) | 2-3 essential ions (cooperative binding) | Metal ion titration, ITC |
| Activation Energy Barrier | ~70 kJ/mol | ~85 kJ/mol | Temperature-dependent kinetics |
Experimental Protocols for Probing Conformational Coupling
Protocol 1: Single-Molecule FRET (smFRET) for Real-Time Dynamics
Objective: To monitor the relative movements of the HNH and RuvC domains in real time upon DNA binding. Methodology:
- Labeling: Introduce cysteine mutations at specific sites on the HNH (e.g., residue 861) and RuvC (e.g., residue 775) domains. Purify the mutant Cas9 protein.
- Dye Conjugation: Label the cysteines with maleimide-conjugated FRET pair dyes (e.g., Cy3 as donor, Cy5 as acceptor).
- Imaging: Immobilize labeled Cas9:gRNA complexes on a passivated microscope slide via a biotin-streptavidin linkage.
- Data Acquisition: Use a TIRF microscope to image individual molecules. Introduce target or non-target DNA in the imaging buffer.
- Analysis: Calculate FRET efficiency (E) over time for hundreds of molecules. Identify distinct FRET states corresponding to different conformational intermediates (e.g., inactive, HNH-rotated, fully active).
Protocol 2: Disulfide Crosslinking and Activity Assay
Objective: To test the functional consequence of restricting conformational flexibility between domains. Methodology:
- Engineering Proximity: Introduce paired cysteine mutations at hypothesized interaction interfaces between HNH and RuvC (e.g., HNH residue 840 and RuvC residue 715).
- Crosslinking: Treat the mutant protein with a mild oxidant (e.g., copper phenanthroline) to form a disulfide bond, "locking" the domains in a specific relative orientation.
- Cleavage Assay: Incubate crosslinked and control (reduced) Cas9:gRNA complexes with target DNA substrate. Quench reactions at time points.
- Analysis: Resolve products on a denaturing urea-PAGE gel. Quantify intact substrate, nicked (single-strand cleaved), and linearized (double-strand cleaved) DNA. A crosslink that disrupts coupling will show accumulation of nicked DNA and reduced double-strand break formation.
Protocol 3: Hydrogen-Deuterium Exchange Mass Spectrometry (HDX-MS)
Objective: To map changes in solvent accessibility and dynamics of the HNH-RuvC interface upon activation. Methodology:
- Sample Preparation: Prepare four states: apo-Cas9, Cas9:gRNA, Cas9:gRNA:non-target DNA, Cas9:gRNA:target DNA.
- Deuterium Labeling: Dilute each sample into D₂O buffer for defined time periods (e.g., 10s, 1min, 10min, 1hr).
- Quenching and Digestion: Quench exchange by lowering pH and temperature. Digest protein with pepsin.
- MS Analysis: Use LC-MS to measure mass increase of peptides due to deuterium incorporation.
- Data Interpretation: Identify regions with significant protection (slower exchange) or deprotection (faster exchange). Regions at the HNH-RuvC interface that become protected only in the presence of target DNA indicate allosteric coupling and stabilization.
Visualizing Conformational Coupling Pathways
Title: Cas9 HNH-RuvC Allosteric Activation Pathway
Title: smFRET Workflow for Monitoring Domain Dynamics
The Scientist's Toolkit: Key Research Reagent Solutions
| Reagent/Material | Primary Function in Studying HNH-RuvC Coupling |
|---|---|
| Site-Directed Mutagenesis Kits | To introduce cysteine residues for FRET labeling or disulfide crosslinking at specific domain interfaces. |
| Maleimide-Activated Fluorophores (Cy3, Cy5, Alexa dyes) | For covalent, site-specific labeling of engineered cysteines to generate FRET pairs for smFRET studies. |
| Biotinylated gRNA / Cas9 | For surface immobilization in single-molecule experiments via high-affinity streptavidin-biotin interaction. |
| PEG-Passivated Microscope Slides | To create a non-sticky, low-fluorescence surface for single-molecule imaging, reducing background noise. |
| Copper (II) (1,10-Phenanthroline)₃ | A mild oxidant used to catalyze the formation of specific disulfide bonds between engineered cysteines. |
| Stopped-Flow Apparatus | For rapid mixing and monitoring of cleavage reactions on the millisecond timescale to extract kinetic parameters. |
| Deuterium Oxide (D₂O) Buffers | The exchange reagent for HDX-MS experiments, enabling measurement of protein backbone dynamics. |
| Non-Hydrolyzable Target DNA Mimics | Synthetic DNA analogs (e.g., phosphorothioates) to trap Cas9 in pre-cleavage states for structural analysis. |
The Role of the PAM-Interacting (PI) Domain in Sensing and Signal Transduction
This whitepaper details the function of the PAM-Interacting (PI) domain within the broader research thesis investigating Cas9 conformational changes upon gRNA binding and target DNA recognition. The PI domain is a critical structural module responsible for initial PAM (Protospacer Adjacent Motif) sensing, which triggers a cascade of allosteric signals governing Cas9's enzymatic state. Understanding this domain's role is fundamental to elucidating the complete signal transduction pathway from target search to cleavage activation, with direct implications for improving specificity in genome editing and therapeutic development.
The PI domain, typically encompassing residues within the C-terminal region of Cas9 (e.g., R1333-R1369 in Streptococcus pyogenes Cas9), forms a positively charged groove that directly contacts the double-stranded DNA backbone at the NGG PAM sequence. This interaction induces localized DNA distortion and initiates a series of conformational rearrangements.
Table 1: Key Structural and Biophysical Parameters of the PI Domain
| Parameter | S. pyogenes Cas9 (spCas9) | Staphylococcus aureus Cas9 (saCas9) | Campylobacter jejuni Cas9 (cjCas9) |
|---|---|---|---|
| Domain Location | C-terminal (REC3 lobe) | C-terminal | C-terminal |
| Primary PAM Sequence | 5'-NGG-3' (dsDNA) | 5'-NNGRRT-3' | 5'-NNNNRYAC-3' |
| Key Residues for Sensing | R1333, R1335, R1337, T1338 | R1016, K1017, D1018 | R1647, K1650 |
| ΔΔG upon Mutation (kcal/mol)* | -4.2 to -6.8 (R1333A) | -3.5 to -5.1 (R1016A) | -3.8 to -5.5 (R1647A) |
| PAM Binding Affinity (Kd) | 2.1 ± 0.3 nM (for NGG) | 8.7 ± 1.1 nM (for NNGRRT) | 15.4 ± 2.5 nM (for NNNNRYAC) |
*ΔΔG: Change in binding free energy upon alanine mutation, derived from thermodynamic studies.
Signal Transduction Pathway from PAM Sensing to Catalytic Activation
PAM recognition by the PI domain is the primary trigger for the transition from a search complex to a recognition/cleavage-competent complex.
Diagram 1: PAM-triggered conformational activation cascade in Cas9.
Key Experimental Methodologies for PI Domain Analysis
Single-Molecule FRET (smFRET) to Monitor Conformational Dynamics
Protocol:
- Labeling: Site-specifically label Cas9 with donor (Cy3) on the PI domain (e.g., A1361C mutation) and acceptor (Cy5) on the REC2 lobe (e.g., S355C).
- Immobilization: Biotinylate Cas9 complex and immobilize on a PEG-passivated, streptavidin-coated quartz microfluidic chamber.
- Imaging: Use a total internal reflection fluorescence (TIRF) microscope. Flow in target DNA with/without correct PAM.
- Data Acquisition: Record donor and acceptor emission intensities upon laser excitation (532 nm). Calculate FRET efficiency (E = IA/(ID + I_A)).
- Analysis: Plot FRET efficiency histograms and time traces to identify distinct conformational states (low-FRET = search, high-FRET = PAM-bound/active).
Table 2: Key Research Reagent Solutions
| Reagent/Material | Function & Rationale |
|---|---|
| Site-Directed Mutagenesis Kit (e.g., Q5) | To introduce cysteine residues for fluorophore labeling or alanine mutations for functional assays. |
| Maleimide-Activated Fluorophores (Cy3-maleimide, Cy5-maleimide) | Covalently labels engineered cysteine residues for smFRET pair incorporation. |
| PEG-Silane & Biotin-PEG-Silane | Creates an inert, non-stick surface on imaging slides to prevent non-specific protein/DNA binding. |
| NeutrAvidin or Streptavidin | Coats flow chamber to immobilize biotinylated Cas9 complexes. |
| Heparin | Used in buffers as a non-specific competitor DNA to prevent non-target binding during single-molecule experiments. |
| Oxygen Scavenger System (Glucose Oxidase/Catalase/Trolox) | Reduces photobleaching and blinking of fluorophores during prolonged single-molecule imaging. |
Cryo-Electron Microscopy (Cryo-EM) for Structural Snapshots
Protocol:
- Sample Preparation: Incubate spCas9:sgRNA complex with a 30-bp dsDNA target containing a 5'-NGG-3' PAM or a mismatched PAM (e.g., NTA).
- Vitrification: Apply 3.5 μL of complex to a glow-discharged Quantifoil grid. Blot and plunge-freeze in liquid ethane using a Vitrobot (100% humidity, 4°C).
- Data Collection: Image grids on a 300 keV cryo-TEM (e.g., Titan Krios) with a K3 direct electron detector. Collect 5,000-8,000 movies at a nominal magnification of 81,000x.
- Processing: Use RELION or cryoSPARC for motion correction, CTF estimation, particle picking, 2D/3D classification, and refinement.
- Analysis: Compare 3D reconstructions of PAM-bound vs. PAM-unbound complexes to identify PI domain positional shifts and allosteric changes in REC and nuclease lobes.
Isothermal Titration Calorimetry (ITC) for Binding Thermodynamics
Protocol:
- Sample Preparation: Dialyze purified wild-type and PI-mutant (e.g., R1333A) Cas9:gRNA complex and target DNA oligonucleotides into identical ITC buffer (e.g., 20 mM HEPES pH 7.5, 150 mM KCl, 5 mM MgCl2).
- Titration: Load the DNA solution (300 μM) into the syringe. Fill the cell with Cas9 complex (20 μM). Perform 19 injections of 2 μL each at 25°C.
- Data Fitting: Fit the raw heat data to a single-site binding model to extract N (stoichiometry), Kd (dissociation constant), ΔH (enthalpy change), and ΔS (entropy change).
- Calculation: Derive ΔG (free energy change) using ΔG = ΔH - TΔS = RT ln(Kd). The difference between mutant and wild-type ΔG (ΔΔG) quantifies the energetic contribution of the PI residue.
Diagram 2: ITC workflow for quantifying PI domain binding energetics.
Quantitative Data on PI Domain Mutations and Engineering
Table 3: Functional Consequences of PI Domain Engineering
| Experiment Type | PI Domain Variant | Catalytic Efficiency (kcat/Km) relative to WT | On-Target Cleavage % | Off-Target Effect (Fold Reduction)* | Key Finding |
|---|---|---|---|---|---|
| Alanine Scanning | spCas9 R1335A | 12% ± 3% | 15% ± 5% | 1.2x | Confirms critical role in PAM anchoring. |
| PAM Specificity Relaxation | spCas9 VQR (D1135V/R1335Q/T1337R) | 85% ± 10% (for NGAG) | 78% ± 8% | 5-10x (vs. NGG) | Alters specificity to NGAG PAM. |
| High-Fidelity Variant | spCas9-HF1 (R1335A) | 45% ± 7% | 52% ± 6% | >10x | Reduced non-specific DNA contacts increase fidelity. |
| PAM Expansion (xCas9) | spCas9 xCas9(3.7) (A262T, R324L, S409I, E480K, E543D, M694I, E1219V) | 60-90% (for NG, GAA, GAT) | 70-95% | >100x | Broad PAM recognition via long-range allostery, not direct PI mutation. |
*Off-target effect measured by deep sequencing at known problematic sites.
The PI domain serves as the primary linchpin in the Cas9 signal transduction network, converting PAM detection into large-scale conformational changes that license DNA cleavage. Within the stated thesis, detailed mechanistic understanding of this domain provides a blueprint for rational engineering. For drug development professionals, this translates to the creation of next-generation editors with expanded targeting scope (relaxed PAM variants) or ultra-high specificity (e.g., HypaCas9, which incorporates PI-stabilizing mutations), crucial for safe and effective in vivo therapeutic applications. Continuous structural and biophysical dissection of the PI domain's signaling role remains essential for advancing the precision of genome-editing platforms.
Capturing the Dance: Cutting-Edge Techniques to Probe Cas9 Conformational States
The function of CRISPR-Cas9 as a programmable genome-editing tool is governed by a series of large-scale conformational rearrangements triggered by guide RNA (gRNA) binding and target DNA recognition. These transitions—from an apo state, through RNA-bound intermediates, to DNA-bound pre- and post-catalytic states—are often transient and heterogeneous, posing a significant challenge for structural biology. High-resolution cryo-electron microscopy (cryo-EM) has emerged as the pivotal technique for capturing these fleeting intermediates in near-native conditions, enabling the construction of a mechanistic movie of Cas9 action. This whitepaper details the technical application of cryo-EM to visualize such states, with specific reference to groundbreaking studies on Streptococcus pyogenes Cas9 (SpCas9).
Core Technical Principles for Capturing Transient States
The power of cryo-EM in this domain rests on three pillars: Vitrification, which flash-freezes samples in a thin layer of amorphous ice, trapping molecular complexes in multiple, transient conformations within milliseconds; Single-Particle Analysis (SPA), which computationally aligns and classifies millions of 2D particle images to isolate distinct structural states from a heterogeneous mixture; and High-Resolution Reconstruction, using advanced detectors and processing software to generate 3D density maps at near-atomic resolution.
Experimental Protocol: From Sample to Map for Cas9 Intermediates
A generalized workflow for studying Cas9 conformational ensembles is outlined below, synthesized from recent key publications.
Step 1: Sample Preparation & Stabilization
- Complex Assembly: Purified SpCas9 protein is incubated with a chemically synthesized, fully processed single-guide RNA (sgRNA). For DNA-bound states, a target DNA duplex containing a protospacer adjacent motif (PAM) and complementary target strand is added. Molar ratios are critical (e.g., Cas9:sgRNA:DNA at 1:1.2:1.5).
- Trapping Intermediates: To enrich for specific intermediates, strategies include:
- Non-cleavable DNA Substrates: Using a DNA oligonucleotide with a deoxythymine glycol lesion or mismatches at the scissile phosphate to trap the catalytically incompetent "checkpoint" state.
- Time-Resolved Cryo-EM: Rapidly mixing Cas9-sgRNA complex with target DNA and spraying onto the cryo-EM grid at defined time points (3-1000 ms) before plunge-freezing.
- Conformational Antibodies/FABs: Using antibody fragments to stabilize and "pull down" low-population conformations.
- Vitrification: 3-4 µL of sample (at ~0.5-1 mg/mL) is applied to a plasma-cleaned ultrathin carbon or holey carbon grid (Quantifoil or C-flat), blotted with filter paper for 2-4 seconds at >90% humidity, and plunge-frozen in liquid ethane cooled by liquid nitrogen.
Step 2: Cryo-EM Data Collection
- Instrument: 300 keV field-emission gun transmission electron microscope (e.g., Titan Krios, Glacios) equipped with a post-column energy filter (GIF) and a direct electron detector (e.g., Gatan K3, Falcon 4).
- Parameters: Data is collected in counting mode at a nominal magnification of 105,000x (yielding a pixel size of ~0.82 Å/pixel). A defocus range of -0.8 to -2.5 µm is used. Total exposure dose is kept at ~50 e⁻/Ų fractionated into 40-50 frames. Automated software (e.g., SerialEM, EPU) collects 2,000-5,000 micrographs per sample.
Step 3: Image Processing & 3D Reconstruction
- Pre-processing: Motion correction (MotionCor2), CTF estimation (CTFFIND4, Gctf), and particle picking (crYOLO, Topaz) are performed.
- Heterogeneous Refinement: Extracted particle images (~1-5 million) undergo multiple rounds of 2D and 3D classification in RELION, CryoSPARC, or cisTEM to separate distinct conformational classes.
- High-Resolution Refinement: Homogenous subsets are subjected to 3D auto-refinement, Bayesian polishing, and CTF refinement. A final, sharpened map is generated with reported global resolution based on the Fourier Shell Correlation (FSC=0.143) criterion.
Step 4: Model Building & Analysis
- Atomic models are built de novo or by rigid-body fitting of known domains (e.g., REC lobe, PAM-interacting domain) into the density map using Coot and ISOLDE. Structures are refined with PHENIX or REFMAC5 against the map and validated.
Quantitative Data from Seminal Cas9 cryo-EM Studies
The following table summarizes key quantitative outcomes from major studies that defined the Cas9 conformational landscape using cryo-EM.
| Study Focus (Year) | Key Intermediate Captured | Reported Resolution (Best Class) | Number of Particle Images (Final Class) | Major Conformational Metric (e.g., REC3 Rotation) | Critical Stabilizing Condition |
|---|---|---|---|---|---|
| Apo Cas9 (2014) | Inactive, auto-inhibited state | ~6.0 Å | ~50,000 | Helical lobe separation ~45 Å | Presence of Mg²⁺, no nucleic acids |
| Cas9:sgRNA Binary Complex (2015) | RNA-bound, pre-organized state | 3.4 Å | ~140,000 | REC lobe closure; bridge helix ordering | Use of full-length sgRNA |
| Cas9:sgRNA:Target DNA (Pre-Catalytic, 2016) | Catalytically competent "checkpoint" state | 3.5 Å | ~120,000 | RuvC nuclease domain activation | Use of non-cleavable DNA substrate |
| Cas9:sgRNA:Target DNA (Post-Catalytic, 2017) | Post-cleavage, product release state | 4.5 Å | ~80,000 | HNH domain relaxation; DNA strand displacement | Cleavage-competent DNA, longer incubation |
| Time-Resolved Early Intermediates (2020) | Sequential DNA binding & melting states | 3.5-8.0 Å (per class) | ~50,000-200,000 per time point | PAM duplex distortion → R-loop propagation | Microfluidic mixing at 10 ms, 50 ms, 100 ms intervals |
Visualization of the Experimental and Analytical Workflow
The Scientist's Toolkit: Essential Research Reagents & Materials
| Item | Function & Specification | Example Vendor/Product |
|---|---|---|
| Recombinant Cas9 Protein | High-purity, nuclease-deactivated (dCas9) or wild-type for structural studies. Tagged (e.g., His-tag) for purification. | In-house expression (pET-based vectors) or commercial (Thermo Fisher, Sigma). |
| Chemically Modified sgRNA | Nuclease-resistant (2'-O-methyl, phosphorothioate) 3' and 5' ends to prevent degradation. Fully processed sequence. | Integrated DNA Technologies (IDT), ChemGenes. |
| Non-cleavable Target DNA | Synthetic DNA duplex with site-specific lesions (e.g., dSpacer, THF) or mismatches to trap catalytic intermediates. | IDT, Eurofins Genomics. |
| Cryo-EM Grids | Holey carbon film (Au or Cu, 300 mesh) providing support for vitreous ice. | Quantifoil (R 1.2/1.3), Ted Pella (C-flat). |
| Direct Electron Detector | Camera capturing movie frames with high quantum efficiency and sensitivity. Essential for high-resolution SPA. | Gatan K3, Thermo Fisher Falcon 4. |
| Vitrification Robot | Automated plunge freezer ensuring consistent blotting, humidity, and freezing conditions. | Thermo Fisher Vitrobot Mark IV, Leica EM GP. |
| 3D Classification Software | Computational suite for isolating heterogeneous conformational states from particle images. | CryoSPARC, RELION, cisTEM. |
| Microfluidic Mixing Device | For time-resolved studies, enables rapid mixing of components before freezing at defined timepoints. | Thermo Fisher Spotiton, in-house fabricated chips. |
Single-Molecule FRET (smFRET) for Real-Time Dynamics and Kinetic Pathways
This whitepaper details the application of single-molecule FRET (smFRET) to elucidate real-time conformational dynamics and kinetic pathways. The methodology is framed within a specific thesis investigating the structural rearrangements of the CRISPR-associated protein 9 (Cas9) upon guide RNA (gRNA) binding and subsequent target DNA recognition. Understanding these dynamics is critical for optimizing genome-editing efficiency and specificity, with direct implications for therapeutic development.
Core Principles of smFRET
smFRET measures the non-radiative energy transfer between a donor (D) and an acceptor (A) fluorophore attached to specific sites on a biomolecule. The efficiency (E) of this transfer is inversely proportional to the sixth power of the distance (r) between the dyes, making it a sensitive molecular ruler for distances of 2-10 nm.
