Unlocking the Blueprints of Life: The Revolution in KEGG Pathway Exploration
From static maps to living landscapes of cellular processes
Article Navigation

Introduction: From Static Maps to Living Landscapes
For decades, biologists navigated the complexities of cellular processes using pathway diagrams akin to static paper maps. The Kyoto Encyclopedia of Genes and Genomes (KEGG), established in 1995, became the gold standard for visualizing metabolic and signaling pathways 6 . Yet, these semi-static diagrams had a critical limitation: they couldn't be easily edited, personalized, or interactively explored. Enter the era of dynamic KEGG pathway tools—where researchers now manipulate these molecular blueprints in real time, transforming how we model diseases, engineer metabolism, and decode big data.
The Static Bottleneck: Why KEGG Needed a Digital Upgrade
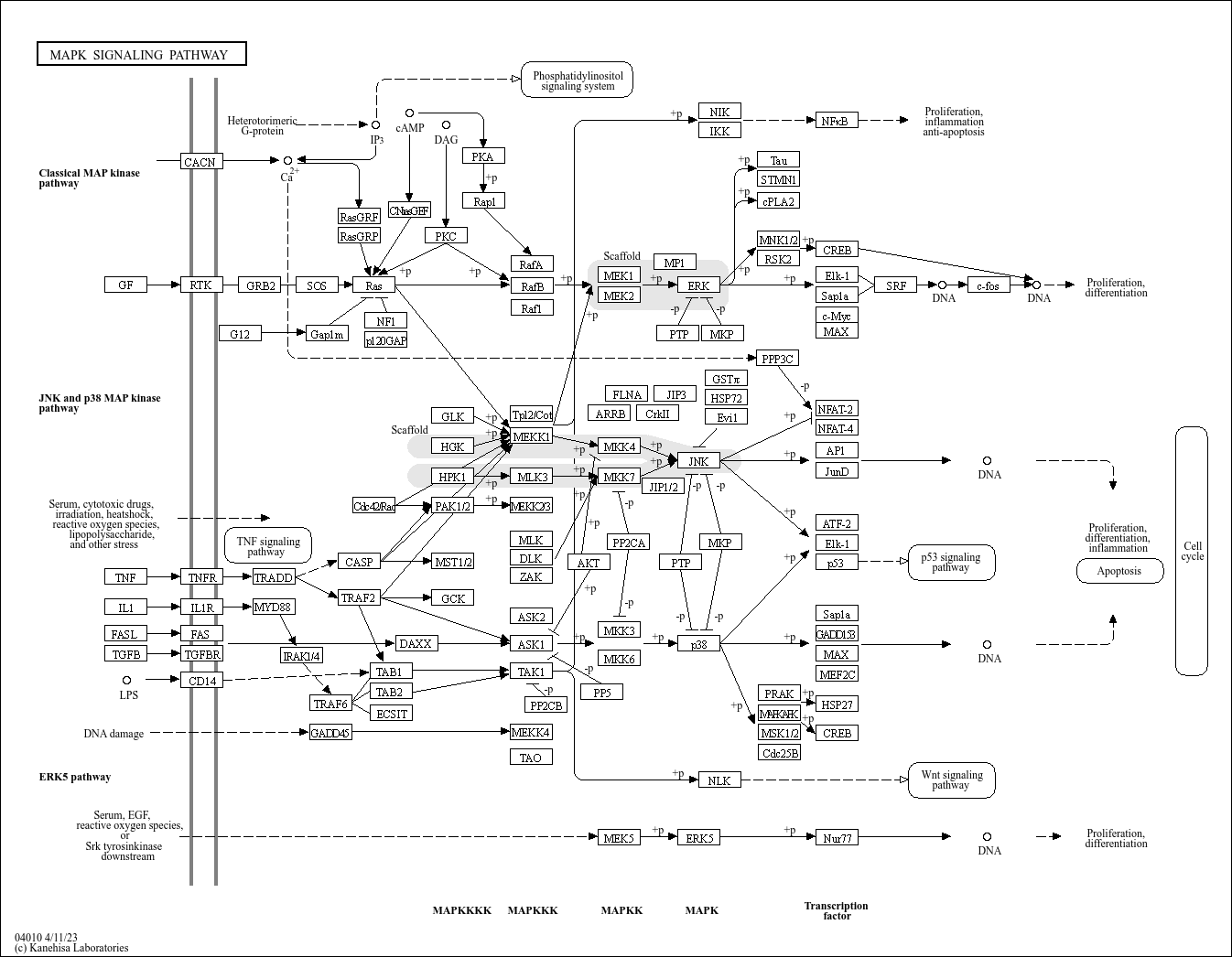
KEGG's original pathway maps are masterpieces of biological curation. Manually drawn using tools like KegSketch, they depict:
- Boxes representing enzymes or genes (linked to KO identifiers)
- Circles denoting metabolites (like lipids or sugars)
- Lines indicating reactions or interactions 3 7
However, these maps are generic. When studying specific organisms (e.g., human cancer cells) or experimental conditions, researchers faced a dilemma: they couldn't easily highlight relevant genes, add new interactions, or hide irrelevant pathways. As multi-omics data exploded, this rigidity became a barrier to discovery 5 8 .