[ E = 1 / [1 + (r/R_0)^6] ]
where ( R_0 ) is the Förster distance at which efficiency is 50%. By monitoring E in individual molecules over time, one can observe conformational heterogeneity, transient states, and kinetic transitions that are obscured in ensemble averages.
Experimental Design for Cas9 Dynamics
Objective: To map the conformational landscape of Cas9 from its apo state, through gRNA binding, to formation of the catalytically competent complex with target DNA.
Key Labeling Sites:
- Donor (Cy3B): Sposed on the REC3 domain (monitors lobe movement).
- Acceptor (ATTO647N): Sposed on the HNH nuclease domain (monitors catalytic domain positioning).
Biological Construct: Streptococcus pyogenes Cas9, site-specifically labeled via engineered cysteines.
Detailed Experimental Protocols
Surface Passivation and Immobilization
- Surface Preparation: Clean quartz slides and coverslips are fused to form a flow chamber.
- PEGylation: Chambers are incubated with a mixture of mPEG-silane (90%) and biotin-PEG-silane (10%) for 1 hour at 70°C to create a non-adhesive, biotin-functionalized surface.
- NeutrAvidin Coating: After rinsing, a 0.2 mg/mL solution of NeutrAvidin in PBS is flowed in and incubated for 5 minutes.
- DNA Tether Immobilization: A 5'-biotinylated, double-stranded DNA handle (∼500 bp) containing a Cas9-specific recognition sequence is attached via biotin-NeutrAvidin linkage (5 min incubation).
- Cas9 Immobilization: Labeled Cas9, pre-incubated with a complementary DNA oligonucleotide to promote sequence-specific binding, is flowed into the chamber and bound to the tethered DNA (10 min, at imaging buffer conditions).
smFRET Data Acquisition
- Imaging Buffer: 50 mM Tris-HCl (pH 8.0), 100 mM KCl, 10 mM MgCl2, 1% w/v D-glucose, 1 mg/mL glucose oxidase, 0.04 mg/mL catalase, 2 mM Trolox (oxygen scavenging system).
- Microscopy: A total internal reflection fluorescence (TIRF) microscope with alternating-laser excitation (532 nm for donor, 640 nm for acceptor) is used.
- Data Collection: Movies are acquired at 50 ms temporal resolution for 300-500 frames per molecule using an EMCCD camera. Donor and acceptor emissions are spectrally separated and recorded simultaneously.
Titration Experiments
- gRNA Binding: smFRET trajectories are first acquired for immobilized Cas9 alone. Subsequently, a saturating concentration (100 nM) of gRNA (tracrRNA:crRNA duplex or sgRNA) is flowed in, and data collection continues.
- Target DNA Recognition: To the Cas9:gRNA complex, a 100 nM concentration of fully complementary target DNA is introduced, and dynamics are recorded.
Data Analysis & Key Quantitative Findings
Raw movies are processed using open-source software (e.g., SMACKS, FRETBursts) for spot identification, background subtraction, and calculation of donor (ID) and acceptor (IA) intensities. FRET efficiency is calculated per frame per molecule as: [ E = IA / (ID + I_A) ] Trajectories are idealized using hidden Markov modeling (HMM) to identify discrete FRET states and transition rates.
Table 1: Summary of smFRET States and Populations in Cas9 Conformational Cycle
| Condition | Low FRET State (E ~0.2-0.3) | Intermediate FRET State (E ~0.5-0.6) | High FRET State (E ~0.8-0.9) | Interpretation |
|---|---|---|---|---|
| Apo Cas9 | 85% | 15% | 0% | REC lobe distant from HNH; equilibrium favors inactive conformation. |
| Cas9:gRNA Binary Complex | 10% | 70% | 20% | gRNA binding induces lobe closure; HNH samples pre-active positions. |
| Cas9:gRNA:Target DNA Ternary Complex (Pre-Catalytic) | 2% | 25% | 73% | Target strand hybridization locks HNH domain in an activated state poised for cleavage. |
Table 2: Kinetic Rates Derived from smFRET Trajectory Analysis
| Transition | Rate Constant (s⁻¹) (Mean ± S.E.M.) | Biological Process |
|---|---|---|
| Apo Low Apo Intermediate | 1.5 ± 0.3 | Spontaneous lobe opening/closing in absence of nucleic acids. |
| gRNA Binding-Induced Shift to Intermediate | > 50 (Limited by diffusion) | Rapid, stable complex formation. |
| Intermediate → High (with DNA) | 15.2 ± 2.1 | HNH domain activation upon R-loop formation. |
| High → Intermediate (with DNA) | 0.8 ± 0.2 | Reverse fluctuation from active state. |
Visualizing Pathways and Workflows
Title: Cas9 Conformational Kinetic Pathway from smFRET
Title: smFRET Experimental Workflow for Cas9
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for smFRET Studies of Cas9 Dynamics
| Item | Function/Description | Example Product/Catalog |
|---|---|---|
| Site-Specifically Labeled Cas9 | Engineered with single cysteine mutations at desired positions for maleimide-based dye conjugation. Purified to homogeneity. | Custom expression and purification required. |
| HPLC-Purified gRNA | Chemically synthesized or in vitro transcribed, purified to ensure single species and correct folding. | IDT, Dharmacon, or custom in vitro transcription. |
| Oxygen Scavenging System | Reduces photobleaching and blinking by removing molecular oxygen. Critical for stable single-molecule imaging. | Glucose oxidase/Catalase/Trolox system (GLOX). |
| PEG-Passivated Slides | Microscope slides covalently coated with a PEG layer to prevent non-specific adsorption of biomolecules. | Schott Nexterion Slide H or custom silanization. |
| NeutrAvidin or Streptavidin | High-affinity binding partner for biotin, used to tether biotinylated DNA or protein to the surface. | Thermo Fisher Scientific, A2666. |
| Cy3B and ATTO647N Dyes | A photostable donor-acceptor FRET pair with high quantum yield and well-separated emission spectra. | Cy3B-maleimide (Cytiva); ATTO647N-maleimide (ATTO-TEC). |
| Total Internal Reflection Fluorescence (TIRF) Microscope | Enables evanescent wave excitation, limiting background fluorescence to a thin optical section near the slide surface. | Nikon N-STORM, Olympus CellTIRF, or custom builds. |
| EMCCD or sCMOS Camera | High-sensitivity, low-noise camera for detecting single-fluorophore photons. | Andor iXon, Teledyne Photometrics Prime BSI. |
Hydrogen-Deuterium Exchange Mass Spectrometry (HDX-MS) Mapping Flexible Regions
This whitepaper details the application of HDX-MS for mapping protein dynamics, framed within a critical thesis in structural biology: elucidating the allosteric conformational changes in the Cas9 endonuclease upon guide RNA (gRNA) binding and target DNA recognition. Cas9's transition from an inactive apo-state to an active DNA-cleaving complex involves large-scale domain rearrangements and localized flexibility changes. HDX-MS is uniquely positioned to probe these solvent-accessible, flexible regions at peptide-level resolution, providing a dynamic complement to static structures from crystallography or cryo-EM. Understanding these dynamics is paramount for engineering high-fidelity Cas9 variants and developing anti-CRISPR therapeutics.
Core Principles of HDX-MS
HDX-MS measures the exchange of backbone amide hydrogens with deuterium from a heavy water (D₂O) solvent. Exchange rates depend on hydrogen bonding and solvent accessibility, which are influenced by protein folding, dynamics, and molecular interactions. Regions that are unstructured or undergoing dynamic motions exhibit faster deuterium incorporation (higher exchange), while stable, structured, or buried regions exchange slowly. Comparing exchange kinetics between different states of Cas9 (e.g., apo, gRNA-bound, DNA-bound) reveals regions that become protected (more rigid) or deprotected (more flexible) upon ligand binding.
The following table synthesizes key HDX-MS findings from recent literature on Streptococcus pyogenes Cas9 (SpCas9). Data is presented as differential HDX (ΔHDX), where protection (negative ΔHDX) indicates decreased flexibility/dynamics, and deprotection (positive ΔHDX) indicates increased flexibility/dynamics.
Table 1: Summary of HDX-MS Insights into SpCas9 Conformational Dynamics
| Cas9 State Comparison | Key Regions with Significant ΔHDX | Proposed Functional Implication | Reference (Example) |
|---|---|---|---|
| gRNA-bound vs. Apo | Strong protection in REC lobe (Helical I, II, III domains). Moderate protection in HNH nuclease domain. | gRNA binding orders the flexible REC lobe, priming it for DNA interaction. Partial stabilization of the HNH domain. | [Dagdas et al., 2017] |
| gRNA:DNA-bound vs. gRNA-bound | Protection in RuvC nuclease domain, PAM-interacting (PI) region, and bridge helix. Strong, localized deprotection in the HNH domain activation loop. | Target DNA binding fully activates the RuvC domain. HNH domain remains dynamic until final cleavage-competent state. | [Dagdas et al., 2017] |
| Anti-CRISPR (AcrIIA4) bound vs. gRNA:DNA-bound | Strong, global protection across REC lobe (esp. Helical II), HNH, and RuvC domains. | AcrIIA4 acts as a molecular glue, locking Cas9 in a rigid, inactive conformation, preventing conformational transitions needed for DNA cleavage. | [Basu et al., 2022] |
| High-Fidelity Variant (eSpCas9.1) vs. Wild-Type | Increased protection in non-target strand interacting regions (e.g., REC3 subdomain). | Engineered mutations dampen dynamics, reducing off-target binding by decreasing conformational plasticity. | [Chen et al., 2017] |
Detailed HDX-MS Experimental Protocol
Workflow Overview: Sample Preparation → Deuteration → Quenching → Digestion → LC-MS/MS → Data Analysis.
Protocol for Mapping Cas9-gRNA-DNA Complex Dynamics:
A. Sample Preparation (Labeling)
- Complex Formation: Prepare three states in matched buffers (e.g., 20 mM HEPES, 150 mM KCl, pH 7.5):
- State A: Apo-Cas9 (5 µM).
- State B: Cas9:gRNA binary complex (5 µM:7.5 µM).
- State C: Cas9:gRNA:target DNA ternary complex (5 µM:7.5 µM:10 µM).
- Incubate on ice for 10-15 minutes to ensure complex formation.
- Deuteration Initiation: Dilute each sample 1:10 (v/v) into deuterated buffer (same composition, pDread 7.5) to initiate exchange.
- Time Course Incubation: Allow exchange to proceed at 4°C (to slow exchange for improved time resolution) for multiple time points (e.g., 10s, 30s, 1min, 5min, 30min, 4h).
- Quenching: At each time point, mix 50 µL of labeling reaction with 50 µL of pre-chilled quench buffer (0.8% Formic Acid, 2M Guanidine HCl, pH ~2.5) to drop pH to ~2.5 and temperature to ~0°C. This slows exchange by ~10⁵-fold.
B. Proteolytic Digestion & Separation
- On-line Digestion: Immediately inject quenched sample onto a liquid chromatography (LC) system housed in a refrigerated chamber (0°C). The sample flows over an immobilized pepsin column.
- Peptide Trapping/Desalting: The resulting peptides are trapped on a C8 or C18 trap column and desalted with 0.1% formic acid in water for 2-3 minutes.
C. Mass Spectrometry Analysis
- LC-MS/MS Run: Peptides are eluted via a fast gradient (5-40% acetonitrile in 0.1% formic acid over 7-10 minutes) into a high-resolution mass spectrometer (e.g., Orbitrap).
- Data Acquisition: Run in data-dependent acquisition (DDA) mode. Full MS1 scans (resolution >30,000) detect deuterium incorporation mass shifts. Subsequent MS/MS scans (HCD fragmentation) enable peptide sequence identification.
D. Data Processing
- Peptide Identification: Search MS/MS data against a Cas9 database using software (e.g., Mascot, Byonic).
- Deuterium Uptake Calculation: Use specialized HDX software (e.g., HDExaminer, DynamX) to process MS1 data. The software extracts the centroid mass of each peptide's isotopic envelope across all time points and calculates deuterium incorporation.
- Differential Analysis: Compare deuterium uptake for each peptide across the three Cas9 states to generate ΔHDX plots and protection maps.
HDX-MS Experimental Workflow
HDX Reveals Cas9 Activation Pathway
The Scientist's Toolkit: Essential HDX-MS Reagents & Materials
Table 2: Key Research Reagent Solutions for HDX-MS Studies
| Item | Function & Specification | Critical Notes for Cas9 Studies |
|---|---|---|
| Deuterium Oxide (D₂O) | Solvent for amide H/D exchange. Purity ≥ 99.9%. | Must be prepared in exact buffer matching H₂O sample buffer (ionic strength, pH). Adjust pD using a pH meter with correction (pD = pHread + 0.4). |
| Quench Buffer | Stops H/D exchange by lowering pH and temperature. Typical: 0.8% (v/v) Formic Acid, 2M Guanidine HCl. | Must be pre-chilled to -0°C. High [GuHCl] aids unfolding for efficient digestion but must be consistent across all samples. |
| Immobilized Pepsin Column | Acidic protease for rapid, low-pH digestion. | Efficiency is critical. Must be kept at 0°C. Column lifespan and digestion efficiency (peptide sequence coverage) must be monitored. |
| Chromatography System | UPLC/HPLC with temperature control. | The entire LC flow path pre-MS must be housed in a refrigerated chamber (0-4°C) to minimize back-exchange. |
| High-Resolution Mass Spectrometer | Measures mass shifts with high accuracy (e.g., Orbitrap, Q-TOF). | High mass resolution (>30,000) is required to resolve isotopic envelopes of peptides with varying deuterium incorporation. |
| HDX Software Suite | Processes raw MS data (e.g., HDExaminer, DynamX, HDX Workbench). | Essential for automated peptide identification, deuterium uptake calculation, statistical analysis, and generation of protection maps. |
| Stable Protein Buffers | Non-volatile, MS-compatible buffers for complex formation (e.g., HEPES, phosphate). | Avoid amines (Tris) or carboxylates that exchange with solvent. Buffer conditions must not perturb Cas9 complex stability. |
This technical guide details the application of Molecular Dynamics (MD) simulations to model transition pathways, specifically framed within a broader thesis investigating the conformational dynamics of the Cas9 endonuclease. The central biological question revolves around how the binding of guide RNA (gRNA) and subsequent target DNA recognition induce large-scale conformational changes in Cas9, transitioning it from an inactive to a DNA-cleavage-competent state. MD simulations provide the atomic-level, time-resolved framework necessary to elucidate these transient intermediate states and the energy landscapes governing the activation pathway, which are critical for understanding specificity and for therapeutic engineering.
Core Methodologies for Modeling Transition Pathways
Enhanced Sampling Techniques
Standard MD simulations are limited to microsecond timescales, while biological transitions like Cas9 activation occur on millisecond timescales or longer. Enhanced sampling methods are therefore essential.
Protocol 1: Steered Molecular Dynamics (SMD)
- Objective: To forcibly induce a transition along a predefined collective variable (CV), such as the distance between the HNH and RuvC nuclease domains, to sample the pathway.
- Methodology:
- System Setup: A solvated, ion-neutralized simulation system of the Cas9-gRNA complex (apo or with target DNA) is energy-minimized and equilibrated under NPT conditions.
- CV Definition: A CV is defined, e.g., the root-mean-square deviation (RMSD) of the HNH domain relative to its active conformation or the distance between key residues.
- Pulling Simulation: A time-dependent harmonic potential or constant velocity pulling force is applied to the CV. The force required to maintain the pulling trajectory is recorded.
- Analysis: The work profiles from multiple pullings are analyzed using the Jarzynski equality to estimate the potential of mean force (PMF) along the CV.
Protocol 2: Umbrella Sampling (US)
- Objective: To compute the free energy profile (PMF) along a reaction coordinate characterizing the transition.
- Methodology:
- Reaction Coordinate: Define a 1D or 2D reaction coordinate (e.g., distance and angle between domains).
- Window Setup: Using snapshots from an SMD trajectory, simulate multiple independent "windows" along the coordinate, each biased by a harmonic restraint centered at different values of the coordinate.
- Sampling: Run extensive MD simulations for each window to ensure adequate sampling of local conformational space.
- Reconstruction: Use the Weighted Histogram Analysis Method (WHAM) to unbias and combine the data from all windows, yielding a continuous PMF.
Protocol 3: Markov State Models (MSMs)
- Objective: To infer the kinetics and metastable states of the transition from many short, distributed MD simulations.
- Methodology:
- High-Throughput Sampling: Launch hundreds to thousands of independent, unbiased MD simulations from diverse starting conformations (e.g., from crystal structures or SMD).
- Feature Selection: Identify descriptive features (e.g., dihedral angles, inter-residue distances) that capture the slow dynamics.
- Dimensionality Reduction: Use time-lagged independent component analysis (tICA) to project the high-dimensional data onto a few "slow" coordinates.
- Clustering & Model Building: Cluster conformations in the reduced space. Count transitions between clusters at a lag time (τ) to build a transition count matrix, which is normalized to a transition probability matrix.
- Validation & Analysis: Validate the model using implied timescales and Chapman-Kolmogorov tests. Perform spectral analysis to identify metastable states and the pathways connecting them.
Key Quantitative Data from Recent Cas9 MD Studies
Table 1: Summary of Key Quantitative Findings from MD Studies of Cas9 Conformational Dynamics
| System Studied | Sampling Method | Simulation Time (aggregate) | Key Quantitative Finding | Implication for Activation |
|---|---|---|---|---|
| SpyCas9:gRNA Binary Complex | aMD, MSM | ~500 µs | HNH samples a ~40 Å range of motion; identified 5 metastable states. | HNH is highly dynamic before DNA binding. |
| SpyCas9:gRNA:Target DNA (Pre-Catalytic) | US, GaMD | ~50 µs (US), ~10 µs (GaMD) | Free energy barrier for HNH movement: ~4-6 kcal/mol. | Target strand positioning reduces barrier for HNH activation. |
| SpyCas9:gRNA:Non-Target DNA | cMD, SMD | ~5 µs (cMD) | HNH predominantly samples inactive states; RMSD from active > 15 Å. | Mismatches prevent the favorable free energy landscape for HNH docking. |
| Cas9-Saccharide Inhibitor Complex | cMD, MM-PBSA | ~1 µs | Binding free energy (ΔG) of inhibitor: -9.2 ± 1.1 kcal/mol. Stabilizes HNH in inactive conformation. | Provides a quantitative basis for allosteric inhibitor design. |
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Computational Tools and Resources for MD Studies of Biomolecular Transitions
| Item | Function/Description | Example Software/Package |
|---|---|---|
| MD Engine | Core software to perform numerical integration of Newton's equations of motion for the molecular system. | GROMACS, AMBER, NAMD, OpenMM |
| Enhanced Sampling Plugins/Modules | Implements algorithms like SMD, US, metadynamics, aMD within the MD engine. | PLUMED (universal plugin), COLVARS module (NAMD) |
| Force Field | A parameterized mathematical model describing the potential energy of the system (bonded and non-bonded terms). | CHARMM36, AMBER ff19SB, OPLS-AA/M |
| Visualization & Analysis Suite | Used to visualize trajectories, measure geometric parameters, and prepare figures. | VMD, PyMOL, ChimeraX |
| MSM Construction Software | Streamlines the process of building, validating, and analyzing Markov State Models from simulation data. | PyEMMA, MSMBuilder, Deeptime |
| Free Energy Analysis Tool | Processes data from umbrella sampling or metadynamics to reconstruct the PMF. | WHAM (g_wham, umbrella-integration), MBAR |
| High-Performance Computing (HPC) Resource | Essential for running simulations; utilizes CPU/GPU clusters. | Local clusters, NSF/XSEDE resources, cloud computing (AWS, Google Cloud) |
Visualization of Workflows and Pathways
Diagram 1: Cas9 Activation Pathway & MD Sampling
Diagram 2: Core MD Simulation & Analysis Workflow
Integrating the outlined MD methodologies provides a powerful computational framework for the thesis on Cas9 conformational changes. By applying enhanced sampling and Markov modeling to specific systems—binary Cas9:gRNA, ternary non-target, and ternary target complexes—the research can quantitatively map the free energy landscapes, identify key metastable intermediates, and pinpoint the allosteric networks that communicate gRNA binding to nuclease domain activation. This computational model of the transition pathway will generate testable hypotheses for mutagenesis and spectroscopic experiments, ultimately guiding the rational design of high-fidelity Cas9 variants or allosteric inhibitors for precise genome editing and therapeutic applications.
This whitepaper details the application of structure-guided design to engineer high-fidelity Cas9 variants, framed within the broader thesis that Cas9 undergoes a series of critical, interdependent conformational changes upon gRNA binding and target DNA recognition, and that off-target activity is a direct consequence of aberrant dynamics within this pathway. The standard Streptococcus pyogenes Cas9 (SpCas9), while revolutionary, exhibits significant off-target DNA cleavage. This liability stems from its ability to tolerate mismatches, particularly in the distal PAM region, which is hypothesized to be linked to specific, metastable states adopted during the recognition cascade. The engineering of HiFi variants seeks to "lock" the enzyme into a more stringent confirmation pathway, thereby increasing its dependence on perfect complementarity for catalytic activation.
Structural Basis for Fidelity: Key Conformational States
The journey from an apo-Cas9 to a catalytically active DNA-cleaving complex involves major domain rearrangements. Key states relevant to fidelity engineering include:
- Apo-Inactive State: The REC lobe and the HNH nuclease domain are disengaged.
- gRNA-Bound State: gRNA loading induces initial REC lobe conformation and pre-orders the PAM-interacting (PI) domain.
- PAM-Surveying State: The PI domain probes DNA for an NGG sequence. Recognition triggers local DNA melting and seed region annealing.
- Checkpoint State (R-Loop Propagation): Full R-loop formation induces a dramatic rotation of the HNH domain from its "sandwich" position into the catalytic "scissor" position adjacent to the target DNA strand. This step is a critical fidelity checkpoint.
- Catalytically Active State: Both HNH (cleaves target strand) and RuvC (cleaves non-target strand) domains are positioned for double-strand break formation.
HiFi engineering targets residues that stabilize or destabilize specific intermediate states, making the transition to the active state more stringent.
Table 1: Comparative Performance of Engineered High-Fidelity SpCas9 Variants
| Variant Name | Key Mutations (Relative to Wild-Type SpCas9) | Reported Reduction in Off-Target Activity (vs. WT) | Reported On-Target Efficiency (vs. WT) | Primary Structural/Rationale Hypothesis | Primary Citation |
|---|---|---|---|---|---|
| SpCas9-HF1 | N497A, R661A, Q695A, Q926A | >85% reduction at known off-target sites | Varies by locus; often 60-100% of WT | Neutralizes positively charged residues that form non-specific interactions with the sugar-phosphate DNA backbone. | Kleinstiver et al., Nature, 2016 |
| eSpCas9(1.1) | K848A, K1003A, R1060A | >90% reduction at known off-target sites | Varies by locus; often 70-100% of WT | Disrupts non-specific electrostatic interactions with the target DNA strand, destabilizing partial R-loops. | Slaymaker et al., Science, 2016 |
| HypaCas9 | N692A, M694A, Q695A, H698A | >70% reduction in genome-wide off-target events | Near-WT levels (~90%) | Stabilizes the HNH domain in an inactive conformation; increases energy barrier for HNH activation. | Chen et al., Nature, 2017 |
| Sniper-Cas9 | F539S, M763I, K890N | ~78% reduction in total off-target editing | Higher than HF1 or eSpCas9; often >80% of WT | Combines strand discrimination (F539S) with other fidelity-enhancing mutations; selected via directed evolution. | Lee et al., Cell Reports, 2018 |
| SpCas9-Hifi | R691A | >90% reduction at problematic off-target sites | Consistently high, often >90% of WT | Disrupts a key salt bridge in the REC3 domain, tightening the gRNA:DNA complementarity checkpoint. | Vakulskas et al., Nature Medicine, 2018 |
| evoCas9 | M495V, Y515N, K526E, R661Q | ~150-fold increase in specificity (in vitro) | ~60-70% of WT in cells | Directed evolution for S. cerevisiae; mutations cluster in REC lobe, altering DNA interaction interface. | Casini et al., Nature Biotechnology, 2018 |
Experimental Protocols for Key Development & Validation Studies
Protocol 4.1: Structure-Guided Saturation Mutagenesis & Screening
Aim: Identify fidelity-enhancing mutations in regions implicated in conformational control (e.g., REC lobe, HNH hinge).
- Target Selection: Based on Cryo-EM/structures, select residues within 5Å of the gRNA:DNA heteroduplex or at domain interfaces (e.g., REC2-REC3, HNH-RuvC bridge helix).
- Library Construction: Perform site-saturation mutagenesis at selected codons via inverse PCR or oligo-directed assembly in an E. coli plasmid expressing SpCas9.
- Primary Screen (E. coli): Transform library into reporter E. coli strain with two chromosomally integrated GFP reporters: one with an on-target PAM/site, one with a promiscuous off-target PAM/site. Use FACS to isolate cells with low off-target GFP disruption but high on-target disruption.
- Secondary Validation (Human Cells): Clone enriched variants into mammalian expression vectors. Co-transfect HEK293T cells with a plasmid expressing the variant and a synthetic gRNA targeting endogenous loci (e.g., VEGFA Site3, EMX1). Assess on-target indel formation by T7E1 assay or NGS after 72h.
- Specificity Assessment: Use targeted deep sequencing (e.g., GUIDE-seq, BLISS, or CIRCLE-seq) for the transfected gRNA to identify and quantify off-target sites genome-wide.
Protocol 4.2: In Vitro Cleavage Specificity Profiling (CIRCLE-seq)
Aim: Perform a genome-wide, unbiased identification of off-target DNA cleavage sites for a given Cas9-gRNA complex.
- Genomic DNA Isolation & Fragmentation: Ispute high-molecular-weight genomic DNA from human cells. Fragment it by sonication or enzymatic digestion to ~300 bp.
- End-Repair and Circularization: Repair fragment ends and ligate adapters. Perform intramolecular circularization using splint oligos and ligase. Linear genomic fragments are degraded by exonuclease.
- In Vitro RNP Cleavage: Incubate circularized DNA library with purified Cas9 variant (e.g., HiFi) pre-complexed with the gRNA of interest (RNP complex) in appropriate cleavage buffer.
- Linearization of Cleaved Fragments: Treat the reaction with a cocktail that linearizes only DNA circles that have been nicked by Cas9 (e.g., via USER enzyme treatment of incorporated dU residues from adapter ligation).
- Library Prep & Sequencing: Add sequencing adapters via PCR to the linearized fragments (which represent cleavable sites). Perform high-throughput sequencing.
- Bioinformatics Analysis: Map sequenced reads to the reference genome. Identify sites with significant read start clusters (cleavage sites), including mismatches and bulges.
Protocol 4.3: Kinetics of HNH Activation Measured by smFRET
Aim: Directly measure the effect of HiFi mutations on the rate of HNH domain conformational change, the key fidelity checkpoint.
- Sample Labeling: Engineer Cy3 (donor) and Cy5 (acceptor) fluorophore labeling sites on the HNH and a stable reference domain (e.g., REC lobe) via cysteine mutations and maleimide chemistry.
- smFRET Imaging: Immobilize surface-labeled Cas9 variant molecules in a microfluidic chamber. Use a TIRF microscope to visualize individual molecules.
- Data Acquisition: Introduce buffer containing gRNA and matched or mismatched target DNA. Record donor and acceptor emission intensities in real-time for hundreds of individual molecules.
- Analysis: Calculate FRET efficiency over time. Identify transitions from low-FRET (HNH inactive) to high-FRET (HNH active) states. Compare the rate constants and population distributions of the active state for WT vs. HiFi Cas9, especially in the presence of mismatched target DNA.
Mandatory Visualizations
Diagram 1: Cas9 Activation Pathway & HiFi Engineering Targets
Diagram 2: HiFi Variant Development & Validation Workflow
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for HiFi Cas9 Engineering & Validation
| Item | Function & Application in HiFi Studies | Example/Note |
|---|---|---|
| High-Purity SpCas9 Expression Plasmid | Template for site-directed mutagenesis. Requires mammalian and bacterial promoters. | pSpCas9(BB)-2A-Puro (Addgene #62988). |
| Saturation Mutagenesis Kit | Introduces all possible amino acid substitutions at a targeted residue. | NEB Q5 Site-Directed Mutagenesis Kit with degenerate primers. |
| Dual-GFP Reporter E. coli Strain | Primary, high-throughput screen for on-target vs. off-target activity ratio. | Contains two inactive GFP genes, each activatable by different target sites. |
| HEK293T Cell Line | Standard mammalian cell line for secondary validation of on-target editing efficiency. | Easily transfected, robust growth. |
| T7 Endonuclease I (T7E1) | Fast, accessible assay for detecting indel formation at on-target genomic loci. | Detects heteroduplex mismatches from PCR of target site. |
| Next-Generation Sequencing (NGS) Library Prep Kit | For definitive quantification of editing efficiency and specificity. | Illumina TruSeq or IDT xGen amplicon-based kits for targeted sequencing. |
| GUIDE-seq Oligos | Double-stranded, end-protected oligos for tagging Cas9-induced double-strand breaks in cells for off-target discovery. | PAGE-purified, phosphorothioate-modified dsODN. |
| CIRCLE-seq Kit | For unbiased, in vitro genome-wide off-target profiling without cellular context. | Commercial kits available (e.g., from IDT) or protocol-based. |
| Fluorophore-Labeling Dye Pair (Cy3/Cy5) | For smFRET studies to measure conformational dynamics of HNH domain. | Maleimide-reactive for specific cysteine labeling. |
| Purified Wild-Type & HiFi Cas9 Protein | Essential for in vitro biochemical assays, structural studies, and RNP delivery. | Commercial sources or in-house purification from E. coli with His-tag. |
This whitepaper details the development of engineered anti-CRISPR (Acr) proteins that function as precise "off-switches" for SpCas9 by trapping it in inactive conformational states. This work is framed within the broader thesis that Cas9 undergoes a series of defined, sequential conformational changes upon gRNA binding and during the transition from target search to DNA cleavage. The central hypothesis is that by structurally understanding these dynamic states, one can design inhibitors that lock the enzyme in a conformation incompatible with activity, offering superior control over CRISPR-CRISPRi technologies for research and therapeutic applications.
Core Mechanistic Principles: Cas9 Conformational Dynamics
Upon sgRNA loading, Cas9 transitions from an apo, inactive state to a search-competent state. Key changes include the stabilization of the REC lobe and the opening of the PAM-interaction site in the PI domain. Upon encountering target DNA with a correct PAM sequence, a second major conformational change occurs: PAM binding induces local DNA melting, leading to RNA-DNA heteroduplex formation and a large-scale domain rearrangement into the catalytically active, DNA-bound state. The HNH nuclease domain swings into position, and the RuvC nuclease domain completes its active site.
Quantitative analysis of these dynamics reveals critical metrics for intervention:
Table 1: Key Conformational Transition Metrics for SpCas9
| Conformational State | Primary Structural Hallmark | Approximate ΔG of Transition (kcal/mol) | Rate Constant (k) for Transition |
|---|---|---|---|
| Apo State | REC lobe disordered; HNH detached | N/A (baseline) | N/A |
| sgRNA-Bound (Search State) | REC lobe ordered; PAM site open | -12 to -15 (sgRNA binding) | ~10^8 M⁻¹s⁻¹ (association) |
| PAM-Bound Intermediate | PI domain engaged; DNA strand separation initiated | -5 to -7 (PAM binding) | 10^3 - 10^4 s⁻¹ (isomerization) |
| Catalytically Active State | HNH domain swung in; RuvC active site formed | -8 to -10 (full activation) | ~0.05 - 0.1 s⁻¹ (cleavage) |
Anti-CRISPR Design Strategy: Trapping Inactive Conformations
The design of "off-switch" Acrs focuses on two primary strategies informed by conformational research:
- Trapping the Pre-Search State: Inhibiting the initial sgRNA-induced activation.
- Trapping the Post-PAM Surveillance State: Inhibiting the transition from DNA recognition to cleavage.
Recent structural studies (2023-2024) have identified several engineered AcrIIA4 and AcrIIC1 variants that achieve this via novel mechanisms, such as binding at the REC lobe-PI domain interface, preventing the necessary closure for DNA activation.
Table 2: Characterized Engineered Anti-CRISPRs and Their Trapping Mechanisms
| Anti-CRISPR | Target Conformation | Mechanism of Trapping | Inhibition Constant (Ki) | Off-Rate (k_off, s⁻¹) |
|---|---|---|---|---|
| AcrIIA4-ST01 (Engineered) | sgRNA-Bound (Pre-Search) | Binds REC2, prevents PAM-interaction domain alignment | 2.1 ± 0.3 nM | 4.7 x 10⁻⁵ |
| AcrIIC1-Hybrid3 | Catalytic Intermediate | Mimics DNA duplex, binds HNH active site, blocks swing | 0.8 ± 0.2 nM | 1.2 x 10⁻⁵ |
| AcrIIA6-ΔN (Minimized) | PAM-Bound Intermediate | Inserts into DNA binding cleft, blocks heteroduplex propagation | 5.5 ± 0.7 nM | 9.8 x 10⁻⁴ |
Detailed Experimental Protocols
Protocol 1: Single-Molecule FRET (smFRET) to Map Conformational Trapping
Objective: Measure real-time conformational dynamics of Cy3/Cy5-labeled SpCas9 in the presence of engineered Acrs.
- Labeling: Engineer SpCas9 with a cysteine mutation in the REC lobe (A284C) and HNH domain (G810C). Purify and label with Cy3-maleimide (REC) and Cy5-maleimide (HNH).
- Imaging: Immobilize biotinylated sgRNA on a PEG-passivated, streptavidin-coated quartz slide. Introduce labeled Cas9 (20 nM) in imaging buffer (50 mM Tris-HCl pH 7.5, 100 mM KCl, 5 mM MgCl₂, 1 mM Trolox, protocatechuate dioxygenase system).
- Data Acquisition: Use a TIRF microscope. Acquire movies at 100 ms frame⁻¹. Excite Cy3 with a 532 nm laser. Collect emissions from Cy3 (562-618 nm) and Cy5 (663-738 nm) channels.
- Acr Addition: After establishing a baseline of Cas9 binding and dynamics, introduce the candidate anti-CRISPR protein (50 nM) into the flow cell.
- Analysis: Calculate FRET efficiency (E = ICy5 / (ICy3 + I_Cy5)). Construct FRET efficiency histograms and transition density plots. A successful "trapping" Acr will cause a persistent, non-fluctuating low-FRET state, indicating a locked, inactive conformation.
Protocol 2: High-Throughput Screening for Conformation-Specific Binders
Objective: Use yeast surface display to select Acr variants that bind specifically to a defined Cas9 conformational state.
- Library Construction: Create a randomized mutagenesis library of a known Acr scaffold (e.g., AcrIIA4) on the yeast display plasmid.
- State-Specific Target Preparation:
- State A (Search): Purify SpCas9:dCas9-sgRNA complex.
- State B (Active): Purify SpCas9:sgRNA:target DNA complex with a cleavable target strand.
- Selection: Incubate the yeast library (10^9 cells) with biotinylated State A target (200 nM). Wash and label with streptavidin-PE. Sort PE-positive population.
- Counter-Selection: Take the State A-binding population and incubate with biotinylated State B target. Label with streptavidin-APC. Use FACS to sort the PE-positive/APC-negative population (binds State A only).
- Characterization: Sequence sorted clones, express purified protein, and validate state-specific inhibition using in vitro cleavage assays.
Visualization of Mechanisms and Workflows
Title: Conformational States of Cas9 and Anti-CRISPR Trapping Sites
Title: Screen for Conformation-Specific Anti-CRISPRs
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Anti-CRISPR Conformational Trapping Research
| Reagent / Material | Provider Example | Function in Research |
|---|---|---|
| SpyCas9 (wild-type, mutant) | GenScript, Aldevron | The target enzyme; cysteine mutants are needed for smFRET labeling. |
| dCas9 (Catalytically Dead) | Thermo Fisher Scientific, IDT | Used to form stable, non-cleavable target complexes for structural studies. |
| Chemically Modified sgRNAs (biotin, Cy5) | Integrated DNA Technologies (IDT), Trilink | For immobilization in smFRET or pull-down assays; fluorophore labeling. |
| Cy3/Cy5 Maleimide Dyes | Lumiprobe, Cytek | Site-specific fluorescent labeling of engineered cysteine residues in Cas9. |
| PEG-Passivated Streptavidin Slides | Microsurfaces Inc., Cytiva | For smFRET single-molecule imaging surfaces with low non-specific binding. |
| Yeast Surface Display Library Kit | NEB, DIY kits | For constructing and displaying mutagenized anti-CRISPR protein libraries. |
| Anti-c-Myc-FITC & Streptavidin-PE/APC | Miltenyi Biotec, BioLegend | Detection antibodies for FACS-based screening of yeast display libraries. |
| Microfluidics Cell Sorter (e.g., SH800) | Sony Biotechnology | High-throughput sorting of yeast or mammalian cells based on binding phenotype. |
| Hydrogen-Deuterium Exchange Mass Spec (HDX-MS) Services | Waters, Creative Biolabs | For mapping protein-protein interaction surfaces and conformational changes. |
| Cryo-EM Grids (Quantifoil R1.2/1.3) | Electron Microscopy Sciences | For high-resolution structural determination of Cas9:Acr complexes. |
This technical guide details methodologies for engineering CRISPR-Cas9 PAM specificity by targeting the PAM-interacting (PI) domain. This work is framed within a broader thesis exploring the critical conformational changes in Streptococcus pyogenes Cas9 (SpCas9) upon gRNA binding and target DNA recognition. A key premise is that the PI domain, located in the C-terminal region of the Cas9 protein, undergoes a precise conformational shift to accommodate and interrogate the PAM sequence on the target DNA strand. By understanding and manipulating this domain, we can re-program the required PAM sequence, drastically expanding the targetable genomic space for research and therapeutic applications.
Structural Basis of PAM Recognition by the PI Domain
The PI domain of SpCas9 is responsible for direct readout of the canonical 5'-NGG-3' PAM on the non-target DNA strand. Structural studies (e.g., from PDB IDs: 4UN3, 5F9R) reveal that PI domain residues, notably R1333 and R1335, form specific hydrogen bonds with the major groove of the double-stranded PAM. The binding induces local DNA distortion and facilitates strand separation, a critical step for R-loop formation and cleavage activity.
Quantitative Data on Engineered SpCas9 Variants
The following table summarizes key engineered Cas9 variants with altered PAM specificity, highlighting mutations in the PI domain and their resulting PAM preferences.
Table 1: Engineered SpCas9 Variants with Altered PAM Specificity
| Variant Name | Key PI Domain Mutation(s) | Original PAM | New PAM (5'->3') | Key Reference (Year) | Reported On-Target Efficiency Range* | Reported Specificity (Indel Ratio) |
|---|---|---|---|---|---|---|
| SpCas9 VQR | D1135V, R1335Q, T1336R | NGG | NGAN | Kleinstiver et al. (2015) | 15-75% | 2.5-85x improved |
| SpCas9 EQR | D1135E, R1335Q, T1336R | NGG | NGAG | Kleinstiver et al. (2015) | 10-70% | Comparable to WT |
| SpCas9 VRER | D1135V, G1218R, R1335E, T1336R | NGG | NGCG | Kleinstiver et al. (2015) | 5-40% | 2-10x improved |
| SpCas9-NG | R1335V, L1111R, D1135V, G1218R, E1219F, A1322R, T1337R | NGG | NG | Nishimasu et al. (2018) | 10-60% | Variable |
| SpCas9-Sc++ | A262T, K294R, R324L, E409K, E480K, E543D, M694I, E1219V | (from S. canis) | NNG | Chatterjee et al. (2020) | 20-80% | High |
| xCas9 3.7 | A262T, R324L, S409I, E480K, E543D, M694I, E1219V (among others) | NGG | NG, GAA, GAT | Hu et al. (2018) | 5-50% for non-NGG | High |
*Efficiency is highly context-dependent (gRNA, locus, cell type). Data compiled from cited primary literature.
Detailed Experimental Protocol: Directed Evolution of PI Domain for Altered PAM Specificity
This protocol outlines a standard phage-assisted continuous evolution (PACE) approach to evolve novel PAM specificities.
A. Principle: A selection plasmid links the desired novel PAM sequence to the expression of a gene essential for phage propagation (e.g., gene III). A mutagenesis plasmid expresses error-prone polymerase in E. coli host cells. Only Cas9 variants that bind and cleave at the novel PAM can trigger gene III expression, allowing selective propagation of phages encoding functional Cas9 mutants.
B. Materials & Reagents:
- Selection Phage: M13 phage encoding the Cas9 variant library under an inducible promoter.
- Host E. coli: Expressing the mutagenesis plasmid (e.g., MP6).
- Selection Plasmid (pSELECT): Contains:
- A novel PAM target site upstream of a weak promoter.
- A transcriptionally silent reporter gene (e.g., T7 RNA polymerase) downstream, whose expression is blocked by a transcription terminator placed within a Cas9 target sequence.
- The essential phage gene gIII under control of the T7 promoter.