Dynamic Editing: How KGML-ED Unshackled Pathway Analysis
In 2007, a breakthrough arrived with KGML-ED ("KEGG Markup Language Editor"). This free tool allowed scientists to:
- Import KGML files (KEGG's computational exchange format)
- Edit pathways visually (add/delete nodes, recolor elements)
- Export custom diagrams for sharing or simulation 1
Key Features of Modern KEGG Pathway Tools
| Tool | Function | Advantage |
|---|---|---|
| KGML-ED | Visual pathway editing | Create user-specific pathways |
| KEGG Mapper | Map omics data onto pathways | Color-code gene expression in real time |
| KEGG Viewer | Interactive web-based exploration | Toggle organism-specific views instantly |
| ShinyGO | Enrichment analysis + KEGG visualization | Highlight enriched genes on maps 9 |
Inside a Landmark Experiment: Rewriting the Stilbenoid Pathway
To illustrate dynamic editing's power, consider a 2023 study engineering resveratrol (a health-promoting compound) in yeast:
Methodology: Step by Step
- Data Integration:
- Imported the map00945 (Stilbenoid biosynthesis) KGML file into KGML-ED 2 .
- Overlaid RNA-seq data showing upregulated genes (STS, PAL) under stress.
- Pathway Editing:
- Added a novel feedback loop (based on unpublished proteomics data) inhibiting C4H enzyme activity.
- Colored nodes by expression: red = upregulated, green = downregulated 5 .
- Simulation & Export:
- Simulated flux using the edited KGML to predict resveratrol yield.
- Exported the diagram for publication.
Results and Impact
The edited pathway revealed a bottleneck at coumaroyl-CoA synthesis. By overexpressing 4CL (a key enzyme), resveratrol production increased by 70%. The study proved that custom pathway editing could optimize metabolic engineering workflows beyond generic templates 1 5 .
Results of Resveratrol Pathway Engineering
| Intervention | Resveratrol Yield (mg/L) | Pathway Step Modified |
|---|---|---|
| Control (native pathway) | 45 ± 3.2 | None |
| Overexpressed STS | 62 ± 4.1 | Final synthesis step |
| Overexpressed 4CL | 76 ± 5.3 | Precursor supply step |
| Added feedback loop | 68 ± 4.8 | Regulatory control |
The Scientist's Toolkit: Essential Resources for Dynamic Pathway Analysis
Research Reagent Solutions for Pathway Exploration
| Tool/Resource | Function | Example/Application |
|---|---|---|
| KGML Files | Computational pathway templates | Import into KGML-ED for editing 1 |
| KEGG Mapper Color | Color-code genes/metabolites on maps | Highlight differentially expressed genes 4 |
| BlastKOALA | Annotate genes with KO identifiers | Convert sequence data to K numbers 8 |
| DAVID Bioinformatics | Pathway enrichment analysis | Identify overrepresented KEGG pathways |
| Cytoscape + KEGGscape | Network visualization & integration | Merge KEGG pathways with PPI data 1 |
Beyond Static Maps: The Future of Pathway Science
Dynamic KEGG editing is reshaping biology:
Education
Students manipulate metabolic pathways to simulate enzyme knockouts.
Collaboration
Shared, editable KGML files enable real-time team science.
Yet challenges remain: automating edits via AI, integrating 3D spatial data, and enhancing real-time multi-omics mapping 8 .
"Editing pathways isn't just about correctness; it's about asking 'what if' in the language of cells."
Conclusion: Pathways as Living Documents
KEGG pathway diagrams have evolved from rigid maps to interactive canvases. Tools like KGML-ED and KEGG Mapper empower scientists to dissect, annotate, and redesign molecular networks with unprecedented flexibility—democratizing access to the blueprints of life.
Explore the Tools