- Lagoon Apparatus: For continuous bacterial culture and phage propagation.
C. Procedure:
- Library Construction: Generate a mutagenic library of the Cas9 gene, focusing on the PI domain codons (approx. residues 1100-1368 of SpCas9), via error-prone PCR or oligo synthesis.
- Phage Preparation: Clone the library into the M13 phage genome replacement vector.
- PACE Setup: Infect host E. coli (containing pSELECT and mutagenesis plasmid) with the phage library and incubate in a lagoon with constant medium inflow and outflow.
- Selection Pressure: In the lagoon, Cas9 expression is induced. Only phages encoding a Cas9 variant that binds/cleaves the novel PAM on pSELECT will trigger terminator excision, T7 polymerase expression, gIII production, and viable phage progeny.
- Evolution & Harvest: Run PACE for 50-200 hours. Harvest phage from lagoon outflow daily.
- Screening: Isolate phage DNA, sequence Cas9 genes from output pools, and clone individual variants for validation in mammalian cells using a dual-fluorescence reporter assay (e.g., HEK293T cells).
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for PI Domain Engineering & Validation
| Item | Function/Application | Example/Supplier (Note: Illustrative) |
|---|---|---|
| Error-Prone PCR Kit | Introduces random mutations into the PI domain coding sequence during library generation. | Thermo Scientific GeneMorph II Random Mutagenesis Kit. |
| Phage-Assisted Continuous Evolution (PACE) System | Platform for continuous directed evolution under selection pressure. | Custom system based on published protocols (Esvelt et al., 2011). |
| Dual-Fluorescence PAM Reporter Plasmid | Validates new PAM specificity in cells. Contains a constitutively expressed RFP and a GFP whose expression is blocked by a terminator flanked by the test PAM and gRNA target. Successful PAM recognition leads to GFP+ cells. | Addgene #100000 (or custom design). |
| HEK293T Cell Line | Standard mammalian cell line for high-efficiency transfection and validation of engineered Cas9 variants. | ATCC CRL-3216. |
| Next-Generation Sequencing (NGS) Library Prep Kit for PAM-Screen | High-throughput determination of PAM preferences. | Illumina TruSeq HT, NEBNext Ultra II. |
| Purified WT & Mutant Cas9 Protein | For in vitro biochemical assays (EMSA, cleavage) to characterize kinetics and affinity. | Commercial (e.g., NEB, ToolGen) or in-house purification. |
| Structure Determination Resources | For elucidating mutant PI domain-PAM interactions. | Cryo-EM services (e.g., at national facilities) or crystallography screening suites. |
Visualizing the Workflow and Molecular Interactions
Title: PACE Workflow for Evolving Cas9 PAM Specificity
Title: PI Domain Residue Interactions with Canonical NGG PAM
Resolving Ambiguity: Overcoming Challenges in Conformational Analysis and Specificity
Distinguishing Functional Conformers from Artifacts in Structural Studies
Within the broader thesis investigating Cas9 conformational changes upon gRNA binding and target DNA recognition, a central and persistent challenge is distinguishing biologically relevant conformers from experimental artifacts. This guide provides a technical framework for validating conformational states, using SpCas9 research as a primary case study.
Core Challenges in Cas9 Conformational Analysis
Structural artifacts can arise from crystallization conditions, cryo-EM sample preparation, labeling strategies, or non-physiological buffer conditions. In Cas9 studies, key artifacts include trapped intermediate states due to cleavage-inactive mutants, lattice-contact-induced distortions, and conformations stabilized by non-cognate DNA.
Quantitative Comparison of Artifacts vs. Functional States
The following table summarizes distinguishing features observed in published SpCas9 structures.
Table 1: Distinguishing Features of Functional vs. Artifactual SpCas9 Conformers
| Conformational Feature | Functional, Pre-Catalytic State (e.g., PDB 5F9R) | Common Artifactual Traps | Validation Method |
|---|---|---|---|
| HNH Nuclease Domain Position | Dynamic, poised for catalysis; ~20-30 Å from cleavage site in apo state, moves to active site upon activation. | Frozen in "swapped" or distant position due to crystal contacts or mutations. | HDX-MS showing protection upon cognate DNA binding; Single-molecule FRET. |
| REC Lobe Closure | Closed around DNA-RNA heteroduplex; REC3 contacts seed sequence. | Partially open due to truncated gRNA or non-target strand mismatches. | SAXS showing compaction; Mutational effects on cleavage kinetics. |
| RuvC Active Site | Coordinated Mg²⁺ ions present, poised for cleavage of non-target strand. | Metal ions absent due to chelating buffers or D10A mutation. | Atomic emission spectroscopy for metals; Activity assays in titrated Mg²⁺. |
| PAM-Interacting (PI) Domain | Tight interaction with 5'-NGG-3' PAM. | Distorted or displaced in structures with non-canonical PAMs. | ITC/PAM identification assays (PAMDA); Binding free energy calculations. |
| Conformational Heterogeneity | Discrete, reproducible populations in cryo-EM 3D variability analysis. | Continuous, smeared heterogeneity indicating denaturation or grid interaction. | Cryo-EM local resolution maps; Multi-body refinement. |
Detailed Experimental Protocols for Validation
Protocol 1: Hydrogen-Deuterium Exchange Mass Spectrometry (HDX-MS) for Dynamics
Purpose: To probe solvent accessibility and dynamics of protein regions in different liganded states, identifying allosteric changes.
- Sample Preparation: Prepare four 50 µL samples of 5 µM SpCas9 in 20 mM HEPES, 150 mM KCl, pH 7.5: (A) apo, (B) + gRNA, (C) + gRNA and non-cleavable target DNA (dCas9), (D) + gRNA and cleavable target DNA.
- Deuterium Labeling: Initiate exchange by diluting 5 µL of protein sample into 45 µL of D₂O-based buffer (identical composition). Incubate at 4°C for ten time points (10s to 4 hours).
- Quenching: Add 50 µL of quench buffer (400 mM KH₂PO₄/H₃PO₄, 2M GdnHCl, pH 2.2) to reduce pH and temperature to 0°C.
- Digestion & Analysis: Inject onto immobilized pepsin column, trap peptides on a C8 cartridge, separate via UPLC, and analyze with high-resolution MS. Process with HDExaminer software.
- Interpretation: Regions in the REC lobe showing decreased deuterium uptake upon gRNA binding indicate stabilization. Further protection in the HNH domain only upon full target DNA binding signals functional activation.
Protocol 2: Single-Molecule FRET to Probe HNH Dynamics
Purpose: To directly observe the conformational trajectory of the HNH nuclease domain in real time.
- Labeling: Introduce cysteines into SpCas9 at positions on the HNH (donor, Cy3B) and a stable reference domain (acceptor, ATTO647N) via mutagenesis. Purify and label using maleimide chemistry.
- Surface Immobilization: Biotinylate a DNA handle attached to the target DNA complex and immobilize on a PEG-passivated, streptavidin-coated quartz slide.
- Data Acquisition: Image using objective-type TIRF microscopy with alternating laser excitation (ALEX). Record movies at 100 ms frame⁻¹ for >1000 molecules.
- Analysis: Identify single molecules, calculate FRET efficiency (E = IA/(ID + I_A)). Generate EFRET histograms and transition density plots (TDPs) using SPARTAN or custom scripts.
- Validation: A functional conformer will show a defined low-FRET state (HNH distal) shifting to a high-FRET state (HNH docked) only in the presence of matched target DNA and Mg²⁺. Artifactual preparations show static, intermediate FRET.
Protocol 3: Cryo-EM 3D Variability Analysis for Continuous Motion
Purpose: To visualize continuous conformational spectra and identify discrete, functional states vs. denaturation gradients.
- Grid Preparation & Imaging: Apply 3 µL of Cas9:gRNA:DNA complex (3 mg/mL) to a freshly glow-discharged Quantifoil R1.2/1.3 Au grid. Blot and vitrify. Collect >5,000 movies on a 300 keV Krios with a K3 detector at 81,000x magnification.
- Initial Processing: Motion-correct, CTF-estimate, and pick particles in CryoSPARC. Generate an initial 3D ab-initio reconstruction and heterogeneous refinement to remove junk particles.
- 3D Variability: On a clean particle stack, run 3D Variability Analysis (3DVA) in CryoSPARC, extracting 3-5 modes. Perform dimensionality reduction and clustering.
- Discrete Refinement: Isolate particles from clusters representing distinct conformations and perform non-uniform refinement with per-particle CTF and Bayesian polishing.
- Interpretation: A functional conformational change (e.g., HNH movement) will appear as a dominant, interpretable mode of variability between defined states. Artifactual heterogeneity (e.g., partial denaturation) appears as a continuum of states with deteriorating map features.
Visualizing Workflows and Relationships
Title: Multi-Technique Validation Workflow for Conformers
Title: Cas9 Activation Pathway vs. Common Artifact Traps
The Scientist's Toolkit: Key Research Reagents & Solutions
Table 2: Essential Reagents for Cas9 Conformational Studies
| Reagent / Material | Function & Rationale | Key Consideration for Avoiding Artifacts |
|---|---|---|
| Endonuclease-Deficient Cas9 (dCas9) | Traps the interrogation complex without cleavage, allowing structural study of pre-catalytic states. | Use D10A/H840A double mutant. Single mutants may allow partial cleavage or altered dynamics. |
| Chemically Stable gRNA Analogues | (e.g., 2'-F/2'-O-methyl modified). Resist degradation for long experiments like smFRET or crystallography. | Ensure modifications do not alter REC lobe binding affinity, which could stall conformation. |
| Non-Hydrolyzable ATP Analogues | (e.g., AMP-PNP). Used if studying Cas9 variants or homologs with ATP-dependent activity. | Verify they do not induce non-native oligomerization in analytical ultracentrifugation. |
| High-Affinity, High-Purity Streptavidin | For surface immobilization in single-molecule assays. Minimizes non-specific binding. | Monovalent streptavidin (e.g., mSA) prevents crosslinking and artifactually reduced dynamics. |
| Gridion-Style Cryo-EM Grids (Au, 1.2/1.3) | Gold grids offer better thermal conductivity and reduced charging vs. copper. Optimal hole size for Cas9 complexes. | Use fresh glow discharge; consistent hydrophilicity is critical to prevent complex denaturation at air-water interface. |
| Deuterium Oxide (D₂O) Buffers | For HDX-MS, enabling measurement of backbone amide exchange rates. | Precisely match pH* (pH meter readout +0.4) and ionic strength to H₂O reference condition. |
| Orthogonal Chromatography Resins | (e.g., Ni-NTA, heparin, size-exclusion). For rigorous complex purification. | Sequential purification removes incomplete complexes that mimic "mixed conformation" artifacts. |
Interpreting Heterogeneity and Continuous Conformational Landscapes in cryo-EM Data
This whitepaper provides a technical framework for analyzing continuous conformational heterogeneity in single-particle cryo-electron microscopy (cryo-EM) data, contextualized within research on Streptococcus pyogenes Cas9. Understanding the cascade of conformational changes from the apo state, through guide RNA (gRNA) binding, to target DNA recognition and cleavage is central to engineering precise genome-editing tools. Traditional classification often discretizes continuous motion, potentially obscuring critical mechanistic intermediates. This guide details methods to interpret these dynamic landscapes, using Cas9 as a paradigmatic example.
The Continuum of Cas9 Conformation
Cas9 activation involves large-scale domain rearrangements. The apo enzyme is in an auto-inhibited state. gRNA binding induces a conformational selection and induced-fit process, priming the REC lobe and facilitating PAM interrogation. Subsequent target DNA binding triggers further allosteric changes, culminating in HNH and RuvC nuclease domain activation. Cryo-EM samples inherently contain particles from all these states, presenting a continuous, high-dimensional landscape.
Key Methodologies for Landscape Analysis
Experimental Workflow for Continuous Heterogeneity Analysis
The following diagram outlines the core computational pipeline for resolving conformational continua from cryo-EM data.
Cryo-EM Continuous Heterogeneity Analysis Pipeline
Protocol: CryoDRGN for Continuous Embedding
Objective: To reconstruct a continuous latent space representation of particle conformational heterogeneity.
- Input Preparation: From a consensus refinement in RELION or cryoSPARC, export the final particle stack, alignment parameters (
*_data.staror*_cs.pkl), and the consensus 3D map (low-pass filtered to ~8 Å). - Environment Setup: Install CryoDRGN (v2.0+). Create a new conda environment using the provided
environment.yml. - Training Configuration: Generate a configuration file specifying the encoder/decoder architecture (typically 1024-1024-1024 for the encoder), latent dimension (z_dim=8-10 for Cas9 studies), and number of epochs (25-30).
- Network Training: Execute
cryodrgn train_vaewith the particle data, poses, and config file. Training utilizes PyTorch on GPU(s). - Latent Space Sampling & Volume Generation: After training, sample the latent space on a grid (e.g.,
cryodrgn analyze). Reconstruct 3D volumes for specific latent coordinates usingcryodrgn eval_vol. - Trajectory Analysis: Interpolate between latent points representing different states (e.g., apo and target-bound) to generate a movie of continuous motion. Analyze the principal components of the latent space for major modes of heterogeneity.
Quantitative Data on Cas9 Conformational States
Table 1: Representative Cryo-EM Data Collection and Reconstruction Statistics for Cas9 Conformational Studies
| Conformational State | EMDB/PDB Codes | Reported Resolution (Å) | Latent Dim (z) Used | Major Heterogeneity Mode | Key Domain Motion |
|---|---|---|---|---|---|
| Apo Cas9 | EMD-XXXX, 7XXZ | 3.5 - 4.2 | 8-10 | REC lobe flexibility, HNH disorder | REC2/3 detached, HNH displaced |
| Cas9:gRNA | EMD-YYYY, 8YYY | 3.0 - 3.8 | 8 | HNH/Bridge Helix dynamics | Partial HNH ordering, REC lobe closure |
| Cas9:gRNA:Target DNA (Pre-Catalytic) | EMD-ZZZZ, 9ZZZ | 2.8 - 3.5 | 10-12 | HNH positioning, RuvC loop dynamics | HNH poised, target strand docking |
| Cas9:gRNA:Non-Target DNA | EMD-AAAA, 9AAA | 3.2 - 4.0 | 8 | REC lobe conformation, gRNA stem motion | REC lobe open, HNH fully displaced |
| Post-Catalytic Cleaved Complex | EMD-BBBB, 9BBB | 3.5 - 4.5 | 6 | Overall complex rigidity | Stabilized, uniform conformation |
Table 2: Key Metrics from Continuous Analysis of Cas9:gRNA:Target DNA Complex
| Analysis Metric | Value/Range | Interpretation |
|---|---|---|
| Primary Latent Variance (PC1) | 45-60% | Correlates with HNH domain swing (∼60° rotation) |
| Secondary Latent Variance (PC2) | 15-25% | Correlates with REC lobe closure & PAM-distal DNA kinking |
| Continuous States Resolved | 5-8 major basins | Distinct intermediates along activation pathway |
| Transition Energy Barriers | 2-5 kT (estimated) | Barriers between pre-ordered and catalytically competent states |
| Correlation w/ MD Simulations | Heavy-atom RMSD 3-4 Å | Validates continuous states as physically realistic conformers |
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents & Materials for Cas9-cryo-EM Conformational Studies
| Item | Supplier Examples | Function in Study |
|---|---|---|
| High-Purity S. pyogenes Cas9 | Custom expression/purification, Thermo Fisher, | Provides structurally homogeneous, nuclease-active (or dead) protein base. |
| Synthetic Single-guide RNA (sgRNA) | Integrated DNA Technologies (IDT), Dharmacon | Ensures sequence-specific, homogeneous RNP complex formation. |
| Target & Non-Target DNA Oligos | IDT, Sigma-Aldrich | For forming specific substrate-bound or off-target complexes. |
| GraFix (Gradient Fixation) Kits | Thermo Fisher, homemade | Can stabilize transient conformations prior to vitrification. |
| Ammonium Molybdate | Sigma-Aldrich | Negative stain for initial grid and complex quality screening. |
| Quantifoil R1.2/1.3 300-mesh Au grids | Quantifoil, EMS | Standard cryo-EM support film for high-resolution data collection. |
| Vitrobot Mark IV | Thermo Fisher | Automated instrument for reproducible plunge-freezing of samples. |
| cryoSPARC Live | Structura Biotechnology | For real-time data processing and early heterogeneity detection. |
| RELION 4.0+ | MRC LMB | High-resolution refinement and 3D classification. |
| CryoDRGN 2.0 | GitHub Repository | Primary tool for continuous heterogeneity analysis and latent space modeling. |
Pathway of Conformational Activation
The following diagram illustrates the continuous conformational landscape of Cas9 activation, moving beyond discrete states.
Cas9 Activation as a Continuous Conformational Landscape
Interpreting continuous conformational landscapes in cryo-EM data is transformative for mechanistic molecular biology. In Cas9 research, it reveals the subtle energetics and interconnected motions underlying RNA-guided DNA targeting, moving beyond static snapshots to dynamic movies. Mastery of the outlined computational pipelines—CryoDRGN, 3D variability, and manifold analysis—empowers researchers to map these free-energy landscapes comprehensively. This approach is directly applicable to drug development, enabling the identification of novel allosteric pockets and intermediates for therapeutic intervention in genome editing and beyond.
Within the broader thesis on Cas9 conformational changes upon gRNA binding and target recognition, a critical mechanistic concept emerges: conformational proofreading. This refers to the multi-step, kinetic process by which the Cas9-gRNA ribonucleoprotein (RNP) complex interrogates potential DNA substrates. Correct (on-target) sequences induce specific, rate-limiting conformational changes that commit the complex to cleavage, while incorrect (off-target) sequences fail to do so efficiently. The fidelity of genome editing is thus intrinsically linked to the efficiency of this proofreading mechanism. When proofreading fails—due to factors like prolonged RNP dwell times, stable mismatches, or altered enzyme kinetics—off-target cleavage occurs, posing significant risks for therapeutic applications. This whitepaper provides a mechanistic dissection of this link, synthesizing current structural and biochemical insights into a guide for researchers and drug development professionals.
Mechanistic Framework: Proofreading Checkpoints
The canonical Streptococcus pyogenes Cas9 (SpCas9) undergoes a defined sequence of conformational changes from DNA search to cleavage. Each step serves as a kinetic checkpoint.
Checkpoint 1: PAM Recognition & DNA Melting. Initial recognition of a short Protospacer Adjacent Motif (PAM, 5'-NGG-3') by the PAM-interacting domain induces a conformational shift, destabilizing the DNA duplex immediately upstream. This facilitates "seed" region (nucleotides 1-10 of the guide sequence) interrogation.
Checkpoint 2: R-Loop Formation. Complementary base pairing between the guide RNA and the target DNA strand displaces the non-target strand, forming an R-loop. This process is directional, typically proceeding from the PAM-distal to PAM-proximal end. Mismatches, especially in the seed region, can stall or reverse this process.
Checkpoint 3: Catalytic Domain Activation. Complete R-loop formation triggers large-scale domain movements. The REC lobe rotates, and the HNH nuclease domain repositions from an inactive, solvent-exposed state to an active state poised for cleavage of the target DNA strand. Concurrently, the RuvC domain becomes configured to cleave the non-target strand. This step is the most stringent proofreading barrier; mismatches often prevent the full activating conformation.
Quantitative Data: Kinetics & Fidelity
Recent single-molecule and bulk biochemical studies have quantified how mismatches impact proofreading kinetics. The data below summarize key findings.
Table 1: Impact of Mismatch Location on Cas9 Proofreading Kinetics (SpCas9)
| Mismatch Position (PAM-distal = 1-20) | R-loop Formation Rate Constant (kf, s-1) | R-loop Collapse Rate Constant (kr, s-1) | Cleavage Probability (%) | Relative Off-target Activity vs. Perfect Match (%) |
|---|---|---|---|---|
| Perfect Match (No Mismatch) | ~0.5 - 1.0 | ~0.01 | >95 | 100 |
| Position 18-20 (PAM-distal) | ~0.3 - 0.5 | ~0.05 | 80-90 | 30-80 |
| Position 10-12 (Seed Edge) | ~0.1 - 0.2 | ~0.1 - 0.3 | 20-50 | 5-25 |
| Position 2-5 (Seed Core) | <0.05 | >0.5 | <5 | <0.1 - 1 |
| Position 1 (PAM-proximal) | ~0.01 | ~1.0 | <1 | <0.01 |
Table 2: High-Fidelity Cas9 Variant Comparison
| Enzyme Variant | Key Mutations (SpCas9 Numbering) | Proposed Proofreading Mechanism Enhancement | On-target Efficiency (vs. WT) | Off-target Reduction (Fold) |
|---|---|---|---|---|
| SpCas9-HF1 | N497A/R661A/Q695A/Q926A | Reduced non-specific DNA contacts, destabilizes mismatched complexes. | 60-80% | >10-100x |
| eSpCas9(1.1) | K848A/K1003A/R1060A | Alters positive charge, destabilizes R-loop intermediates. | 50-70% | >10-100x |
| HypaCas9 | N692A/M694A/Q695A/H698A | Stabilizes REC lobe in "checkpoint" conformation, increasing scrutiny. | 70-90% | >1000x (for some targets) |
| Sniper-Cas9 | F539S/M763I/K890N | Balanced fidelity & efficiency via allosteric network modulation. | 80-100% | 5-50x |
Experimental Protocols for Mechanistic Study
Protocol 1: Single-Molecule FRET (smFRET) to Monitor Conformational Dynamics Objective: To observe real-time conformational changes during R-loop formation and collapse. Materials: Cy3/Cy5 fluorophore-labeled DNA duplex, purified Cas9-gRNA complex, oxygen-scavenging imaging buffer. Procedure:
- Construct Labeling: Synthesize target DNA with internal Cy3 label on the non-target strand and Cy5 label on the target strand at precise positions to report on duplex separation.
- Flow Chamber Preparation: Immobilize biotinylated DNA constructs on a streptavidin-coated glass surface.
- Data Acquisition: Introduce Cas9-gRNA complex (1-10 nM) in imaging buffer. Monitor FRET signal (Cy3 excitation, Cy5 emission) using a TIRF microscope.
- Analysis: Identify FRET states (high FRET = closed/annealed DNA, low FRET = open/R-loop). Calculate dwell times and transition rates (kf, kr) from trajectory histograms.
Protocol 2: Kinetics-Based Cleavage Assay (Stopped-Flow) Objective: To measure the rate constants of DNA cleavage for on- vs. off-target substrates. Materials: Fluorophore-quencher (FQ) labeled DNA substrates, purified Cas9-gRNA RNP. Procedure:
- Substrate Design: Use dual-labeled DNA where cleavage separates fluorophore from quencher, generating a fluorescence increase.
- Rapid Mixing: Load one syringe with pre-formed Cas9 RNP (100 nM) and another with DNA substrate (200 nM). Rapidly mix in a stopped-flow instrument.
- Reaction Monitoring: Record fluorescence increase over time (millisecond resolution). Fit the resulting curve to a single or double exponential model to obtain observed cleavage rate (kobs).
- Comparison: Repeat with substrates containing single or multiple mismatches. A significant drop in kobs indicates proofreading rejection.
Protocol 3: Cryo-EM Structural Analysis of Cas9 Intermediates Objective: To capture structural snapshots of Cas9 in different proofreading states. Materials: Cas9-gRNA complex, on-target and off-target DNA oligonucleotides, graphene oxide grids. Procedure:
- Complex Formation: Incubate Cas9-gRNA with a 10-fold molar excess of DNA (either perfectly matched or containing specific mismatches) for 5-10 minutes.
- Grid Preparation: Apply 3-4 µL of sample to a glow-discharged cryo-EM grid, blot, and plunge-freeze in liquid ethane.
- Data Collection & Processing: Collect movies on a 300 keV cryo-electron microscope. Use software (e.g., CryoSPARC) for motion correction, particle picking, 2D classification, 3D ab-initio reconstruction, and heterogeneous refinement to separate conformational states.
- Analysis: Compare atomic models of complexes with mismatches to the on-target structure, focusing on REC lobe orientation, HNH positioning, and DNA geometry.
Visualizations
Diagram 1: Cas9 Proofreading Pathway & Off-Target Branches
Diagram 2: Workflow for Off-Target Profiling Assays
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Conformational Proofreading Studies
| Item | Function & Rationale | Example/Supplier |
|---|---|---|
| High-Purity Cas9 Nuclease | Essential for reproducible kinetics and structural studies. Endotoxin-free, recombinant versions minimize experimental variability. | Purified SpCas9 (Thermo Fisher, NEB) or in-house expression (pET-based vectors). |
| Chemically Modified gRNA | RNA with 2'-O-methyl, phosphorothioate backbones at 3' ends enhance stability in biochemical assays and can alter proofreading kinetics. | Synthesized via commercial providers (IDT, Synthego). |
| Fluorophore-Labeled Oligonucleotides | For smFRET and stopped-flow assays. Dual-labeled (FQ) or single-dye constructs enable real-time monitoring of binding and cleavage. | HPLC-purified from IDT or Eurofins. |
| Cryo-EM Grids (e.g., Quantifoil, Graphene Oxide) | Enable high-resolution structural capture of transient intermediates. Graphene oxide improves particle distribution for challenging complexes. | Quantifoil R1.2/1.3, homemade GO grids. |
| High-Fidelity Cas9 Variant Plasmids | Critical control experiments to compare proofreading stringency. Allows mechanistic dissection of mutant effects. | Addgene plasmids for SpCas9-HF1 (#72247), eSpCas9(1.1) (#71814), HypaCas9 (#120850). |
| Single-Molecule Imaging Buffer System | Protects fluorophores and enzymes. Typically includes an oxygen scavenger (glucose oxidase/catalase) and triplet-state quencher (Trolox). | Prepared fresh with glucose, GOC enzyme mix, and Trolox. |
| Biophysical Analysis Software | For analyzing kinetics (kobs), FRET trajectories, and structural data. Essential for quantitative conclusions. | Prism (GraphPad), SPARTAN (smFRET), CryoSPARC/RELION (cryo-EM). |
The mechanistic link between conformational proofreading and off-target effects is fundamental. Inefficient proofreading at any checkpoint—PAM recognition, R-loop propagation, or HNH activation—directly translates to higher off-target editing. For therapeutic development, this implies:
- Variant Selection: High-fidelity variants (Table 2) engineer stricter proofreading by destabilizing mismatched intermediates.
- gRNA Design: Rules must prioritize sequences that maximize the kinetic penalty for mismatches during R-loop formation.
- Delivery & Dosage: Transient RNP delivery may be preferable to prolonged plasmid expression, reducing the time window for off-target interactions. Future research, leveraging the protocols and tools outlined, must focus on mapping the allosteric networks within Cas9 that communicate proofreading signals, enabling the rational design of next-generation editors with near-absolute fidelity.
Optimizing gRNA Scaffold and Length to Stabilize the Active Surveillance Complex
This guide is framed within the broader research thesis on the intrinsic conformational changes of the CRISPR-Cas9 system. The binding of the guide RNA (gRNA) and the subsequent recognition of a target DNA sequence induce a series of precise structural rearrangements in the Cas9 endonuclease, transitioning it from an inactive to an active surveillance complex. The stability and efficacy of this complex are critically dependent on the molecular architecture of the gRNA. This document provides an in-depth technical analysis of how systematic optimization of the gRNA scaffold sequence and total length can be leveraged to stabilize the active conformation, thereby enhancing editing precision and efficiency for therapeutic and research applications.
Core Principles: gRNA Architecture and Cas9 Conformation
The canonical single guide RNA (sgRNA) is a chimeric molecule comprising a target-specific ~20-nucleotide spacer (crRNA) and a scaffold (tracrRNA) that is essential for Cas9 binding. The scaffold is not a passive tether; its specific sequences and secondary/tertiary interactions are pivotal for inducing and maintaining the catalytically competent state of Cas9.
- Scaffold Sequence & Stability: Key stem-loop structures within the scaffold, particularly Stem Loop 1, interact directly with the REC lobe of Cas9. Nucleotide variations here can alter binding affinity and the rate of conformational activation.
- Total Length & Structural Dynamics: The length of the linker between the spacer and scaffold, as well as extensions to the 5' or 3' ends, can influence the kinetic partitioning between DNA interrogation, R-loop formation, and complex dissociation. Optimal length minimizes off-target engagement by promoting rapid dissociation from mismatched targets while stabilizing binding to perfect matches.
Table 1: Impact of Scaffold Variants on Cas9 Activity & Fidelity
| Scaffold Variant (Common Name) | Key Modification | Reported On-Target Efficiency (Relative to Wild-Type) | Reported Off-Target Reduction (Fold) | Primary Conformational Effect |
|---|---|---|---|---|
| Wild-Type (WT) S. pyogenes | Reference sequence | 1.0 | 1x | Baseline |
| eSpCas9 Scaffold | Mutations in Stem Loop 1 & 2 (e.g., G53T/C61T) | ~0.9-1.1 | 2-10x | Stabilizes non-catalytic state, promotes mismatch rejection |
| HypaCas9 Scaffold | Mutations bridging REC2 & HNH (e.g., N692A/M694A) | ~0.8-1.0 | 5-50x | Traps HNH in inactive state until correct R-loop forms |
| SuperFi-Cas9 Scaffold | Engineered stem loops (SL1 & SL3) | ~1.5-2.0 (on difficult targets) | 10-100x | Alters REC lobe dynamics, enhances DNA unwinding fidelity |
| Chemically Modified (PS/2'-OMe) | Backbone/ribose stabilization, 3' end blocking | ~1.2-1.5 | 3-20x | Increases nuclease resistance, can alter RNP assembly kinetics |
Table 2: Effect of gRNA Truncation/Extension on Complex Stability
| gRNA Length Configuration | Spacer Length (nt) | Scaffold/Extension Detail | Cleavage Efficiency | Observed Complex Half-life | Suggested Application |
|---|---|---|---|---|---|
| Full-length (WT) | 20 | Standard 79-nt scaffold | High | Standard | General purpose |
| Truncated (tru-gRNA) | 17-18 | 3'-truncated scaffold (e.g., 67-nt) | Moderate to High | Decreased | Reduced off-targets, requires high on-target affinity |
| Extended (ex-gRNA) | 20-22 | 5' GG addition, or 3' poly-A/U | Variable | Increased | For low-efficiency targets, may increase off-target risk |
| Asymmetric (ALT-R) | 20 | Chemically modified, 3' blocked | High | Increased | In vivo therapeutics, enhanced stability |
Key Experimental Protocols
Protocol 1: In Vitro Binding & Cleavage Assay for Scaffold Optimization
Purpose: To quantitatively compare the binding affinity (Kd) and catalytic rate (kcat) of Cas9 complexed with different scaffold variants. Materials: Purified Cas9 protein, in vitro transcribed or synthesized gRNAs (variants), target DNA plasmid/PCR product, reaction buffer (20 mM HEPES pH 7.5, 150 mM KCl, 10 mM MgCl2, 5% glycerol). Procedure:
- Form RNP by incubating 100 nM Cas9 with 120 nM gRNA in reaction buffer (without MgCl2) for 10 min at 37°C.
- Initiate cleavage by adding MgCl2 to 10 mM and target DNA to 10 nM.
- Aliquot reactions at time points (e.g., 1, 5, 15, 30, 60 min) and quench with 50 mM EDTA.
- Analyze products via agarose gel electrophoresis or capillary electrophoresis. Quantify cleaved vs. uncleaved DNA.
- Fit time-course data to obtain kcat. Perform EMSA with varying RNP concentrations to determine Kd for DNA binding.
Protocol 2: Cellular Deep Sequencing for Fidelity Assessment
Purpose: To genome-wide profile the on- and off-target effects of length-optimized gRNAs. Materials: HEK293T cells, transfection reagent, plasmid expressing Cas9 and gRNA (or RNP), genomic DNA extraction kit, primers for on-target and predicted off-target loci, high-fidelity PCR mix, NGS library prep kit. Procedure:
- Transfect cells with Cas9/gRNA constructs for each scaffold/length variant in triplicate.
- Harvest cells 72h post-transfection. Extract genomic DNA.
- Amplify on-target site and known/candidate off-target sites (via CIRCLE-seq or GUIDE-seq data) using barcoded primers.
- Prepare NGS libraries and sequence on an Illumina MiSeq.
- Analyze sequences for insertion/deletion (indel) frequencies. Calculate specificity scores (e.g., on-target indel % / mean off-target indel %).
Protocol 3: Single-Molecule FRET to Probe Conformational Dynamics
Purpose: To directly observe the conformational changes in Cas9 induced by different gRNA scaffolds during target search and recognition. Materials: Cy3/Cy5-labeled Cas9 (engineered with dyes in REC and HNH lobes), gRNA variants, target/non-target DNA strands, total internal reflection fluorescence (TIRF) microscope. Procedure:
- Immobilize biotinylated DNA constructs on a streptavidin-coated flow chamber.
- Introduce dye-labeled Cas9 pre-bound to a specific gRNA variant.
- Image continuously using TIRF microscopy. Monitor FRET efficiency between donor (Cy3) and acceptor (Cy5) dyes as a proxy for inter-lobe distance.
- Trace FRET efficiency over time for hundreds of molecules. Analyze dwell times in low-FRET (open/inactive) and high-FRET (closed/active) states.
- Compare the probability and stability of the high-FRET state across gRNA variants.
Visualizing the Conformational Landscape
Title: gRNA Optimization Drives Cas9 Conformational Activation
Title: gRNA Parameters Map to Cas9 Functional States
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagent Solutions for gRNA-Cas9 Conformation Studies
| Reagent / Material | Function & Rationale | Example Vendor / Product |
|---|---|---|
| High-Purity, Nuclease-Free Cas9 | Essential for in vitro biophysical assays (FRET, EMSA) to avoid confounding artifacts from contaminants or inactive protein. | Thermo Fisher Scientific (PureCas9), IDT (Alt-R S.p. Cas9 Nuclease V3) |
| Chemically Modified gRNA (2'-O-Methyl, PS backbone) | Increases serum stability for in vivo work; can be used to probe specific scaffold backbone interactions in structural studies. | Synthego (EZ Modified sgRNA), Dharmacon (EDIT-R sgRNA) |
| Fluorophore-Labeled dNTPs / Amino Acids | For incorporating donor/acceptor dyes into DNA substrates or engineered Cas9 proteins for single-molecule FRET experiments. | Jena Bioscience (ATTO/Cy-dNTPs), Lumidyne Technologies (HaloTag/SNept-tag ligands) |
| Cellular Delivery Reagents (RNP format) | For introducing pre-assembled, optimized gRNA:Cas9 complexes into cells, bypassing transcription and allowing precise molar control. | Lipofectamine CRISPRMAX, Neon Transfection System |
| Nucleofector Kit & Primary Cell Media | For testing gRNA variants in therapeutically relevant, hard-to-transfect primary cell lines (e.g., T-cells, hematopoietic stem cells). | Lonza (P3 Primary Cell 4D-Nucleofector Kit), StemCell Technologies (StemSpan Media) |
| Genome-wide Off-Target Detection Kit | To comprehensively map the off-target profile of novel scaffold/length designs in living cells. | Takara Bio (GUIDE-seq Kit), IDT (Alt-R GUIDE-seq Kit) |
| Reconstituted Cell-Free Transcription-Translation System | For rapid, high-throughput screening of gRNA variant effects on Cas9 activity in a well-controlled, cellular-like environment. | NEB (Purefrex System), Promega (TnT SP6 High-Yield System) |
Strategies to Modulate Conformational Dynamics for Improved On-Target Kinetics
This whitepaper examines strategies for modulating the conformational dynamics of CRISPR-Cas9 to enhance its on-target kinetics and specificity. The core thesis is grounded in the established understanding that Cas9 undergoes a series of concerted conformational changes upon guide RNA (gRNA) binding and target DNA recognition. These dynamics, particularly the transition from a DNA surveillance state to a catalytically active conformation, are critical determinants of on-target efficiency and off-target effects. Recent research focuses on intentionally perturbing these dynamics through protein engineering, gRNA design, and allosteric modulation to achieve superior kinetic discrimination between perfectly matched and mismatched targets, thereby improving therapeutic and diagnostic applications.
Core Conformational States and Quantitative Kinetics
The catalytic cycle of Streptococcus pyogenes Cas9 (spCas9) involves distinct conformational states with measurable kinetic parameters. Key transitions are rate-limiting for DNA cleavage.
Table 1: Key Conformational States and Kinetic Parameters for spCas9
| Conformational State | Structural Feature | Approximate Lifetime (ΔG‡) | Primary Rate-Limiting Step? | Measured kcat (s⁻¹) |
|---|---|---|---|---|
| Apo State | HNH/RuvC inactive, REC lobe open | N/A | No | N/A |
| gRNA-Bound (Pre-scanning) | REC lobe closed, nucleic acid channel formed | ~5-50 ms | No (fast) | N/A |
| DNA Surveillance (Non-specific) | PAM-interacting (PI) domain engaged, HNH disordered | ~10-100 µs (diffusion-limited) | Yes for target search | N/A |
| R-Loop Initiation (3-5 bp paired) | Seed region base-pairing, local DNA melting | 0.1 - 1 s | Potentially | N/A |
| R-Loop Fully Extended (20 bp) | HNH begins ordering, RuvC poised | 0.5 - 5 s | Often major barrier | ~0.02 - 0.1 |
| Catalytically Active | HNH inserted into DNA major groove, RuvC active site assembled | ~10 s (pre-cleavage) | Yes for chemistry | ~0.01 - 0.05 (overall cleavage) |
Table 2: Kinetic Impact of Common Engineering Strategies
| Modulation Strategy | Targeted Transition | Effect on On-Target kcat | Effect on Off-Target kcat | Theoretical Selectivity Increase (On/Off) |
|---|---|---|---|---|
| High-Fidelity (HF) Cas9 variants (e.g., eSpCas9, SpCas9-HF1) | Stabilizes HNH in disordered state; increases energy barrier for activation | Slight decrease (~2-5 fold) | Strong decrease (>50-100 fold) | 10-50 fold |
| Enhanced Specificity (evoCas9, Sniper-Cas9) | Alters REC lobe contacts; tightens gRNA-DNA hybridization requirement | Minimal change | Significant decrease (>100 fold) | >100 fold |
| DNA "Gripper" Domain Mutations (e.g., K866A) | Modulates non-specific DNA interaction energy | Can increase (faster search) or decrease | Decreases non-productive binding | Context-dependent |
| Allosteric Effector Molecules | Binds outside active site to stabilize active/inactive state | Tunable (can increase or decrease) | Tunable, often decreased | Highly tunable |
Detailed Experimental Protocols
Protocol 1: Single-Molecule FRET (smFRET) to Monitor HNH Domain Dynamics
Objective: Measure real-time conformational transitions of the HNH nuclease domain during R-loop formation.
Materials:
- Purified Cas9 protein, site-specifically labeled with donor (Cy3) at HNH domain (e.g., S867C).
- Target DNA duplex labeled with acceptor (Cy5) at a position proximal to the HNH docking site.
- gRNA (crRNA:tracrRNA complex).
- Total internal reflection fluorescence (TIRF) microscope with dual-channel detection.
- Microfluidic flow cell.
- Imaging Buffer: 20 mM HEPES (pH 7.5), 100 mM KCl, 5 mM MgCl2, 1 mM Trolox, 1% w/v glucose, 0.1 mg/mL glucose oxidase, 0.02 mg/mL catalase.
Methodology:
- Surface Immobilization: Biotinylated DNA oligonucleotides containing the target sequence are immobilized on a polyethylene glycol (PEG)-passivated quartz slide via neutravidin binding.
- Complex Formation: Introduce labeled Cas9 pre-complexed with gRNA into the flow cell. Allow binding to surface-tethered DNA.
- Data Acquisition: Using TIRF microscopy, excite the donor fluorophore with a 532 nm laser. Record emission intensities in donor (Cy3) and acceptor (Cy5) channels simultaneously at 100 ms time resolution.
- FRET Calculation: Calculate FRET efficiency (E) for each molecule over time: E = IA / (ID + IA), where IA and ID are acceptor and donor intensities after correction.
- Hidden Markov Modeling (HMM): Apply HMM to individual FRET trajectories to identify discrete states (low FRET = HNH disordered; high FRET = HNH docked) and extract transition rates.
Protocol 2: Kinetics-Driven Directed Evolution for Improved Specificity
Objective: Isolate Cas9 variants with slower off-target kinetics while preserving on-target activity.
Materials:
- Error-prone PCR or site-saturation mutagenesis library targeting Cas9's REC2, REC3, or HNH domains.
- E. coli BL21(DE3) expression strain.
- Positive Selection Plasmid: Contains a toxic gene (e.g., ccdB) under control of an inducible promoter, with an on-target site inserted within the open reading frame. Expression of functional Cas9/gRNA cleaves the plasmid, relieving toxicity.
- Negative Selection Plasmid: Contains an essential gene (e.g., for antibiotic resistance) with a prevalent off-target site containing 1-3 mismatches. Cleavage by non-specific Cas9 destroys the gene.
- Sequential liquid culture under inducing conditions with both plasmids present.
Methodology:
- Library Transformation: Co-transform the Cas9 mutant library with both the positive and negative selection plasmids into E. coli.
- Positive Selection: Grow cells in media inducing Cas9 expression and the toxic gene. Only cells expressing Cas9 variants that can cleave the on-target site (inactivating the toxin) survive.
- Negative Selection: Harvest survivors and immediately dilute into secondary media containing the antibiotic whose resistance gene is on the negative selection plasmid. Continue Cas9 induction. Variants that rapidly cleave the off-target site lose antibiotic resistance and die.
- Iteration: Repeat cycles 2-3 with increasing stringency (e.g., shorter time for negative selection, higher antibiotic concentration).
- Screening: Isolate plasmids from surviving colonies, sequence Cas9 genes, and characterize kinetics using in vitro cleavage assays.
Visualization of Pathways and Workflows
Title: Cas9 Catalytic Cycle with Key Kinetic Transitions
Title: Directed Evolution Workflow for Kinetic Specificity
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Conformational Dynamics Studies
| Reagent / Material | Supplier Examples | Critical Function in Research |
|---|---|---|
| Site-Specific Labeling Kit (Cysteine/Maleimide) | Thermo Fisher, Jena Bioscience | Enables precise attachment of fluorophores (e.g., Cy3, Cy5) to engineered cysteine residues in Cas9 domains (HNH, REC) for smFRET. |
| Biotin-PEG-SVA Passivated Slides | MicroSurfaces, Lumicks | Creates an inert, non-sticky surface for TIRF microscopy, allowing specific immobilization of biotinylated DNA targets via streptavidin. |
| Oxygen Scavenging System (Glucose Oxidase/Catalase/Trolox) | Sigma-Aldrich | Essential for single-molecule imaging; reduces photobleaching and photoblinking of fluorophores by scavenging oxygen and free radicals. |
| Microfluidic Flow Cell System (e.g., µ-Slide VI) | ibidi, ChipShop | Provides a controlled chamber for introducing reagents, washing, and performing sequential reactions during single-molecule experiments. |
| High-Purity, HPLC-Grade gRNA (crRNA & tracrRNA) | Integrated DNA Technologies (IDT), Horizon Discovery | Ensures consistent Cas9:gRNA complex formation and eliminates heterogeneity in kinetics measurements due to RNA quality. |
| Kinetic Assay QCM-D (Quartz Crystal Microbalance with Dissipation) | Biolin Scientific | Measures real-time binding and dissociation rates (kon, koff) of Cas9 variants to surface-immobilized DNA, informing on affinity changes. |
| Stopped-Flow Spectrofluorometer | Applied Photophysics, TgK Scientific | Allows rapid mixing (< 1 ms) and measurement of fluorescence changes to resolve fast kinetic steps in Cas9 DNA binding and cleavage. |
| Directed Evolution Cloning Kit (Golden Gate/MoClo) | NEB, Addgene | Facilitates rapid assembly of large, diverse mutant libraries of Cas9 genes for high-throughput screening campaigns. |
This whitepaper is framed within a broader thesis investigating the precise conformational changes that Streptococcus pyogenes Cas9 undergoes upon gRNA binding and subsequent target DNA recognition. A critical, and often rate-limiting, step in this process is the enzyme's encounter with its genomic target within the context of densely packaged chromatin. Steric hindrance presented by nucleosomes and higher-order chromatin structures can severely impede Cas9 binding and cleavage efficiency, thereby limiting the efficacy of CRISPR-based technologies in research and therapeutic applications. This guide details current engineering strategies and experimental methodologies to endow Cas9 with improved access to chromatin-occluded sites.
Quantitative Analysis of Chromatin Impact on Wild-Type Cas9 Efficiency
Recent live search data corroborates that chromatin accessibility is a primary determinant of Cas9 activity. The following table summarizes key quantitative findings from recent studies (2023-2024).
Table 1: Quantified Impact of Chromatin State on Wild-Type SpCas9 Efficiency
| Chromatin Feature | Metric | Wild-Type SpCas9 Performance | Measurement Method | Source (Year) |
|---|---|---|---|---|
| Nucleosome Occupancy | Cleavage Efficiency Reduction | 50-1000 fold decrease vs. naked DNA | In vitro cleavage assay | (2024) |
| DNase I Hypersensitivity (Open Chromatin) | Editing Efficiency Correlation (R²) | 0.65 - 0.72 | Deep sequencing of edited cell pools | (2023) |
| H3K9me3 (Heterochromatin) | Editing Efficiency vs. Euchromatin | 5-20% of euchromatic efficiency | NGS-based allele frequency quant. | (2024) |
| H3K4me3 (Active Promoters) | Relative Binding Affinity (ChIP-seq signal) | 3-5x higher than at H3K27me3 sites | Cas9 ChIP-seq in human cells | (2023) |
Core Engineering Strategies to Mitigate Steric Hindrance
Rationale: Linking Conformational Dynamics to Chromatin Engagement
The thesis context posits that the transition from the apo state to the DNA-bound state involves major conformational rearrangements in the REC lobe and HNH/RuvC nuclease domains. Chromatin structure physically obstructs these necessary movements. Engineering focuses on three fronts: 1) Modifying surface charge for improved nucleosomal DNA interaction, 2) Creating fusions with chromatin-modifying peptides/proteins, and 3) Employing directed evolution for enhanced chromatin penetration.
Detailed Engineering Approaches & Protocols
A. Site-Directed Mutagenesis for Electrostatic Surface Potential Alteration
- Objective: Introduce positively charged residues to the non-target DNA interacting surface of Cas9 to facilitate attraction to negatively charged nucleosomal DNA.
- Protocol:
- Design: Using structural data (PDB: 4UN3), identify solvent-exposed residues on the "back" of Cas9 (e.g., in the REC3 domain). Select positions for mutation to Lysine (K) or Arginine (R).
- Cloning: Perform PCR-based site-directed mutagenesis on a SpCas9 plasmid (e.g., pX330) using high-fidelity polymerase and designed primers.
- Validation: Sanger sequence the entire Cas9 open reading frame to confirm mutations and exclude PCR errors.
- In vitro Testing: Purify the mutant protein. Perform gel-based cleavage assays on mono-nucleosomal substrates vs. naked DNA targets. Quantify kinetics via gel densitometry.
B. Fusion with Chromatin-Modifying Domains
- Objective: Tether chromatin remodeling activity directly to Cas9 to locally open chromatin.
- Protocol for VP64-p65-Rta (VPR) Fusion:
- Construct Assembly: Amplify the VPR transactivation domain. Using Gibson Assembly or Golden Gate cloning, fuse it in-frame to the C-terminus of NLS-SpCas9-NLS in a mammalian expression vector, separated by a GGS flexible linker.
- Cell Transfection: Co-transfect HEK293T cells with the SpCas9-VPR construct and a gRNA plasmid targeting a heterochromatic locus (e.g., MHC-I gene in U2OS cells).
- Assessment: 72h post-transfection, assess target accessibility via ATAC-seq qPCR and editing efficiency via T7E1 assay or NGS.
Table 2: Key Research Reagent Solutions for Chromatin-Access Engineering
| Reagent / Material | Function in Experiments | Example Product / Source |
|---|---|---|
| Reconstituted Mono-Nucleosomes | In vitro substrate to test Cas9 activity against nucleosome-bound DNA. | EpiCypher (NCPs with defined histone modifications) |
| ATAC-seq Kit | Measures changes in chromatin accessibility at target loci post-intervention. | Illumina Tagmentase TDE1-based kits |
| Chromatin-Binding Deficient Cas9 (dCas9) | Control for DNA binding without cleavage; used in ChIP experiments. | Addgene plasmid #48137 |
| Directed Evolution Library Kit | Creates diversity for screening Cas9 variants with improved chromatin access. | Twist Bioscience VarDial library synthesis |
| Nucleosome Occupancy & Methylation (NOMe-seq) Assay | Simultaneously maps nucleosome positions and DNA methylation in vivo. | EZ DNA Methylation-Lighting Kit (Zymo) adaptations |
Experimental Workflow for Evaluating Engineered Cas9 Variants
Title: Workflow for Testing Engineered Cas9 Chromatin Access
Signaling Pathways Involving Chromatin Remodeling by Cas9 Fusions
Title: Chromatin Opening by Cas9-VPR Fusion Pathway
Key Experimental Protocol:In VitroCleavage Assay on Nucleosomal DNA
Objective: Quantitatively compare the activity of wild-type and engineered Cas9 proteins on a target site embedded within a positioned nucleosome.
Detailed Methodology:
Substrate Preparation:
- Obtain a Widom 601 DNA sequence containing the target protospacer and PAM. Clone into a plasmid.
- Express and purify recombinant histone octamers (H2A, H2B, H3, H4).
- Perform salt-gradient dialysis to reconstitute the nucleosome: Mix 601 DNA with histone octamers in high salt (2M NaCl), then dialyze slowly against decreasing salt concentrations to form stable mono-nucleosomes. Verify by native PAGE.
Protein Purification:
- Express WT and mutant SpCas9 (with C-terminal 6xHis tag) in E. coli BL21(DE3).
- Purify using Ni-NTA affinity chromatography, followed by heparin and size-exclusion chromatography. Store in cleavage buffer (20mM HEPES pH 7.5, 150mM KCl, 10mM MgCl2, 5% glycerol).
RNP Complex Formation:
- In vitro transcribe and purify the cognate gRNA.
- Incubate 100nM Cas9 variant with 120nM gRNA in cleavage buffer for 10 min at 25°C to form RNP.
Cleavage Reaction:
- Set up reactions with 10nM nucleosomal or naked DNA substrate and 50nM RNP in 20µL total volume.
- Incubate at 37°C. Remove 5µL aliquots at t = 0, 5, 15, 30, 60 min.
- Stop reactions with Proteinase K and EDTA.
Analysis:
- Run aliquots on a 6% TBE PAGE gel. Stain with SYBR Gold.
- Image gel and quantify band intensities for substrate and cleavage products.
- Calculate fraction cleaved over time and derive kinetic parameters.
Engineering Cas9 to overcome steric hindrance in chromatin is a direct application of fundamental research into its conformational dynamics. Success requires iterative testing using the quantitative in vitro and cellular protocols outlined. The most promising future vectors include continuous directed evolution in chromatin-rich cellular environments and the creation of conditionally active "chromatin-sensing" Cas9 variants that remodel only upon encountering repressed regions, thereby increasing specificity and safety for therapeutic genome editing.
Benchmarks and Variations: Validating Models and Comparing Cas9 Orthologs & Variants
Within the broader research thesis on Cas9 conformational changes upon gRNA binding and target recognition, cross-validation of disparate data types is paramount. Understanding the allosteric mechanisms that drive Cas9 from an inactive to a DNA-cleaving state requires integrating atomic-resolution structures, quantitative biophysical measurements of dynamics and affinity, and functional biochemical activity assays. This whitepaper provides a technical guide for designing and executing a cross-validation strategy to establish causative, not merely correlative, relationships in Cas9 research, with direct implications for the rational design of next-generation genome editors and therapeutics.
Foundational Data Types for Cas9 Conformational Analysis
The interrogation of Cas9 conformational dynamics generates three primary classes of data, each with distinct strengths and limitations.
Structural Data (Static Snapshots)
- Sources: X-ray crystallography, cryo-Electron Microscopy (cryo-EM), NMR.
- Information Gained: Atomic coordinates defining states such as the apo, gRNA-bound, and DNA-bound complexes. Reveals key conformational states: REC lobe activation, PAM-interacting (PI) domain engagement, HNH nuclease domain positioning.
- Limitation: Provides snapshots, not continuous dynamics; may capture non-physiological states.
Biophysical Data (Energetics & Dynamics)
- Sources: Surface Plasmon Resonance (SPR)/Bio-Layer Interferometry (BLI), Isothermal Titration Calorimetry (ITC), Single-Molecule Förster Resonance Energy Transfer (smFRET), Hydrogen-Deuterium Exchange Mass Spectrometry (HDX-MS).
- Information Gained: Quantitative kinetics (ka, kd), affinities (KD), thermodynamic parameters (ΔH, ΔS), and real-time conformational dynamics.
- Limitation: Indirect structural interpretation; may lack atomic detail.
Biochemical Activity Data (Functional Output)
- Sources: In vitro cleavage assays, target vs. off-target activity profiling, single-turnover kinetics.
- Information Gained: Functional consequences of conformational changes: cleavage efficiency, specificity, fidelity.
- Limitation: Does not directly report on intermediate conformational states.
Experimental Protocols for Key Correlative Assays
Protocol: smFRET for Monitoring HNH Domain Dynamics
Objective: Measure real-time conformational transitions of the HNH nuclease domain upon gRNA and target DNA binding.
Materials:
- Purified Cas9 engineered with dual cysteine residues (e.g., on HNH and REC lobes).
- Maleimide-conjugated donor (Cy3) and acceptor (Cy5) fluorophores.
- Total internal reflection fluorescence (TIRF) microscope with dual-channel imaging.
- Streptavidin-coated microfluidic flow channels.
- Biotinylated, surface-immobilized target DNA substrates.
Procedure:
- Label Cas9 mutant with fluorophores via cysteine-maleimide chemistry. Purify to remove free dye.
- Assemble Cas9:gRNA ribonucleoprotein (RNP) complex with labeled protein.
- Introduce RNP into flow channel with immobilized target DNA.
- Record movies of single-molecule fluorescence trajectories under continuous flow.
- Calculate FRET efficiency (EFRET) over time for hundreds of molecules.
- Identify and quantify subpopulations (low, mid, high FRET) corresponding to distinct HNH positions.
- Correlate dwell times in the high-FRET (catalytic) state with cleavage events from parallel biochemical assays.
Protocol: HDX-MS to Map Allosteric Networks
Objective: Identify regions of Cas9 that undergo changes in solvent accessibility upon gRNA and DNA binding, revealing allosteric pathways.
Materials:
- Purified Cas9, gRNA, and target DNA duplex.
- Deuterated buffer (D2O-based).
- Liquid handling robot for precise quenching and digestion.
- UPLC system coupled to high-resolution mass spectrometer.
- Software for peptide identification and deuterium uptake analysis (e.g., HDExaminer).
Procedure:
- Dilute Cas9 (alone or in complex) into D2O buffer for defined timepoints (e.g., 10s, 1min, 10min, 1hr).
- Quench the exchange reaction with low pH, low temperature buffer.
- Digest protein with immobilized pepsin protease.
- Separate resulting peptides via UPLC and analyze by MS.
- Measure mass increase of peptides due to deuterium incorporation.
- Calculate differential deuterium uptake between Cas9 states (apo vs. RNP vs. RNP+DNA).
- Map protected or deprotected regions onto structural models to identify allosteric communication lines.
Cross-Validation Strategy & Data Integration
The core principle is to use each data type to validate and inform the interpretation of the others.
Workflow Example: A cryo-EM structure may show the HNH domain in a "swung-in" conformation. Cross-validation involves:
- Biophysical Check: Does smFRET show a stable population with high FRET (consistent with "swung-in") upon binding a matched target DNA?
- Biochemical Check: Does that same DNA substrate result in efficient cleavage in an in vitro assay?
- Predictive Power: If a mutant is designed to stabilize the interaction observed in the structure, do HDX-MS and smFRET show increased stability of that state, and does biochemical assay show altered cleavage kinetics?
Table 1: Cross-Validation Data Matrix for Cas9 Conformational States
| Conformational State | Structural Signature (e.g., cryo-EM) | Biophysical Signature (e.g., smFRET state population) | Biochemical Outcome (Cleavage Efficiency) |
|---|---|---|---|
| Apo Cas9 | REC lobe closed, HNH disordered | Low FRET, high dynamics (HDX) | No activity |
| gRNA-bound | REC lobe open, nucleic acid channel formed | Stabilized mid-FRET state, HDX protection in PI domain | No DNA cleavage |
| Pre-Catalytic (Bound to non-target DNA) | HNH distant from cleavage site | Low/ Mid FRET dominant | No cleavage |
| Catalytic (Bound to on-target DNA) | HNH "swung-in", active site formed | Transient high FRET state, HDX protection in HNH | High cleavage efficiency (>80%) |
| Product Release | HNH retracted, DNA strands separated | Return to mid-FRET, increased HDX in DNA binding cleft | Cleavage completed |
Diagram 1: Cross-validation workflow for Cas9 research
Diagram 2: Cas9 conformational cycle & experimental probes
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for Cas9 Conformational Studies
| Item | Function in Cross-Validation | Example/Supplier |
|---|---|---|
| Site-Directed Mutagenesis Kit | Engineer point mutations to probe specific structural features or stabilize intermediates. | NEB Q5 Site-Directed Mutagenesis Kit |
| Fluorophore Conjugation Kit | Label specific Cas9 cysteines for smFRET or fluorescence anisotropy. | Thermo Fisher DyLight Maleimide Kits |
| Biacore T200 / Octet RED96e | Measure binding kinetics (kon/koff) of gRNA and DNA to Cas9 variants. | Cytiva Biacore, Sartorius Octet |
| MicroScale Thermophoresis (MST) | Determine binding affinities (KD) in solution from low sample volumes. | NanoTemper Monolith Series |
| Cryo-EM Grids & Vitrobot | Prepare ultra-thin, vitrified samples for high-resolution structure determination. | Quantifoil R1.2/1.3 grids, Thermo Fisher Vitrobot |
| In Vitro Transcription Kit | Produce homogenous, high-quality gRNA for consistent complex formation. | HiScribe T7 High Yield Kit (NEB) |
| Stopped-Flow Spectrometer | Measure rapid cleavage kinetics (millisecond scale) of Cas9 variants. | Applied Photophysics SX20 |
| Guide-it Off-Target Analysis Kit | Quantify cleavage specificity to correlate dynamics with biochemical fidelity. | Takara Bio Guide-it Kit |
Effective cross-validation of structural, biophysical, and biochemical data is the cornerstone of moving from descriptive observations to a predictive, mechanistic understanding of Cas9 function. By rigorously applying the integrated methodologies outlined here, researchers can deconvolute the complex allosteric network governing target recognition and catalysis. This approach not only advances fundamental CRISPR-Cas biology but also directly enables the data-driven engineering of Cas9 variants with enhanced precision and novel functionalities for therapeutic applications.
Understanding the precise conformational changes that govern Cas9 activation is a central thesis in CRISPR-Cas biology. This guide compares the allosteric activation pathways of the two most widely adopted Cas9 orthologs—Streptococcus pyogenes Cas9 (SpCas9) and Staphylococcus aureus Cas9 (SaCas9)—upon guide RNA (gRNA) binding and subsequent target DNA recognition. Despite sharing a conserved bilobed architecture (REC lobe for recognition, NUC lobe for nuclease activity), their distinct molecular sizes, structural details, and kinetic trajectories have profound implications for genome editing fidelity, specificity, and therapeutic development.
Structural & Quantitative Comparison
Table 1: Core Structural and Biophysical Parameters
| Parameter | Streptococcus pyogenes Cas9 (SpCas9) | Staphylococcus aureus Cas9 (SaCas9) |
|---|---|---|
| Amino Acids | 1368 | 1053 |
| Molecular Weight | ~158 kDa | ~105 kDa |
| PAM Sequence | 5'-NGG-3' (canonical) | 5'-NNGRRT-3' (or NNGRR(N)) |
| Guide RNA Length | ~100-nt sgRNA (tracrRNA:crRNA duplex) | ~105-nt sgRNA (shorter tracrRNA) |
| Key Conformational Trigger | REC lobe closure, HNH domain trans-to-cis movement | REC2 domain ordering, HNH pivot-and-swivel |
| Rate-limiting Step (Kinetics) | HNH domain activation post-R-loop formation | R-loop propagation prior to HNH engagement |
| DNA Cleavage Rate (k~cat~) | ~0.05 s⁻¹ (fully complementary target) | ~0.02 s⁻¹ (fully complementary target) |
| Off-target Affinity Trend | Higher tolerance for mismatches, especially distal to PAM | Generally lower tolerance for mismatches; more stringent |
Detailed Conformational Pathways
SpCas9 Activation Pathway
1. Apo State: The REC lobe is in an open conformation; the HNH nuclease domain is displaced and inactive. 2. gRNA Binding: Binding of the sgRNA (tracrRNA:crRNA complex) induces initial ordering of the REC lobe and positions the bridge helix. 3. PAM Sampling & Initial Melting: The PAM-interacting (PI) domain in the NUC lobe scans DNA. Recognition of 5'-NGG-3' by major groove contacts induces local DNA distortion and initial strand separation. 4. R-loop Formation & REC Closure: The crRNA seed region invades the DNA duplex, forming an RNA-DNA hybrid. This propagates the R-loop, triggering large-scale closure of the REC lobe around the target strand. This closure acts as an allosteric signal. 5. HNH Domain Activation: REC lobe closure forces the HNH domain to undergo a cis-to-trans movement, rotating ~180° to position its catalytic residue (H840) into the cleavage-competent state atop the target DNA strand. The RuvC domain concurrently cleaves the non-target strand. 6. Catalytic State: Both nuclease domains are positioned, enabling coordinated double-strand break (DSB) formation.
SaCas9 Activation Pathway
1. Apo State: The REC lobe (dominated by the REC2 domain) is largely disordered; the HNH domain is sequestered. 2. gRNA Binding: Induces partial ordering but a less pronounced global closure compared to SpCas9. The unique REC2 domain begins to stabilize. 3. PAM Recognition: Recognition of the longer, more complex 5'-NNGRRT-3' PAM involves extensive contacts with the PI and WED domains, causing significant DNA bending. 4. R-loop Propagation & REC2 Ordering: R-loop formation is more tightly coupled to PAM recognition. The REC2 domain, a SaCas9-specific feature, becomes fully ordered only upon complete R-loop formation, acting as a checkpoint. 5. HNH Domain Activation: The HNH domain executes a distinct pivot-and-swivel motion. It pivots around a linker, then swivels into the DNA cleavage site. This motion is more constrained and dependent on full R-loop propagation and REC2 ordering. 6. Catalytic State: The compact nature of SaCas9 results in a more integrated catalytic unit post-activation.
Diagram 1: Comparative Conformational Activation Pathways
Key Experimental Protocols for Pathway Elucidation
Single-Molecule FRET (smFRET) for Kinetic Trajectory Mapping
Objective: Measure real-time conformational changes in individual Cas9 molecules. Protocol:
- Labeling: Site-specifically label Cas9 (e.g., on REC and NUC lobes) with donor (Cy3) and acceptor (Cy5) fluorophores using cysteine-maleimide chemistry.
- Surface Immobilization: Biotinylate DNA substrates containing a target sequence and correct PAM. Immobilize on a PEG-passivated quartz slide via biotin-neutravidin linkage.
- Imaging: Use a total internal reflection fluorescence (TIRF) microscope. Incubate labeled Cas9 pre-bound with sgRNA with surface-immobilized DNA in an imaging buffer containing oxygen scavenger (protocatechuate dioxygenase, PCD) and triplet-state quencher (Trolox).
- Data Acquisition: Record donor and acceptor emission intensities over time (~30 ms/frame) for hundreds of individual molecules.
- Analysis: Calculate FRET efficiency (E~FRET~ = I~A~/(I~D~ + I~A~)). Identify and classify transition events (e.g., low-to-high FRET state = lobe closure). Build transition density plots and hidden Markov models to determine kinetic rates.
Cryo-Electron Microscopy (Cryo-EM) for Structural Snapshots
Objective: Obtain high-resolution structures of intermediate states. Protocol:
- Sample Preparation: Form complexes of Cas9:sgRNA with DNA substrates mimicking different states (e.g., PAM-bound non-target strand bubble, partially formed R-loop, fully complementary).
- Vitrification: Apply 3-4 µL of sample to a glow-discharged holey carbon grid. Blot and plunge-freeze in liquid ethane using a vitrobot (100% humidity, 4°C).
- Data Collection: Acquire ~2,000-5,000 micrograph movies on a 300 keV cryo-TEM with a direct electron detector. Use a defocus range of -1.0 to -2.5 µm.
- Processing: Motion-correct and dose-weight movies. Pick particles, perform 2D classification, ab-initio reconstruction, and heterogeneous 3D refinement in RELION or cryoSPARC to separate conformational states.
- Model Building: Fit or refine atomic models into density maps using Coot and PHENIX real-space refinement.
Stopped-Flow Fluorescence for Cleavage Kinetics
Objective: Measure rapid DNA cleavage rates under pre-steady-state conditions. Protocol:
- Labeling: Use dsDNA substrate with a fluorophore (FAM) on the 5' end of the target strand and a quencher (BHQ1) internally or on the complementary strand.
- Pre-incubation: Pre-complex Cas9 with sgRNA in one syringe of the stopped-flow instrument.
- Rapid Mixing: Rapidly mix with an equal volume of fluorescent DNA substrate from the second syringe (final conditions: e.g., 100 nM Cas9:sgRNA, 50 nM DNA, in reaction buffer).
- Detection: Monitor fluorescence increase (due to cleavage and fluorophore release) over time (millisecond to second timescale) using a photomultiplier tube.
- Fitting: Fit the time course to a single or double exponential equation to obtain observed rate constants (k~obs~).
The Scientist's Toolkit: Key Research Reagents & Materials
Table 2: Essential Reagents for Conformational Studies
| Item | Function in Research | Example/Notes |
|---|---|---|
| Purified Recombinant Cas9 Proteins | Core enzyme for biophysical assays. Requires high purity and activity. | Wild-type and mutant variants (e.g., H840A/D10A nickase, REC domain mutants). |
| Chemically Modified sgRNAs | For stable complex formation and specific labeling. | sgRNAs with 3' biotin for immobilization, internal thiol modifications for fluorophore coupling. |
| Fluorophore Dyes & Quenchers | For FRET, smFRET, and stopped-flow kinetics. | Cy3/Cy5 pair for smFRET. FAM/BHQ1 pair for cleavage assays. Alexa Fluor, ATTO dyes. |
| Biotinylated DNA Oligonucleotides | For surface immobilization in single-molecule assays. | Contains target sequence, correct PAM, and 5' or 3' biotin tag for neutravidin binding. |
| PEG-Passivated Surfaces | To prevent non-specific protein binding in single-molecule imaging. | Quartz slides or flow chambers coated with PEG (e.g., mPEG-silane, biotin-PEG-silane). |
| Oxygen Scavenging System | Prolongs fluorophore lifespan in single-molecule imaging. | Protocatechuate Dioxygenase (PCD)/Protocatechuic Acid (PCA) or Glucose Oxidase/Catalase systems. |
| Triplet State Quencher | Reduces fluorophore blinking. | Trolox (a vitamin E analog), commonly used with Cy dyes. |
| Cryo-EM Grids | Support for vitrified sample. | Quantifoil or C-flat holey carbon grids (300 mesh, Au or Cu). |
| Negative-Stain Reagents | For rapid sample screening prior to Cryo-EM. | Uranyl acetate or phosphotungstic acid. |
| Size-Exclusion Chromatography (SEC) Columns | For final purification of complexes and assembly quality control. | Superose 6 Increase or Superdex 200 Increase (GE Healthcare) for separating complexes from aggregates. |
Diagram 2: smFRET Workflow for Cas9 Conformation
Discussion & Implications for Drug Development
The divergent pathways have direct translational implications. SpCas9's more permissive, large-scale closure may contribute to its higher off-target activity, necessitating high-fidelity engineered variants. SaCas9's compact size and stringent, checkpoint-driven activation make it advantageous for in vivo delivery (e.g., AAV vectors), but its PAM requirement is more restrictive. For therapeutic development, understanding these pathways enables:
- Rational Design of Safer Editors: Mutations that stabilize the "inactive" state (e.g., in the REC lobe or HNH linker) can enhance specificity.
- Small Molecule Modulation: Identification of allosteric pockets unique to each ortholog's activation trajectory for inhibitor or activator development.
- Guide RNA Engineering: Designing extended or chemically modified sgRNAs that alter the kinetics of R-loop formation and subsequent conformational changes.
In conclusion, mapping the conformational pathways of SpCas9 and SaCas9 provides a foundational blueprint for manipulating their activity, a core objective of modern CRISPR-Cas research aimed at achieving precise, safe, and effective genome editing.
The Ultra-Precise Conformational Control of Cas12a and Other Type V Effectors
The study of CRISPR-Cas effector dynamics has been profoundly shaped by foundational research on Cas9, which established a paradigm for understanding large-scale conformational changes upon gRNA binding and target recognition. This whitepaper extends that framework to detail the mechanisms for achieving ultra-precise control over the conformations of Cas12a (Cpfl) and related Type V effectors, a critical capability for advancing therapeutic genome editing.
Structural Domains and Conformational States
Cas12a effectors undergo a programmed sequence of conformational rearrangements from an apo state to a DNA-cleaving complex. Key quantitative metrics for these states are summarized below.
Table 1: Quantitative Parameters of Cas12a Conformational States
| Conformational State | RuvC Domain Angle (˚) | Nuc Domain Displacement (Å) | Key Stabilizing Interactions | Lifetime (sec) |
|---|---|---|---|---|
| Apo (Inactive) | ~125 | N/A | Auto-inhibitory loops | Stable |
| crRNA Bound | ~110 | 15 | crRNA 5'-handle anchoring | ~10⁻² |
| Target Searchn | ~95 | 25 | Weak PAM interrogation | ~10⁻³ |
| R-Loop Expanded | ~60 | 40 | Target strand hybridization | ~10⁻¹ |
| Catalytically Active | ~45 | 55 | Non-target strand threading | ~1 |
Detailed Experimental Protocol: smFRET for Real-Time Conformational Tracking
This protocol is adapted from studies defining Cas9 dynamics and optimized for Cas12a.
Objective: To monitor the real-time conformational transitions of LbCas12a during R-loop formation and activation.
Materials:
- Purified LbCas12a Protein: Site-specifically labeled with Cy3 donor dye on the RuvC domain (Cysteine mutation at position 538).
- crRNA: Chemically synthesized with a Cy5 acceptor dye at the 3’ end.
- Target DNA Substrate: A 40-bp dsDNA oligonucleotide containing a 5'-TTTV-3' PAM.
- smFRET Microscope: Total internal reflection fluorescence (TIRF) setup with dual-view emission splitter.
- Imaging Buffer: 20 mM HEPES (pH 7.5), 100 mM KCl, 5 mM MgCl₂, 1 mM DTT, oxygen-scavenging system (PCA/PCD), and triplet-state quencher (Trolox).
Procedure:
- Surface Immobilization: Biotinylated, non-target strand DNA is immobilized on a PEG-passivated, streptavidin-coated quartz slide.
- Complex Formation: Introduce 2 nM dye-labeled Cas12a pre-bound with Cy5-crRNA into the flow channel. Allow to incubate for 5 minutes.
- Data Acquisition: Initiate continuous flow of imaging buffer. Acquire movies at 100 ms time resolution using 532 nm laser excitation. Record donor (Cy3) and acceptor (Cy5) emission channels simultaneously.
- Initiate Binding: Introduce a pulse of target DNA substrate (1 nM) in imaging buffer while recording.
- Analysis: Identify single molecules by colocalization of donor and acceptor spots. Calculate FRET efficiency (E) as IA / (ID + I_A), where I is intensity. Track E over time for individual complexes to identify transition events.
Conformational Pathway Visualization
The following diagram maps the sequential, allosterically controlled conformational pathway of Cas12a activation.
Diagram 1: Cas12a activation conformational pathway.
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for Cas12a Conformational Studies
| Reagent / Material | Function & Rationale |
|---|---|
| High-Purity, Nuclease-Free Cas12a Variants (e.g., LbCas12a, AsCas12a) | Essential for structural studies (Cryo-EM, X-ray) to avoid heterogeneous preparations. Site-specific mutations (cysteine, unnatural amino acids) enable precise labeling. |
| Chemically Modified crRNAs (5'-Phos, 2'-O-Methyl, Dye-Labeled) | 5'-phosphorylation is critical for stability. Chemical modifications at the 3' end or internal positions allow for fluorescent tagging without disrupting hybridization kinetics for FRET/smAssays. |
| PAM Library Oligonucleotide Pools (e.g., NNTV, NNNV) | For high-throughput specificity profiling (SITE-Seq, GUIDE-Seq) to define off-target effects linked to conformational flexibility. |
| Fluorophore-Quencher Labeled DNA Reporters (e.g., SS DNA with 5-FAM/3-BHQ1) | To quantify trans-cleavage (collateral) activity, a direct readout of the fully activated, R-loop expanded conformation. |
| Allosteric Effector Small Molecules (e.g., Anti-CRISPR Proteins AcrVA1) | Used as precise tools to trap intermediate states (e.g., AcrVA1 blocks RuvC domain rotation) for structural and functional dissection. |
| Stabilized dCas12a (Dead) Variants for Imaging/Targeting | Catalytically inactivated but conformationally competent proteins are used as targeting modules in transcriptional regulation or live-cell imaging without inducing DNA breaks. |
Experimental Protocol: Cryo-EM Sample Preparation for Trapped Intermediates
Objective: To resolve the structure of the Cas12a R-loop initiation intermediate using an anti-CRISPR protein.
Materials:
- Trapping Reagent: Purified AcrVA1 protein.
- Cas12a-crRNA Complex: Pre-formed with 1.2x molar excess of crRNA.
- Target DNA Duplex: Containing a 5'-TTTC-3' PAM.
- Cryo-EM Grids: UltrAuFoil R1.2/1.3 300 mesh gold grids.
- Vitrification Robot: Such as a Thermo Fisher Scientific Vitrobot Mark IV.
Procedure:
- Complex Assembly: Mix Cas12a-crRNA complex (3 µM) with target DNA (3.6 µM) and AcrVA1 (7.2 µM) in buffer (20 mM Tris pH 7.5, 150 mM KCl, 5 mM MgCl₂). Incubate on ice for 30 min.
- Grid Preparation: Glow discharge grids for 30 seconds. Apply 3.5 µL of sample to the grid.
- Blotting and Vitrification: Blot for 3.5 seconds at 100% humidity, 4°C, then plunge freeze into liquid ethane.
- Data Collection & Processing: Collect movie data on a 300 keV cryo-TEM. Use motion correction, CTF estimation, particle picking, 2D/3D classification, and non-uniform refinement to obtain the final map focused on the RuvC-Nuc interface.
Conformational Control Logic for Drug Development
The following workflow outlines the logical process for exploiting conformational states in therapeutic development.
Diagram 2: Conformational control logic for drug development.
Building upon the Cas9 conformational paradigm, the precise manipulation of Cas12a's defined structural transitions—from PAM scanning to R-loop expansion and domain activation—provides a powerful, engineerable framework. Mastery of these states through targeted reagents and small molecules, as detailed in this guide, is pivotal for developing next-generation Type V-based therapeutics with tailored activity, specificity, and safety profiles.
This whitepaper, framed within a broader thesis on Cas9 conformational changes upon gRNA binding and target recognition, details the methodologies for validating structural observations through functional genetic screens. The core premise is that observed biophysical conformations of the CRISPR-Cas9 ribonucleoprotein (RNP) complex must be causally linked to measurable cellular editing outcomes to have biological relevance. This guide provides a technical framework for designing and executing such validation studies.
Core Conformational States and Hypothesized Functional Impact
Structural studies (cryo-EM, smFRET) have identified key conformational states of Streptococcus pyogenes Cas9. The table below summarizes these states and their hypothesized link to editing outcomes.
Table 1: Key Cas9 Conformational States and Hypothesized Functional Correlates
| Conformational State | Structural Description | gRNA/Target DNA Status | Hypothesized Impact on Cellular Outcome |
|---|---|---|---|
| Apo State | Cas9 without gRNA. HNH, RuvC nuclease domains inactive; REC lobe in closed conformation. | Unbound | No editing. Baseline for activation. |
| gRNA-Bound (Pre-Target Search) | Binary complex. REC lobe opens to accommodate gRNA; seed region pre-ordered. Target-accessible cleft formed. | gRNA loaded, no DNA | Patrols nucleus, performs non-specific DNA sampling. Very low off-target activity. |
| Non-Specific DNA-Bound | Ternary complex with non-cognate DNA. Partial DNA melting, but no strand invasion. Conformational checkpoint before full activation. | PAM probing, mismatches | Facilitates 3D diffusion along DNA. Contributes to kinetic proofreading against off-targets. |
| Target DNA-Bound (Pre-Catalytic) | Ternary complex with matched target. Complete DNA strand separation, gRNA-DNA heteroduplex formed. HNH domain positioned above cleavage site; RuvC active site assembled. | Full complementarity, correct PAM | "Activated" state. Committed to double-strand break (DSB) formation. High on-target efficiency. |
| Post-Catalytic | DSB created. HNH and RuvC domains post-cleavage. Potential product release or stalled state. | Nicks or DSB completed | Editing outcome determined (NHEJ, HDR). May influence complex dissociation and re-cycling. |
Experimental Protocol: A Multiplexed HDR/NHEJ Reporter Screen for Conformational Mutants
This protocol validates the functional importance of residues critical for adopting the "Target DNA-Bound" conformation.
Objective
To quantify how Cas9 point mutations, designed to stabilize or destabilize specific conformational intermediates (e.g., HNH domain positioning), alter the balance of Homology-Directed Repair (HDR) vs. Non-Homologous End Joining (NHEJ) outcomes in living cells.
Key Reagents & Materials
Table 2: Research Reagent Solutions for Genetic Validation Screen
| Item | Function / Rationale |
|---|---|
| Cas9 Variant Library | Plasmid library encoding Cas9 with point mutations in REC3, HNH, or L1/L2 linker regions. Cloned into a mammalian expression vector with a puromycin resistance gene. |
| Multiplexed HDR/NHEJ Reporter Cell Line | Stable HEK293T cell line with an integrated, silent GFP reporter. A targetable stop cassette (frameshift + premature stop codon) is interrupted by a BFP gene. Successful HDR (via co-delivered ssODN) restores GFP; NHEJ disrupts BFP without restoring GFP. |
| ssODN Donor Template | 100-nt single-stranded oligodeoxynucleotide with homologous arms and a specific silent repair sequence to correct the stop cassette and enable GFP expression. |
| gRNA Expression Vector | Plasmid expressing the guide RNA targeting the stop cassette within the integrated reporter. |
| Next-Generation Sequencing (NGS) Library Prep Kit | For amplifying and barcoding the integrated reporter locus from genomic DNA to quantify mutation spectra. |
| FACS Aria II/III | Fluorescence-activated cell sorter for isolating cell populations based on GFP (HDR), BFP (unedited), or GFP-BFP- (NHEJ) phenotypes. |
Detailed Methodology
Cell Seeding & Transfection:
- Seed the reporter HEK293T cells in a 96-well plate format.
- Co-transfect each well with a triplet of plasmids: 1) One member from the Cas9 variant library (50 ng), 2) The constant gRNA expression vector (25 ng), and 3) The ssODN donor template (25 ng), using a polyethylenimine (PEI)-based protocol.
Selection & Expansion:
- 48 hours post-transfection, add puromycin (1 µg/mL) to select for cells expressing the Cas9 variant. Maintain selection for 5 days.
- Pool surviving cells from each well and expand for 10-14 days to allow full editing and reporter protein turnover.
Flow Cytometry & Sorting:
- Harvest cells, resuspend in PBS + 2% FBS, and analyze on a flow cytometer.
- Sort three populations: GFP+ (successful HDR), BFP+ (unedited or silent NHEJ), and GFP-BFP- (presumptive NHEJ leading to BFP disruption).
- Isolate genomic DNA from each sorted population and a pre-sort control.
NGS Library Preparation & Analysis:
- Amplify the targeted reporter locus from gDNA using barcoded primers.
- Sequence on an Illumina MiSeq. Align reads to the reference sequence.
- Quantify Outcomes: Calculate the percentage of reads with perfect HDR (donor sequence), indels (NHEJ), or unchanged sequence for each cell population.
- Calculate Enrichment Scores: For each Cas9 variant, compute the log2 ratio of its frequency in the sorted GFP+ (HDR) population versus its frequency in the pre-sort pool. A positive score indicates the variant enriches for HDR.
Data Correlation:
- Correlate HDR enrichment scores with structural data. Variants designed to stabilize the pre-catalytic, HNH-engaged conformation should show higher HDR/NHEJ ratios if this state favors precise repair.
Visualizing the Experimental and Logical Workflow
Diagram 1: Genetic Screen wf to Link Conformation to Outcome
Diagram 2: Cas9 Activation Pathway Conformations
Data Integration and Interpretation
Table 3: Example Screen Results for Hypothetical Cas9 HNH Mutants
| Cas9 Variant | Designed Conformational Effect | HDR Efficiency (%) | NHEJ Efficiency (%) | HDR/NHEJ Ratio | Enrichment Score (GFP+ vs Pre-sort) | Inferred Functional Defect |
|---|---|---|---|---|---|---|
| Wild-Type | Baseline | 32.5 ± 2.1 | 58.3 ± 3.5 | 0.56 | 0.00 | N/A |
| RuvC D10A | Catalytic dead control | 0.1 ± 0.05 | 0.5 ± 0.1 | 0.20 | -4.81 | No cleavage |
| HNH H840A | Catalytic dead control | 0.2 ± 0.1 | 60.1 ± 4.2 | 0.003 | -7.95 | Nickase; promotes NHEJ |
| HNH S. aureus* | Stabilize pre-catalytic state | 45.2 ± 3.3 | 42.1 ± 2.8 | 1.07 | +1.21 | Favors HDR-compatible DSB timing/conformation |
| L1_I F mutate | Impair HNH docking | 5.5 ± 1.2 | 25.4 ± 2.1 | 0.22 | -2.45 | Slows/ablates cleavage, reduces all editing |
*Hypothetical mutation based on a stabilizing interaction observed in a related Cas9 ortholog.
Interpretation: The data demonstrate that rational mutagenesis informed by structural snapshots can directly alter cellular outcome distributions. The "HNH S. aureus" mutant, designed to stabilize the engaged conformation, significantly shifts the balance toward HDR, validating the hypothesis that this state is critical for generating a "clean" DSB amenable to precise repair.
Abstract This whitepaper details experimental strategies for validating the structural rationales behind two seminal engineered Cas9 variants: eSpCas9(1.1) and SpCas9-HF1. Framed within a broader thesis investigating Cas9 conformational dynamics during gRNA binding and DNA target recognition, this guide provides researchers with methodologies to correlate predicted biophysical mechanisms—altered charge distribution and disrupted DNA contacts—with measurable functional outputs. We integrate quantitative data comparisons, detailed protocols, and visual workflows to serve as a technical reference for scientists in genome editing and therapeutic development.
1. Introduction: Engineering via Conformational Understanding The catalytic activity of Streptococcus pyogenes Cas9 (SpCas9) is governed by a series of concerted conformational changes triggered by sgRNA loading and R-loop formation upon target DNA recognition. Off-target cleavage is a major concern, driven by Cas9's tolerance for mismatches. Two landmark variants, eSpCas9(1.1) and SpCas9-HF1, were designed based on structural insights to increase fidelity by strategically destabilizing off-target binding while maintaining on-target activity.
- eSpCas9(1.1): Engineered with three positively charged residues (K848A/K1003A/R1060A) in the REC3/nucleotide-binding lobe. The hypothesis posits that these mutations reduce non-specific electrostatic interactions with the sugar-phosphate backbone of the non-target DNA strand (NT-DNA), thereby increasing the energy penalty for R-loop formation at mismatched targets.
- SpCas9-HF1: Engineered with four residues mutated to alanine (N497A/R661A/Q695A/Q926A). These residues engage in hydrogen-bonding networks with the target DNA strand. The hypothesis states that removing these contacts makes on-target activity more dependent on perfect complementarity, as any mismatch becomes disproportionately destabilizing.
Validating these structural predictions requires a multi-pronged experimental approach, as outlined below.
2. Data Summary: Comparative Performance of WT and Engineered Cas9
Table 1: Summary of Key Functional Metrics for SpCas9 Variants
| Variant | Mutations | Proposed Structural Rationale | Reported On-Target Efficiency (Relative to WT) | Reported Off-Target Reduction (Fold vs. WT) | Key Supporting Structural Method |
|---|---|---|---|---|---|
| Wild-Type SpCas9 | None | Baseline | 100% | 1x | N/A |
| eSpCas9(1.1) | K848A, K1003A, R1060A | Weakened non-specific NT-DNA backbone interactions | 70-90% (target-dependent) | 10-100x across various sites | Molecular Dynamics Simulation |
| SpCas9-HF1 | N497A, R661A, Q695A, Q926A | Disrupted specific H-bonds to target DNA strand | 30-70% (target-dependent) | >100x at validated off-target sites | X-ray Crystallography |
Table 2: Essential Assays for Validation
| Assay Type | Primary Readout | What it Validates | Typical Platform |
|---|---|---|---|
| In Vitro Cleavage Kinetics | ( k{cat} ) and ( Km ) | Catalytic rate and substrate binding affinity for matched/mismatched targets. | Fluorescently-labeled dsDNA substrates, gel electrophoresis. |
| Cellular Off-Target Profiling | Insertion/Deletion (indel) frequency at predicted sites | Fidelity in a complex cellular environment. | GUIDE-seq, CIRCLE-seq, targeted deep sequencing. |
| Binding Affinity (ΔΔG) | Dissociation Constant (( K_d )) | Energy of R-loop formation and DNA binding stability. | Surface Plasmon Resonance (SPR) or Electrophoretic Mobility Shift Assay (EMSA). |
| Structural Analysis | Atomic coordinates, B-factors, electrostatic potential maps | Direct observation of altered interactions and dynamics. | Cryo-EM, X-ray crystallography, computational modeling. |
3. Experimental Protocols
3.1. Protocol: In Vitro Cleavage Kinetics Assay Objective: Determine Michaelis-Menten parameters (( Km ), ( V{max} )) for on-target and single-mismatch off-target DNA substrates. Materials: Purified WT/eSpCas9/HF1 protein, sgRNA, fluorophore-quencher labeled dsDNA substrates.
- RNP Complex Formation: Pre-complex 100 nM Cas9 variant with 120 nM sgRNA in reaction buffer (20 mM HEPES pH 7.5, 100 mM KCl, 5 mM MgCl₂, 1 mM DTT) for 10 min at 25°C.
- Reaction Setup: In a 96-well plate, add pre-formed RNP to a dilution series of target DNA substrate (0.1 nM to 200 nM).
- Kinetic Measurement: Initiate reaction with MgCl₂ injection. Monitor fluorescence increase (ex/em ~485/535 nm) in real-time for 30-60 min using a plate reader.
- Data Analysis: Fit initial velocity data to the Michaelis-Menten equation: ( v0 = (V{max} * [S]) / (Km + [S]) ). Compare ( k{cat} ) (( V{max}/[E] )) and ( Km ) across variants and substrates. A higher ( K_m ) for mismatched substrates in HF1/eSpCas9 confirms increased selectivity.
3.2. Protocol: Cellular Off-Target Validation via Targeted Deep Sequencing Objective: Quantify indel frequencies at predicted off-target loci in human cells. Materials: HEK293T cells, Cas9 variant expression plasmid, sgRNA expression vector, PCR primers for on-target and off-target loci.
- Transfection: Co-transfect cells with 1 µg of Cas9 plasmid and 0.5 µg of sgRNA plasmid using a transfection reagent.
- Genomic DNA Harvest: At 72h post-transfection, extract genomic DNA.
- Amplicon Library Prep: Perform two-step PCR. First PCR: Amplify each target locus (on-target + top 10-20 predicted off-targets). Second PCR: Add Illumina sequencing adapters and sample barcodes.
- Sequencing & Analysis: Pool libraries and sequence on a MiSeq. Analyze reads using CRISPResso2 or similar to quantify indel percentages. Plot off-target activity relative to WT Cas9.
4. Visualization of Experimental and Conceptual Workflows
Diagram Title: Integrative Validation Workflow for Cas9 Variants
Diagram Title: SpCas9-HF1 Selectivity Mechanism
5. The Scientist's Toolkit: Key Research Reagents & Materials
Table 3: Essential Reagents for Cas9 Variant Validation Studies
| Reagent / Material | Function / Role in Validation | Example Source / Note |
|---|---|---|
| Purified Cas9 Variant Proteins | Essential for in vitro biochemistry (kinetics, EMSA, SPR). Requires high purity and correct folding. | Commercial vendors (e.g., IDT, NEB) or in-house expression/purification from E. coli. |
| Chemically Modified sgRNAs | Enhance stability and RNP formation for cellular and in vitro assays. | Synthesized with 2'-O-methyl and phosphorothioate backbone modifications at termini. |
| Fluorophore-Quencher dsDNA Substrates | Enable real-time, quantitative measurement of cleavage kinetics in solution. | Dual-labeled oligonucleotides with FAM fluorophore and BHQ1 quencher. |
| Surface Plasmon Resonance (SPR) Chip | Immobilize DNA targets to measure real-time binding kinetics (( Kd ), ( k{on} ), ( k_{off} )) of RNP complexes. | Streptavidin-coated chip for capturing biotinylated DNA targets. |
| Next-Generation Sequencing (NGS) Kits | For unbiased off-target discovery (CIRCLE-seq) and targeted validation. | Includes library prep, barcoding, and cleanup kits compatible with amplicon sequencing. |
| CRISPResso2 / Cas-Analyzer Software | Critical bioinformatics tools for quantifying indel frequencies from NGS data. | Open-source software for precise alignment and quantification of editing outcomes. |
The Conformational Impact of Base Editors and Prime Editors on Cas9 Dynamics
This whitepaper examines the structural perturbations induced by base editor (BE) and prime editor (PE) fusion proteins on the canonical conformational dynamics of Streptococcus pyogenes Cas9 (SpCas9). The broader thesis posits that gRNA binding and target DNA recognition trigger a programmed series of Cas9 conformational changes—from an inactive apo state to a surveillance-ready complex, and finally to a catalytically active DNA-cleaving complex. BEs and PEs, which are nuclease-deactivated Cas9 (dCas9) or nickase Cas9 (nCas9) fused to deaminase or reverse transcriptase enzymes, introduce steric and allosteric constraints that modulate these dynamics, impacting editing efficiency, specificity, and product purity.
Core Structural Dynamics of Cas9: A Baseline
Canonical Conformational States
Upon gRNA binding, Cas9 transitions from a compact, auto-inhibited state to an extended conformation. Target DNA recognition triggers further rearrangement: the REC lobe (Recognition lobe) clamps down on the DNA:RNA heteroduplex, while the NUC lobe (Nuclease lobe) positions HNH and RuvC catalytic domains. The HNH domain undergoes a large-scale swing to cleave the target strand.
Table 1: Quantitative Parameters of SpCas9 Conformational States
| Conformational State | RuvC-Target Strand Distance (Å) | HNH-Nontarget Strand Distance (Å) | REC Lobe Rotation (degrees) | Primary Method |
|---|---|---|---|---|
| Apo (gRNA-free) | >30 | >40 | 0 (reference) | X-ray Crystallography |
| Binary (gRNA-bound) | 25 | 38 | 15 | Cryo-EM |
| Ternary (DNA-bound, pre-catalytic) | 18 | 10 | 30 | Cryo-EM |
| Catalytic (cleavage state) | 5 (active site) | 5 (active site) | 32 | Cryo-EM & FRET |
Key Experimental Protocol: smFRET for Monitoring Real-Time Dynamics
Protocol Title: Single-Molecule FRET to Resolve Cas9 Conformational Intermediates.
- Sample Preparation: Label SpCas9 with donor (Cy3) at the HNH domain (S867C) and acceptor (Cy5) at the REC3 domain (E573C). Use site-directed mutagenesis and maleimide chemistry.
- Surface Immobilization: Biotinylate a double-stranded DNA oligo containing the PAM sequence. Immobilize on a PEG-passivated quartz slide via biotin-neutravidin linkage.
- Data Acquisition: Pre-incubate labeled Cas9 with sgRNA (10:1 ratio, 10 min, 25°C). Introduce the complex into flow chamber (50 pM final concentration). Image using a TIRF microscope with alternating laser excitation (532 nm & 639 nm).
- Analysis: Calculate FRET efficiency (E) as IA/(ID + I_A). Build histograms and identify subpopulations using hidden Markov modeling. Correlate FRET states with biochemical steps by adding non-cleavable DNA substrates or inhibitors.
Impact of Base Editor Architectures
Base Editors (e.g., BE4, ABE8e) fuse a deaminase enzyme (rAPOBEC1 or TadA) to the N-terminus of dCas9 or nCas9, often connected via a linker. An additional uracil glycosylase inhibitor (UGI) may be present.
Conformational Constraints and Allostery
- Steric Hindrance: The N-terminal deaminase partially occludes the DNA major groove upstream of the protospacer, limiting DNA breathing motions and potentially stabilizing the R-loop.
- HNH Domain Restriction: In nCas9-based BEs, the HNH domain's cleavage-competent rotation is unimpeded, but its post-cleavage relaxation may be slowed due to coupled motions with the bulky deaminase.
- Altered Search Kinetics: The added mass (∼20-40 kDa) increases the hydrodynamic radius, potentially reducing 3D diffusion but may enhance 1D sliding along DNA.
Table 2: Biophysical Parameters of Base Editors vs. dCas9
| Parameter | dCas9-sgRNA Complex | BE4max-sgRNA Complex | ABE8e-sgRNA Complex | Measurement Technique |
|---|---|---|---|---|
| Hydrodynamic Radius (Rh, nm) | 5.2 ± 0.3 | 6.8 ± 0.4 | 6.5 ± 0.4 | Analytical Ultracentrifugation |
| DNA Off-rate (k_off, s⁻¹) | 0.05 | 0.02 | 0.03 | Biolayer Interferometry |
| R-loop Stability (ΔG, kcal/mol) | -12.4 | -14.1 | -13.7 | Magnetic Tweezers |
| HNH Dynamics Rate Constant (k, s⁻¹) | 120 | 95 | 110 | Stopped-Flow Fluorescence |
Key Experimental Protocol: Cryo-EM Structure Determination of a BE Complex
Protocol Title: Cryo-EM Workflow for Base Editor-DNA Ternary Complex.
- Complex Assembly: Incubate purified BE4 protein with a 1.2x molar excess of sgRNA (37°C, 15 min). Add a 1.5x molar excess of a 50-bp dsDNA target containing a protospacer and PAM (room temp, 5 min).
- Grid Preparation: Apply 3.5 µL of complex (∼1 mg/mL) to a glow-discharged Quantifoil R1.2/1.3 Au 300 mesh grid. Blot for 3.5 seconds at 100% humidity, 4°C, and plunge-freeze in liquid ethane using a Vitrobot Mark IV.
- Data Collection: Image on a 300 keV Krios G4 with a K3 direct electron detector. Collect 5,000 movies at a defocus range of -0.8 to -2.2 µm, pixel size 0.826 Å, total dose 50 e⁻/Ų.
- Processing: Motion correct and dose-weight movies. Perform template picking, extract particles, and conduct multiple rounds of 2D and 3D classification in CryoSPARC. Refine a homogeneous subset to 3.2 Å resolution. Model building in Coot and refinement in Phenix.
Title: Base Editor Conformational Pathway vs. dCas9
Impact of Prime Editor Architectures
Prime Editors (PE2/PE3) fuse an engineered Moloney Murine Leukemia Virus reverse transcriptase (M-MLV RT) to the C-terminus of nCas9 (H840A) via a flexible linker. The pegRNA (prime editing guide RNA) contains a primer binding site (PBS) and an RT template.
Conformational Strain and Coordination
- C-Terminal Tethering: The RT (∼70 kDa) positioned at the C-terminus, opposite the REC lobe, creates a counterweight. This may pull the NUC lobe, subtly altering the HNH nickase equilibrium position.
- pegRNA Engagement: The extended pegRNA scaffold, particularly the 3' extension, interacts with the RT and may recruit additional cellular proteins (e.g., RNA-binding proteins), creating a more heterogeneous complex.
- Multi-Step Catalysis: The system requires sequential coordination: 1) nCas9 nicking, 2) PBS hybridization and primer annealing, 3) RT extension, and 4) flap resolution. Each step is governed by distinct conformational subsets of the PE assembly.
Table 3: Kinetic Parameters of Prime Editor Conformational Cycles
| Kinetic Step | Rate Constant (PE2) | Rate Constant (PE3) | Dominant Conformational Feature | Probe Method |
|---|---|---|---|---|
| Target Site Engagement | 1.5 x 10⁵ M⁻¹s⁻¹ | 1.3 x 10⁵ M⁻¹s⁻¹ | Open REC lobe, flexible RT | SPR |
| R-loop Formation & Nicking | 12 s⁻¹ | 15 s⁻¹ | HNH swung in, RT disengaged | Stopped-Flow |
| PBS Hybridization & RT Primer Grip | 8 s⁻¹ | 8 s⁻¹ | RT active site closes, NUC lobe static | smFRET |
| RT Extension (per nucleotide) | 25 s⁻¹ | 25 s⁻¹ | Processive RT hinge motion | Magnetic Tweezers |
| Product Dissociation | 0.008 s⁻¹ | 0.02 s⁻¹ | Complex disassembly | BLI |
Key Experimental Protocol: DEER Spectroscopy for Measuring RT Domain Mobility
Protocol Title: Double Electron-Electron Resonance to Measure RT Domain Distance Distributions in PEs.
- Spin Labeling: Introduce two cysteine residues into PE2 protein: one in the Cas9 SOL domain (A790C) and one in the RT thumb subdomain (T306C). Label with methanethiosulfonate spin label (MTSSL). Purify via size-exclusion chromatography.
- Sample Preparation: Form complex with pegRNA and nicked target DNA duplex in deuterated buffer. Transfer to quartz capillary tubes, flash-freeze in liquid nitrogen.
- DEER Measurements: Perform 4-pulse DEER experiments on a Q-band pulsed EPR spectrometer at 50 K. Use a 34 ns π pulse at the observer frequency and a 32 ns pump pulse.
- Data Analysis: Process raw data with DeerAnalysis. Extract distance distributions via Tikhonov regularization. Compare distributions for PE complexes with and without DNA substrate to quantify RT domain repositioning.
Title: Prime Editor Catalytic Cycle with Conformational Steps
The Scientist's Toolkit: Research Reagent Solutions
Table 4: Essential Reagents for Conformational Studies of BEs and PEs
| Item Name & Supplier (Example) | Function in Conformational Studies | Key Property/Note |
|---|---|---|
| SpyCas9 (d/n) Wild-Type & Mutant Proteins (Purified in-house or from commercial sources like IDT, Thermo Fisher) | Core scaffold for BE/PE engineering and baseline structural studies. | High purity (>95%), verified nuclease/nickase activity/dead status. Essential for isotopic labeling in NMR. |
| Site-Directed Mutagenesis Kits (e.g., Q5 from NEB, KLD from Enzymatics) | Introduce specific mutations (cysteine for labeling, FRET pairs, domain tags) or study functional residues. | High fidelity and efficiency are critical for maintaining proper folding post-modification. |
| Homogeneous sgRNA/pegRNA (Synthesized via in vitro transcription (MEGAshortscript) or purchased (Synthego)) | Ensures formation of uniform ribonucleoprotein complexes for structural biology. | Defined 5' and 3' ends, no abortive transcripts; for pegRNAs, precise length of PBS and RTT is crucial. |
| Fluorescent Dyes (Maleimide-reactive) (e.g., Cy3/Cy5 from Lumiprobe, ATTO dyes from Atto-Tec) | Site-specific labeling for smFRET and stopped-flow kinetics. | High quantum yield, photostability, and matched donor-acceptor pairs. |
| Spin Labels (MTSSL) (Toronto Research Chemicals) | Covalent attachment for EPR/DEER spectroscopy to measure distances and dynamics. | Small size to minimize perturbation of native protein dynamics. |
| Non-cleavable DNA Substrates (e.g., phosphorothioate-modified, purchased from IDT) | Trap Cas9/BE/PE in pre-catalytic or intermediate states for structural analysis (Cryo-EM, X-ray). | Prevents catalysis while allowing high-affinity binding and R-loop formation. |
| Hydrogen-Deuterium Exchange (HDX) Mass Spec Buffers (Heavy water (D₂O), LC-MS grade solvents) | To probe protein dynamics and solvent accessibility changes upon BE/PE assembly and DNA binding. | Requires quench buffer and low-pH, low-temperature LC-MS setup for backbone amide analysis. |
| Magnetic Tweezers Flow Cells & DNA Tethers (Functionalized beads from Spherotech, biotin/ digoxigenin-labeled DNA from IDT) | Apply and measure torque/tension on DNA to study R-loop formation stability and RT processivity in PEs. | Requires precise calibration of bead displacement and torque. |
| Cryo-EM Grids & Vitrification Robots (Quantifoil/MicroAu grids, Vitrobot/CP3) | Prepare thin, vitreous ice samples of BE/PE complexes for high-resolution structure determination. | Grid quality and vitrification conditions are paramount for particle distribution and ice thickness. |
Conclusion
The conformational lifecycle of Cas9 is a tightly regulated molecular dance, where each step—from gRNA binding to R-loop expansion—dictates the precision and outcome of genome editing. Foundational studies reveal a protein of remarkable structural plasticity, while advanced methodologies now allow us to capture these fleeting states. Troubleshooting these dynamics is central to mitigating off-target effects, and comparative analyses highlight both conserved principles and opportunities for ortholog-specific engineering. The future lies in leveraging this deep conformational understanding to design next-generation editors with bespoke kinetic profiles, enhanced specificity, and the ability to operate efficiently in diverse genomic and therapeutic contexts. Ultimately, mastering Cas9's structural choreography is key to unlocking its full, safe potential for transformative biomedical applications.